Abstract
Key message
The molecular mechanisms of fatty acid biosynthesis and accumulation during Camellia meiocarpa and Camellia oleifera seed natural drying are characterized through transcriptome analyses.
Abstract
Camellia seed oil has been used as high quality and healthy food for over two thousand years. Seed drying affects oil quality and quantity; however, the molecular mechanisms of fatty acid biosynthesis and accumulation during the drying process remain unknown. In this study, the transcriptomes of Camellia meiocarpa and C. oleifera seed were characterized at five moisture content levels (10–50%) to identify the major processes and reveal genes affecting lipid metabolism in response to natural drying. We found a total of 111,156 unigenes by de novo assembled from RNA-Seq libraries of five moisture content levels during after-ripening of C. meiocarpa (74,016) and C. oleifera (76,374). Ten pathways were closely linked to changes in oil content and composition with 244 genes involved in fatty acid synthesis and accumulation. Gene ontology enrichment of differentially expressed genes (DEGs) indicated that fatty acid synthesis and accumulation are essential in C. meiocarpa while fatty acid accumulation in C. oleifera during natural drying process. Comparative analyses of DEGs between any two consecutive moisture contents identified six and three key unigenes in C. meiocarpa and C. oleifera seeds, respectively, and one additional unigene responsible for the difference between the two species’ fatty acid synthesis and accumulation. Natural drying has improved the quality and quantity of the camellia seed oil. The study provided: (a) global transcriptional profiles at five moisture content levels during seed natural drying, (b) highlighting transcripts putatively involved in the regulation of gene expression program and in specific processes likely essential for lipid metabolism, and (c) discovery of genes associated with oil seed quantity and quality improvement for the studied two camellia species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Camellia, family Theaceae, is well known as a tea and ornamental tree and is also considered as a valuable oil source (Robards et al. 2009). Camellia oil is extracted from a number of different species including Camellia oleifera, C. meiocarpa, C. sinensis, and C. semiserrata and has been used as a culinary oil for over two thousand years (Yang et al. 2016; He et al. 2016). Camellia meiocarpa and C. oleifera cultivated area are the largest in China; however, the former is known for its higher oil quantity and quality than the latter (Yuan et al. 2012; Xie et al. 2013).
Plant oils are of great importance for the agricultural industry and their overall consumption has been increasing by around 5% per year over the past half century (Harwood and Guschina 2013). China’s camellia oil is known as the “Eastern olive oil” (Lee and Yen 2006). Production reached 180,000 tons in 2014 with a monetary value of 42 billion Yuan and by 2020 it is expected to reach more than 250,000 tons with an output value of 100 billion Yuan (National Development and Reform Commission of China 2009). The oleic acid content of camellia oil could reach above 80%, along with high content of monounsaturated lipid, the lowest levels of saturated fats (Haiyan et al. 2007). It is reputed to aid cholesterol reduction, resistance to stress, oxidation and inflammation reduction, and neoplasma improving human immunity (Haro Bailon et al. 2014).
The amount and quality of the oil produced from camellia species can vary according to specie (Su et al. 2014; De et al. 2014), stage of development (Li et al. 2014a, b), as well as extraction (Fang et al. 2016) and drying methods (Liu et al. 2014; Hsieh et al. 2013). Before oil extraction, the drying process is a fundamental factor (Tibaldi et al. 2010) as the seed moisture content should decrease below 10% (Liu et al. 2014). It is proven that the fatty acid synthesis in the dry seed results in improving the quality of fatty acid through increasing the unsaturated fatty acid and decreasing the saturated fatty acid (Hsieh et al. 2013; Wang et al. 2011). Seed drying management not only improves the quality and quantity of oil in camellia species but also other oilseed plants (Tibaldi et al. 2010; Dušková et al. 2016; Evaristo et al. 2016; Tibaldi et al. 2013). Several transcriptome analyses showed that drying management affects transcript abundances and the genetic regulatory networks in seeds, resulting in the selective change of specific transcripts (Rohini et al. 2016; Fu et al. 2016; Pan et al. 2016). However, very little is known about the relationship with expression of genes involved in the fatty acid pathway and oil accumulation during drying management.
The main objective of the present study is to gain knowledge on the molecular mechanisms underlying the role of drying management on the fatty acid pathway and oil accumulation in camellia matured seed and to identify transcripts putatively related to lipid metabolism. To accomplish this goal, we performed comparative transcriptomic analyses to document the dynamics of five different levels of seed moisture content (10, 20, 30, 40 and 50%) in two camellia species C. oleifera and C. meiocarpa under natural drying condition. We expect that this research will provide insights into how natural drying contributes to lipid metabolism for the two camellia species.
Materials and methods
Plant material
In 2012, mature fruits of C. oleifera and C. Meiocarpa, with removed fuzz, deep and shiny color, and showing micro-cracking, were collected from the four crown cardinal directions of superior trees growing at the Minhou Tongkou State Forest Farm (26°05′N, 119°17′E), Fujian Province, China. For each species, one tree located on each of the upper, middle, and lower southeast slopes of the State Forest Farm were selected. Fruits were collected from healthy trees with known good fruit production (>15 kg for 3 consecutive years). The collected fruits were mixed per species, and placed in a ventilated room until they naturally cracked and seeds were extracted by manual shell cutting. The seed moisture content at the time of extraction was close to 50%. Seeds were naturally dried and their moisture content was determined daily. Over time, seed samples were collected at moisture content of 50, 40, 30, 20 and 10% and were sequentially identified as T01 to T05 and T06 and T10 for C. meiocarpa and C. oleifera, respectively, and were flash frozen in liquid nitrogen and stored at −80 °C until RNA extraction.
Ultrastructure
The seeds of different moisture content (10, 20, 30, 40, and 50%) of the two species were immersed in 2.5% glutaraldehyde solution. After three washes with phosphate buffer (0.1 M, pH 7.2), seeds were post-fixed for 4 h at room temperature under 1% osmium tetroxide. Seeds were washed three times with phosphate buffer (0.1 M, pH 7.2), dehydrated in graded ethanol series (30, 50, 70, 80, 90, 95, and 100%, 15 min each), then replaced by acetone. Seeds were infiltrated with epoxy resin 812, and polymerized at the 37 °C oven for 24 h, then at the 60 °C oven for 48 h. Ultrathin sections (100 nm) of seeds were cut with EM UC6 microtome (Leica, Germany), using diamond knives and picked up with copper grids (200 mesh). Sections were double stained with 2% uranyl acetate and 1% lemon lead acid, observed and photographed under Tecnai-12 transmission electron microscope (FEI, America).
To calculate the area fraction of lipid droplets in matured seeds during natural drying, photos were taken in same 72 × 72 lattice system based on calibration curves of 10 μm. Three images from each section were selected and analyzed by Image Analyzer software (1.37). The hit points of lipid droplets were marked manually on blinded images, and counted by the software. The area fractions were estimated by the ratio of hit points/(total points − cell wall points) (Li et al. 2014a, b).
RNA isolation
Following manufacturers’ instructions, total RNA was: (1) extracted from pooled samples of seeds using RNAsimple® kit (Tiangen Biotech, Beijing, China), (2) checked for degradation and contamination after monitoring on 1% agarose gels, (3) purity was determined using the NanoPhotometer® spectrophotometer (IMPLEN, CA, USA), (4) concentration was measured using Qubit® RNA Assay Kit in Qubit® 2.0 Flurometer (Life Technologies, CA, USA), and (5) integrity was assessed using the RNA Nano 6000 Assay Kit using the (Agilent Bioanalyzer 2100 system, Agilent Technologies, CA, USA).
cDNA library construction, and high throughput sequencing
After total RNA extraction, mRNA was purified using poly-T oligo-attached magnetic beads (Connell et al. 2012). A total of 3 μg RNA was used as the input for each RNA sample preparation. Sequencing libraries were generated using a NEBNext UItra RNA Library Prep Kit (Illumina, NEB, USA) following the manufacturer’s instructions. Index codes were assigned to attribute the sequences to each sample. The clustering of the index-coded samples was performed on a cBot Cluster Generation System using TruSeq PE Cluster Kit v3-cBot-HS (Illumina) according to the manufacturer’s instructions. After cluster generation, the library preparations were sequenced on an Illumina Hiseq 2000 platform and paired-end reads were generated (Akazawa et al. 2013).
Data analysis, transcriptome assembly, gene functional annotation
Raw reads of fastq format were first processed through in-house perl scripts. In this step, clean reads were obtained after removing reads containing adapters, low quality reads (with ambiguous “N” bases >5%, and those with >10% of Q value <20 bases) (Gao et al. 2014). At the same time, the clean reads from each library were evaluated for Q20, Q30, GC-content, N-content, CycleQ 20 (a base quality greater than 20 and an error probability of 0.01). All the downstream analyses were based on high quality clean data.
Clean reads from the ten libraries (T01–T10) were assembled using Trinity, which is an efficient method for de novo assembly of full-length transcripts without a reference genome (Grabherr et al. 2013). Then the clean reads were clustered based on nucleotide sequence with parameters set at K-mer length of 25 and a similarity of 80%. First, we combined all the clean reads to form contigs and calculated the distance and relation among these contigs using Trinity software. Then these contigs were connected to obtain transcripts with consensus sequences that could not be extended on either end, and further clustered them into unigenes for annotation. To calculate abundance estimation for each unigene, clean data were mapped back onto the assembled transcriptome, and read count for each unigene was obtained from the mapping results. Fragments per kilobase per transcript per million mapped reads (FPKM) was used to quantify gene expression abundance (Mortazavi et al. 2008).
The program BLAST was used to assign putative functions to the assembled unigenes. Unigenes were aligned against the Nr (NCBI non-redundant protein sequences), Nt (NCBI non-redundant nucleotide sequences), Pfam (Protein family), KOG/COG (Clusters of Orthologous Groups of proteins), Swiss-Prot (A manually annotated and reviewed protein sequence database), KO (KEGG Ortholog database), and GO (Gene Ontology) using the Blast algorithm with a E value cut-off of 10−5. Clean reads were mapped back onto the assembled transcriptome; read count for each unigene was obtained from the mapping results. To annotate the unigenes with GO terms, the best BLASTx hit from Nr database for each transcript was submitted to BLAST2GO platform (Conesa and Götz 2008), and GO terms for each unigene were retrieved based on the relationship between gene names and GO terms, EC number was assigned and parsed based on the BLAST2GO results. To determine metabolic pathways, KEGG mapping was used. KEGG pathways were retrieved from the KEGG web server (KAAS—http://www.genome.jp/kegg/) (Kanehisa et al. 2008). KAAS provides functional annotation of putative genes by BLAST comparison against the KEGG GENES database. The output includes KO (KEGG Orthology) assignments and automatically generated KEGG pathways that are populated with the KO assignments. For coding sequences (CDS) annotation, BLASTx alignments were carried out between unigenes against Nr, KEGG, Swiss-Prot, and COD protein databases, and the transcriptional directions and coding frame of unigenes were predicted from BLASTx results. Unigene CDS with no specific BLASTx matches were predicted by ESTScan (Iseli et al. 1999).
Differential expression analysis
Differential expression analysis of two samples was performed using the DEGseq R package (Wang et al. 2009). The P value was adjusted for multiple testing using the Benjamini–Hochberg method. P value <0.005 and |log2 (foldchange)| ≥1 was set as the threshold for significantly differential expression (Renfro et al. 2011). Gene Ontology (GO) enrichment analysis of the differentially expressed genes (DEGs) was implemented by the GOseq R packages based on Wallenius non-central hyper-geometric distribution was used to test the statistical enrichment of DEGs in KEGG pathways (Mao et al. 2005; Young et al. 2010).
Quantitative RT-PCR analysis
Reverse transcription quantitative real-time PCR (qRT-PCR) of a set of 8 DGEs was carried out to validate the expression profile (Table 1). The concentration of the RNA samples was determined by Nanodrop 2000 and all cDNAs were synthesized from 1.5 μl of total RNA using HiFiScript Quick gDNA Removal cDNA Kit (CWBIO, Beijing, China). Specific primers were designed using Primer Premier 6 software (Zhang et al. 2013). Quantitative real-time PCR experiments were then performed in Bioer 96 plus using UltraSYBR Mixture. For the unigenes tested, 3 biological replicates were used and the reaction mixtures were performed in a final volume of 20 μL containing 10 μL of 2× UltraSYBR Mixture, 500 nM of each primer, 1 μL of cDNA as template and 8 μL of RNase-Free Water. The amplification program was the same for all unigenes tested: 95 °C for 10 min, 40 cycles of 15 s at 95 °C, 1 min at 60 °C. The 2−∆∆Ct method was used to calculate individual unigene’s relative expression levels (Livak and Schmittgen 2001). A calibrator sample was used in each plate to normalize the values obtained and the potential differences among plates. Normalization was carried out with reference gene ACTIN (Reddy et al. 2013; Shi et al. 2011; Zhou et al. 2013). PCR and agarose gel analyses were used to verify the absence of non-specific amplification prior to qRT-PCR. Additionally, following reactions DNA melt curves were created for each primer combination to confirm the presence of a single product. The average of two technical repeats was used for each reaction, and the standard error of the mean was calculated for the three biological replicates. All primers pairs for qRT-PCR are listed in Table 1.
Results
Lipid drops in different moisture content
In both camellia species, the area fraction of lipid droplets showed steady increase in size with deceasing moisture content (Fig. 1; Table 2). Significant difference between all moisture contents’ area fraction of lipid droplets were observed for C. meiocarpa, with the largest change noted during T03-04 (20–30% moisture content), while only significant difference was noted between T07 (40%) and T08 (30%) for C. oleifera (Fig. 1; Table 2). At the 10% moisture content, C. meiocarpa showed more lipid droplets than C. oleifera, but this trend was reversed for the other moisture contents (Fig. 1; Table 2).
Ultrastructure section electron microscope of seeds lipid laden by natural drying treatment in two studied camellia species. a The cells contain accumulations of lipid droplets (LD) in C. meiocarpa seeds. b The cells contain accumulations of LD in C. Oleifera seeds. 1–5 The cells were in 50, 40, 30, 20 and 10% moisture content in two camellia species, respectively. CW cell wall, P plastids containing starch grains. Scale bars 10 μm
Illumina paired-end sequencing and de novo transcriptome assembly
We obtained a total of 78.18 Gb clean data from the ten (T01–T10) C. meiocarpa and C. oleifera libraries, with per library ≥7.19 Gb (Table 3). After stringent quality assessment and data filtering, we obtained >28.56 million reads with no N per library from end of cDNA fragment with 100% Q20 bases. The GC-content averages were 46.51 and 45.56% for C. meiocarpa and C. oleifera, respectively (Table 3). Furthermore, the CycleQ 20% was 100% for all libraries and a base quality score of >30 of clean reads was above 86.15% (Table 3). This suggested that the sequencing was highly accurate.
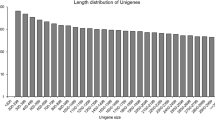
Using the Trinity de novo assembly, all high-quality reads were assembled into transcripts, resulting in 197,381 (mean length of 1006.06 bp) and 209,913 (mean length of 1041.81 bp) for C. meiocarpa and C. oleifera, respectively (Table S1). Then these transcripts were clustered based on nucleotide sequence identity, we harvested a total of 111,156 unigenes with an N50 length of 1149 bp and mean length of 838.1 2 bp, which included 26,282 unigenes (23.64%) with lengths greater than 1 kb. C. meiocarpa and C. oleifera had 74,016 (N50 length of 1038 bp and average length of 789.61) and 76,374 (N50 length of 1046 bp and average length of 798.23) unigenes, respectively (Table 4, S1). These results indicated that the throughput and sequencing quality were high and suitable for the following analysis.
Functional annotation of transcriptome
A total of 98,398 coding sequence (CDS) were extracted from BLASTx results with 111,156 unigene sequences being translated into protein sequences (see Fig. 2a for the length distribution of CDS). To better assign unigene names and annotations, CDS and predicted proteins, all assembled unigenes were first searched against the NCBI Nr and Swiss-Prot databases using BLASTx with an E value cut-off of 10−5. And as a result, 12,932 (29%) unigenes showed significant similarity with sequences of Vitis vinifera, and about 5% of the mapped sequences have a high similarity with sequences of Theobroma cacao (2341 unigenes), Coffea canephora (2300 unigenes) and Nelumbo nucifera (1973 unigenes) (Fig. 2b). Only 44,886 unigenes (40.38%) were annotated with an E value threshold of 10−5 by performing BLASTX search against diverse protein databases, revealing that 11,958 (10.76%) unigenes had significant matches with sequences in COG databases, 23,650 (21.28%) in GO databases, 7931 (7.14%) in KEGG databases, 24,328 (21.89%) in KOG databases, 26,981 (24.27%) in Pfam databases, 29,160 (26.23%) in Swissprot databases, and 44,032 (39.61%) in Nr protein databases, respectively (see Table 5, for the overall functional annotation for the studied camellia species).
Functional classification by gene ontology and eukaryotic orthologous groups
Gene ontology (GO) was used to classify the functions of the assembled transcripts and describe gene products in terms of their associated biological processes, cellular components, and molecular functions. A total of 128,299 predicted proteins were categorized into 52 functional groups with 59,884 (46.68%), 39,149 (30.51%), and 29,266 (22.81%) unigenes grouped in biological processes, cellular components, and molecular functions, respectively (Fig. 3). In the biological processes category, metabolic (12.38%) and cellular processes (10.53%) were the predominant groups, followed by single-organism process (8.12%), response to stimulus (3.43%), and biological regulation (3.35%). In the cellular components category, cell part (7.13%) and cell (7.09%) were the most representative ones, followed by organelle (5.21%) and membrane (4.14%). With regard to molecular function category, the predominant categories were catalytic activity (10.13%) and binding (9.67%), followed by transporter activity (1.22%) (Fig. 3). These results indicated that most of unigenes were responsible for fundamental biological regulation and metabolism.
To further evaluate the completeness of the transcriptome and effectiveness of the annotation process, the annotated unigenes were compared with the Eukaryotic Orthologous Groups (KOG) database for functional prediction and classification. In total, 27,156 annotated putative proteins were classified functionally into 25 KOG groups (Fig. 4). Among the 25 categories, the largest group was “general function prediction only” (26.95%), second was “signal transduction mechanisms” (9.81%), followed by “posttranslational modification, protein turnover, chaperones” (8.64%), and “transcription” (5.24%) (Fig. 4). The three smallest groups were “cell motility” (0.04%), “nuclear structure” (0.40%), and “extracellular structures” (0.46%). However, there were 868 unigenes (3.20%) associated with “lipid transport and metabolism”.
Pathway analysis during camellia seed after-ripen
Pathway-based analysis was used to assist in understanding unigenes’ functions and interactions. Following the Kyoto Encyclopedia of Genes and Genomes (KEGG) database, 7931 unigenes were assigned to 116 pathways (Table S2). Pathway mapping assigned 927 unigenes into 103 functional pathways (KO groups) (Table S3). Plant hormone signal transduction (ko04075, 44, 15.17%) had the largest number of KO identifiers, followed by RNA transport (ko03013, 40, 15.15%), protein processing in endoplasmic reticulum (ko04141, 38, 15.14%) and spliceosome (ko03040, 31, 10.65%) (Table S2). Notably, 10 pathways (85 unigenes) were closely linked to changes in oil content and composition, which were Glycolysis/Gluconeogenesis (19 unigenes), starch and sucrose metabolism (28 unigenes), pyruvate metabolism (20 unigenes), glycerophospholipid metabolism (10 unigenes), fatty acid metabolism (4 unigenes), glycerolipid metabolism (9 unigenes), fatty acid biosynthesis (12 unigenes), biosynthesis of unsaturated fatty acids (7 unigenes) and fatty acid elongation in mitochondria (1 unigenes) (Table 6, S2).
Classifying unigenes by KEGG Orthology resulted in 10 pathway related with lipid metabolism, which had the same KO identifiers to each other (Table 6; Fig. 5). These pathways collectively configured a network for the development of oil body droplet (Fig. 5). The network of lipid metabolism sustained by starch and sucrose metabolism is divided into three parts, one part is related to the fatty acid synthesis supported by pyruvate metabolism which included fatty acid biosynthesis, fatty acid elongation and biosynthesis of unsaturated fatty acid. The second part is dedicated to triacylglycerol synthesis and included glycolysis/gluconeogenesis, glycerolipid metabolism and glycerophospholipid metabolism. These two parts combined to form the oil body droplet. The third part is associated with the fatty acid metabolism (Fig. 5). These lipids pathways provided critical clues to the identification and characterization of key functional genes involved in unsaturated fatty acid (FA) and triacylglyceride (TAG) biosynthesis during the natural seed drying process of the two camellia species.
Diagram depicting lipid metabolism network in the two studied camellia species. Ellipses contained the metabolic pathway and polygon oil droplets. Names on arrows refer to the key intermediate products produced from upstream to downstream metabolism. Brackets refer to the enzyme code roles (K00844 hexokinase, K01810 glucose-6-phosphate isomerase, K00627 dihydrolipoamide acetyltransferase, K00161 pyruvate dehydrogenase E1 component alpha subunit, K00128 aldehyde dehydrogenase, K00162 pyruvate dehydrogenase E1 component beta subunit, K00382 dihydrolipoamide dehydrogenase, K01962 acetyl-CoA carboxylase carboxyl transferase subunit alpha, K02160 acetyl-CoA carboxylase biotin carboxyl carrier protein, K01961 acetyl-CoA carboxylase, K03921 acyl-[acyl-carrier-protein] desaturase, K00059 3-oxoacyl-[acyl-carrier protein] reductase, K13506 glycerol-3-phosphate O-acyltransferase, K00901 diacylglycerol kinase)
Identification and characterization of unigenes involved in fatty acids and triacylglycerols biosynthesis
To identify unigenes showing changes in expression at the studied five moisture content levels, the expression level of each unigene was determined using Fragments Per Kilobase of transcripts per Million mapped reads (FPKM) method (Mortazavi et al. 2008). A total of 44,886 unigenes were annotated, with 244 unigenes involved in FA biosynthesis and TAG assemblage (Table 7, S4).
Differentially expressed genes (DEGs) during after-ripening
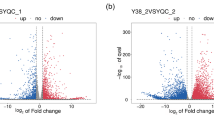
Differentially expressed genes (DEGs) were identified by pairwise sample comparisons, and the majority of unigenes were identified as differentially expressed in at least one comparison. We calculated the number of up- and down-regulated unigenes according to decrease in moisture content (Table 8). The data sets from each moisture content level were compared, and Pearson product–moment correlation was determined between sets. The observed low correlation coefficients (range 0.230–0.972) indicated the presence of significant difference between the investigated moisture content levels (Fig. 6; Table S5). In this analysis, we used the tissues from a specific moisture content level as the control for comparison with its consecutive moisture content (i.e., lower was the control for its next higher moisture content sample). This was done within and across species and in this case C. oleifera served as the control. A total of 17,996 unigenes showed ≥twofold expression difference between the two libraries (Table 8).
In-depth analysis of fatty acid synthesis and accumulation by natural drying
DEGs provide clues about the molecular events related to the functional roles they play during the seed natural drying specifically those related to synthesis and accumulation of fatty acid. To evaluate the potential functions of genes that showed significant transcriptional changes in fatty acid synthesis and accumulation between the two contrast libraries, DEGs were further classified into subsets with Gene Ontology (GO) enrichment analysis. The results showed that 13 and 23 GO terms in biological processes and molecular function, respectively, were related with fatty acid synthesis and accumulation (Tables S6, S7).
Comparative analyses of the DEGs between any two consecutive moisture content samples within and between C. meiocarpa and C. oleifera are illustrated in Fig. 7. For C. meiocarpa, 77 genes were common to all four comparisons, whereas 1255, 777, 936, and 897 genes clusters were restricted to T01 vs T02, T02 vs T03, T01 vs T04, and T01 vs T05 comparisons, respectively (Fig. 7a). For C. oleifera, 24 genes also were common to all four comparisons, whereas 662, 1277, 721, and 1027 gene clusters were restricted to T06 vs T07, T07 vs T08, T08 vs T09, and T09 vs T10 comparisons, respectively (Fig. 7b). Between species comparisons produced 189 genes in common to all five comparisons, whereas 1892, 1208, 1470, 949, and 1248 gene clusters were restricted to T07 vsT02, T06 vs T09, T06 vs T10, and T10 vs T05 comparisons, respectively (Fig. 7c). To get key unigenes associated with fatty acid synthesis and accumulation by natural drying in the two camellia species, significant DEGs within each comparison were selected for further analysis (Tables S4, S8, S9, S10).
Validation of the differential expression profiles
According to the genes annotated in KEGG, eight DEGs were selected for expression analysis, in which 7 transcripts belonged to lipid metabolism and one to RNA regulation (Table 1). Transcript abundance was analyzed by qRT-PCR in C. oleifera and C. meiocarpa seed at the 5 different moisture content levels (Figs. 8, 9, S1; Table S11). Correlation coefficient between qRT-PCR and RNA-Seq in the two species was 0.77 on the average, 0.86 for C. meiocarp and 0.68 for C. oleifera. Most of the genes showed strong or moderately strong correlation. These results confirmed the validity of the transcriptome results.
Validation of the RNA-Seq transcript profiles in C. meiocarpa seed at different moisture content levels. Comparison of transcripts expression patterns from RNA-Seq data and from reverse transcription quantitative real-time PCR (qRT-PCR). The numbers above the graphics correspond to Pearson’s correlation value
Validation of the RNA-Seq transcript profiles in C. oleifera seeds at different moisture content level. Comparison of transcripts expression patterns from RNA-Seq data and from reverse transcription quantitative real-time PCR (qRT-PCR). The numbers above the graphics correspond to Pearson’s correlation value
Discussion
Fatty acids biosynthesis and accumulation
The annotated unigenes were involved in at least 32 plant enzymes in the fatty acid biosynthesis and accumulation (Table S4). These data were integrated and compiled to propose schematic metabolic pathways that lead to oil accumulation in the studied two camellia species (Fig. 10). These results suggested that fatty acid biosynthesis, elongation, desaturation, and TAG biosynthesis were all activated in the seed natural dry process (Fig. 10) (Wang et al. 2014). There were seven fatty acid enzymes (LACS, HCD, SAD, ∆12D, PAP, Oleosin and Caleosin) related to unigenes at an average of FPKM >100 in the two studied camellia species (Table S4), indicating that the C. Meiocarpa and C. Oleifera seed experienced fatty acid modification and accumulation during the natural drying process. This was confirmed by the increase in area fraction of lipid droplets accompanied with decrease in the moisture content in the two camellia species (Fig. 1; Table 2).
Scheme of the reactions involved in fatty acid biosynthesis and triacylglycerol assembly in the studied camellia species. (Lipid substrates abbreviation phosphatidylcholine (PC); fatty acids (FA); lysophosphatidylcholine (LPC); glycerol 3-phosphate (G3P); lysophosphatidic acid (LPA); phosphatidic acid (PA); diacylglycerol (DGA); triacylglycerol (TAG). Enzyme/protein abbreviations malonyl-CoA:ACP transacylase (MAT); biotin carboxyl carrier protein of acetyl-CoA carboxylase (BCCP); biotin carboxylase (BC); acetyl-coenzyme A carboxylase carboxyl transferase subunit (CT); 3-oxoacyl-[acyl-carrier-protein] synthase III (KASIII); 3-oxoacyl-[acyl-carrier-protein] reductase (KER); hydroxyacyl-ACP dehydratase (HAD); enoyl-(acyl-carrier-protein) reductase (EAR); 3-oxoacyl-[acyl-carrier-protein] synthase I (KASI); 3-oxoacyl-[acyl-carrier-protein] synthase II (KASII); acyl-[acyl-carrier-protein] desaturase (∆9D(SAD)); palmitoyl-acyl carrier protein thioesteraseB (FATB); palmitoyl-acyl carrier protein thioesterase A (TATA); long chain acyl-CoA synthetase (LACS); 3-ketoacyl-CoA synthase (KCS); 3-hydroxyacyl-CoA dehydrogenase (HCD); very-long-chain enoyl-CoA reductase (ECR); phospholipase A (PLA); lysophospholipid acyltransferase (LPCAT); omega-6 fatty acid desaturase (∆12D); omega-3 fatty acid desaturase (∆15D); delta(8)-fatty-acid desaturase (∆8D); fatty acid desaturase (4∆5D); sphingolipid delta(4)-desaturase (∆4D); glycerol-3-phosphate acyltransferase (GPAT); glycerol-3-phosphate O-acyltransferase (LPAAT); phosphatidate phosphatase (PAP); phosphatidylcholine:diacylglycerol cholinephosphotransferase (PDCT); diacylglycerol acyltransferase (DGAT); Phospholipid:diacylglycerol acyltransferase (PDAT))
We identified new fatty acid enzymes (BCCP, BC, LPCAT, ∆9D, ∆8D, ∆5D, Oleosin and Caleosin) compared to previously published C. Oleifera transcriptome dataset (Xia et al. 2014), and new fatty acid enzymes (KCS, HCD, CER, ∆9D, ∆15D, ∆8D, ∆5D, Oleosin and Caleosin) compared to C. reticulata transcriptome dataset (Yao et al. 2016). These observations were due to using different study material across studies (i.e., matured seed in the present study vs. buds, leaves, flowers, shoots, and immature fruits in the referenced studies) (Xia et al. 2014; Yao et al. 2016).In the fatty acid biosynthesis, we found 10 BC genes, 5 BCCP and 5 CT, which can freely combine to form the ACCs (Fig. 10; Table 7). A similar expression pattern was observed in tung tree (Galli et al. 2014), Jatropha curcas (Xu et al. 2011), and oil palm seeds (Nakkaew et al. 2008). This step is most likely a feedback inhibition in fatty acid synthesis (Galli et al. 2014; Andre et al. 2012). So expression of genes were low in the fatty acid biosynthesis in this study (Table S4), and BC, BCCP and CT failed to be identified in immature camellia seed (Yao et al. 2016) and free fatty acids were generated from fatty acids acyl-CoA under the action of long chain acyl-CoA synthetase (LACS) (Nobusawa et al. 2013) (Fig. 10). This process is related to 21 LACS during natural seed drying, similar to that observed in Sacha Inchi (Wang et al. 2012) and tree peony (Li et al. 2015). For the formation of unsaturated fatty acids, 34 unigenes encoding fatty acid desaturase were identified, including six kinds of FAD (SAD, ∆15D, ∆12D, ∆9D, ∆8D and ∆5D) (Table 7). However, some oil seed plants failed to identify ∆8D, ∆5D, such as Sacha Inchi (Wang et al. 2012), tung tree (Galli et al. 2014) and Yellow Horn (Liu et al. 2013). There were four unigenes expressed over 100 RPKM, in which three (CL32923Contig1, CL27152Contig1 and CL21434Contig1) were annotated to SAD, and the remaining one (CL32549Contig1) to ∆12D, which are correlated with fatty acid composition and content (Zeng et al. 2014), and corresponded to fast oil accumulation stage (Galli et al. 2014; Wang et al. 2012; Rajwade et al. 2014). This was verified by unsaturated fatty acid content, in which C18:1 was the highest and followed by C18:2 (Yang et al. 2016; Zhang et al. 2015; Long et al. 2012).
The major fate of newly synthesized fatty acids is for TAG assembly as acyl-CoA via two different pathways, catalyzed by phospholipid:diacylglycerolacyltransferase (PDAT) and diacyl-glyceroltransacylase (DGTA) (Simpson and Ohlrogge 2016), in which both pathways were present in the two camellia species’ seeds during the natural drying process. However, lack of cholinephosphotransferase (CPT) was observed compared with the normal biosynthesis of triacylglycerol in seed (Baud and Lepiniec 2010; Lock et al. 2009). DAG was converted to TAG in camellia seed with the involvement of 11 DGAT and 5 PDAT unigenes (Table 7), which indicated that DGAT was the main way to camellia seed oil formation. Drying management may brought about over expression of DGAT (Weselake et al. 2008). Oil bodies are the main form of oil in mature seed, internally containing liquid TAG and externally forming phospholipid monolayer (Jolivet et al. 2013). We identified the presence of oleosins and caleosins proteins in camellia seed, and involved in 4 oleosin unigenes (CL734Contig1, CL422Contig1, CL32127Contig1 and CL25464Contig1) with expression over 1000 RPKM in average (Table 7, S4). So oleosin is the most abundant structural protein in camellia oil bodies like other oilseed crops (Hyun et al. 2013; Yu et al. 2015; Parthibane et al. 2012).
Changes of fatty acids biosynthesis and accumulation in C. meiocarpa during natural drying process
During seed natural drying, C. meiocarpa exhibited 3324 more differentially expressed unigenes in T01 vs. T02 libraries, then the number of DEGs steadily declined to 3000 with seed moisture loss, but in T04 vs. T05 libraries the number of DEGs sharply declined to 1626 unigenes (Table 8). This indicates that seed metabolic activity decreased during the drying process and T04 (20% moisture content) was the turning point. Correlation coefficients of DEGs in T01 vs. T02, T02 vs. T03, T03 vs. T04, and T04 vs. T05 were 0.928, 0.972, 0.728, and 0.966, respectively, with T03 vs. T04 showing the lowest relation during natural drying (Fig. 6; Table S5). Additionally, the correlation of T01 with T04 was 0.595 showing the lowest value among all comparisons (Table S5), indicating that T04 (20% moisture) had the highest metabolism.
To understand the lipid metabolism at different moisture content in C. Meiocarpa seeds, we analyzed the role of DEGs with fatty acids biosynthesis and accumulation based on GO enrichment analysis. In the biological processes, stage T01 vs. T02 (40–50% moisture content) showed the highest DEGs number in lipid biosynthetic, long-chain fatty acid metabolic, phospholipid biosynthetic and glycerolipid biosynthetic processes (Table S7), indicating that the 40–50% moisture content had the highest triacylglycerol synthesis. T02 vs. T03 (30–40% moisture content) had the highest DEGs in the regulation of unsaturated fatty acid biosynthetic process and unsaturated fatty acid metabolic process with the highest unsaturated fatty acid synthetic. Additionally, stage T03 vs. T04 (20–30% moisture content) produced the highest DEGs number in 6 GO terms (fatty acid biosynthetic process, cellular lipid, fatty acid, and glycerolipid metabolic processes, lipid modification, and lipid storage) (Table S7). These indicate that maximum fatty acid synthetic and lipid storage occurred at 20–30% moisture content, which is identified by highest change in area fraction of lipid drops at 20–30% moisture content (Fig. 1; Table 2). However, T04 vs. T05 (10–20% moisture content), showed a dramatic reduction in the number of DEGs compared the other three libraries (Table S7), indicating obvious reduction in lipid metabolism. From the molecular function aspect, 5 GO terms over 2 DEGs were detected (fatty acid synthase, acetyltransferase, O-acyltransferase, and N-acyltransferase activities, and lipid binding), confirming that after-ripening processed the fatty acid synthesis and accumulation (Table S7). So for C. Meiocarpa, quantity increase of fatty acids mainly occurred in 30–50% moisture contents and quality increase mainly in 30–40% moisture content.
To identify key unigenes, we comparatively analyzed the DEGs between any two consecutive moisture content samples. A total of 41 significant DEGs in fatty acid synthesis and accumulation were detected, 12 significant DEGs expressed over 10.00 FPKM, in which Group1_Unigene_BMK.31364 (Oleosin) was largest, followed by Group1_Unigene_BMK.39872 (Oleosin) and CL24499Contig1(HAD) (Table S8). And four DEGs (CL21514Contig1(KAR), CL24499Contig1 (HAD), CL5860Contig1 (∆15D) and Group2_Unigene_BMK.25584 (∆8D)) had significance at three stages (Table S8). So during C. meiocarpa natural seed drying process, six unigenes (Group1_Unigene_BMK.31364, Group1_Unigene_BMK.39872, CL21514Contig1, CL24499Contig1, CL5860Contig1 and Group2_Unigene_BMK.25584) were key unigenes.
Changes of fatty acids biosynthesis and accumulation in C. Oleifera during natural drying process
Significant DEGs were observed during seed natural drying process with 2107 unigenes (1093 up-regulated and 1014 down-regulated) in the T07 vs. T08 libraries (Table 8), indicating that the seed vital metabolisms reached its peak during this moisture content transition (i.e., from 40 to 30%). The correlation coefficients of DEGs in T06 vs. T07, T07 vs. T08, T08 vs. T09, and T09 vs. T10 were 0.947, 0.947, 0.961, and 0.897, respectively (Fig. 6; Table S5), with little differences among them, indicating modest rate of the metabolism in C. oleifera.
To understand the lipid metabolism at different moisture content in C. oleifera seeds, DEGs of fatty acids biosynthesis and accumulation were subjected to GO enrichment analysis. The results indicated that in the biological processes, T07 vs. T08 (30–40% moisture) had the highest DEGs number representing 7 GO terms (lipid biosynthetic, fatty acid biosynthetic, lipid modification, fatty acid metabolic processes, cellular lipid metabolic, long-chain fatty acid metabolic and phospholipid biosynthetic) (Table S7). This confirms that the 30–40% moisture content had the highest lipid metabolism, identified by the highest change in area fraction of lipid droplets (Fig. 1; Table 2). The moisture content transition of 20–30% (T08 vs. T09) produced high DEGs that were related to unsaturated fatty acid biosynthetic and metabolic processes, while 10–20% (T09 vs. T10) DEGs were related to lipid storage and glycerolipid biosynthetic and metabolic processes (Table S7), indicating that 20–30% moisture content had highest fatty acid modification and 10–20% moisture content had highest triacylglycerol synthesis and lipid storage (Table S7). From the molecular function side, there were 3 GO terms over 2 DEGs which were involved in acetyltransferase activity, lipid binding and N-acyltransferase activity, indicating that the main activity was related to fatty acid accumulation (Table S7). So for C. Oleifera, quantity increase of fatty acids mainly occurred in 30–40% moisture contents, and quality increase mainly in 20–30% moisture content.
To identify key unigenes involved in fatty acid synthesis and accumulation, we comparatively analyzed DEGs between any two consecutive moisture content samples and a total of 23 significant DEGs showed the largest expression located in Group1_Unigene_BMK.39872 (Oleosin) followed by Group1_Unigene_BMK.31364 (Oleosin) and CL23502Contig1 (∆12D) (Table S9). So during C. oleifera seed drying process, three unigenes (Group1_Unigene_BMK.31364, Group1_Unigene_BMK.39872 and CL23502Contig1) were key unigenes.
Comparison of fatty acids biosynthesis and accumulation between C. meiocaipa and C. oleifera during natural drying process
The two camellia species differed substantially in the number of DEGs, up-regulated, and down-regulated unigenes with the highest number of 4819 unigenes in the T06 vs. T01 libraries (50% moisture) (Table 8). The correlation coefficients of DEGs in T06 vs, T01, T07 vs. T02, T08 vs. T03, T09 vs. T04, and T10 vs. T05 were 0.347, 0.483, 0.464, 0.901, and 0.703, respectively (Table S5; Fig. 6), with T06 vs. T01 being the lowest, indicating that 50% moisture content had the biggest difference in gene expressions in both species.
The two species utilized special enzymes for their lipid metabolism, with C. meiocaipa utilizing BC, BCCP, MAT, KAR, HAD, and FATB for fatty acid biosynthesis, SAD for fatty acid desaturation, GPAT and LPCAT for TAG biosynthesis, while C. oleifera utilized ∆12D for fatty acid desaturation and LPAAT for TAG biosynthesis. However, both species utilized the same enzymes for the very-long-chain fatty acid elongation and lipid storage (Tables S8, S9), indicating that during the drying process C. meiocarpa had more activity in fatty acid synthesis and elongation, fatty acid desaturation of plastid, but C. oleifera had more activity in fatty acid desaturation of ER. This clearly explains the observed higher oleic acid (C18:1, SAD(∆9D)) in C. meiocarpa and linoleic acid (C18:2, ∆9,12D) and stearic acid (C18:0) in C. Oleifera (Yang et al. 2016; Zhang et al. 2015; Long et al. 2012). There were three DEGs (CL6765Contig1 (PDAT), CL21434Contig1 (SAD), and CL19261Contig1 (LACS)) significantly expressed at all moisture content during drying process in the two species. The highest difference was observed for CL21434Contig1 (SAD) with 546.27 FPKM, followed by CL6765Contig1 (PDAT) with 23.18, FPKM and CL19261Contig1 (LACS) with 23.08 FPKM (Table S10). So CL21434Contig1 (SAD) represented the key unigenes in difference of the two camellia species for FA biosynthesis and TAG assemblage.
Conclusions
RNA-Seq analysis of transcript abundances during seed natural drying process in C. meiocarpa and C. Oleifera has facilitated a global investigation of unigene expression in fatty acid synthesis and accumulation at five specific moisture content levels and comprehensively describe the change of the differential transcriptional events that occurred within the two species. We harvested a total of 111,156 unigenes with 74,016 and 76,374 representing C. meiocarpa and C. Oleifera, respectively. A total of 98,398 coding sequence were extracted from BLASTx and were translated into protein sequences, of which 44,886 were annotated in protein databases. A total of 247 unigenes were involved in fatty acid biosynthesis and accumulation. These unigenes provided a comprehensive molecular biology background for fatty acid biosynthesis and accumulation in C. meiocarpa and C. Oleifera by drying management.
In summary, seed drying was helpful to increase the oil content and improve the quality of fatty acid. For C. Meiocarpa, quantity increase mainly occurred in 30–50% moisture contents and quality increase mainly in 30–40% moisture content. For C. Oleifera, quantity increase occurred in 30–40% moisture contents, and quality increase mainly in 20–30% moisture content. For lipid metabolism pathway, C. Meiocarpa focuses on fatty acid synthesis and accumulation, and C. Oleifera on fatty acid accumulation. For key different unigenes expressed in fatty acid biosynthesis and accumulation during natural drying process, there were six key unigenes (Group1_Unigene_BMK.31364, Group1_Unigene_BMK.39872, CL21514Contig1, CL24499Contig1, CL5860Contig1 and Group2_Unigene_BMK.25584) in C. meiocarpa seeds, three key unigenes (Group1_Unigene_BMK.31364, Group1_Unigene_BMK.39872 and CL23502Contig1) in C. Oleifera seeds, CL21434Contig1 between two species.
Finally, it is interesting to point out that while the two camellia species expressed the same unigenes, their expression level differed at different moisture contents. An explanation to this phenomenon is beyond the scope of the present study as further research related to the regulation of RNA expression during lipid metabolism is expected to shed light on the understanding of fatty acid biosynthesis and accumulation, and simultaneously lay a foundation for its artificial regulation during the after-ripening in oil seed.
Author contribution statement
Conceived and designed the experiment: J-LF, HC and YAK. Performed the experiment: J-LF. Data collection and Figures preparation: J-LF, Z-JY, W-W B, S-P C and W-Q X. Data analysis: J-LF. Wrote the manuscript: J-LF and YAK.
References
Akazawa Y, Isomoto H, Matsushima K, Kanda T, Minami H, Yamaghchi N, Taura N, Shiozawa K, Ohnita K, Takeshima F, Nakano M, Moss J, Hirayama T, Nakao K (2013) Endoplasmic reticulum stress contributes to helicobacter pylori VacA-induced apoptosis. PLoS One. doi:10.1371/journal.pone.0082322
Andre C, Haslam RP, Shanklin J (2012) Feedback regulation of plastidic acetyl-CoA carboxylase by 18:1-acyl carrier protein in Brassica napus. Proc Natl Acad Sci 109:10107–10112
Baud SL, Lepiniec L (2010) Physiological and developmental regulation of seed oil production. Prog Lipid Res 49:235–249
Conesa A, Götz S (2008) Blast2GO: a comprehensive suite for functional analysis in plant genomics. Int J Plant Genom 2008:619832. doi:10.1155/2008/619832
Connell S, Meade KG, Allan B, Lloyd AT, Kenny E, Cormican P, Morris DW, Bradley DG, O’Farrelly C (2012) Avian resistance to campylobacter jejuni colonization is associated with an intestinal immunogene expression signature identified by mRNA sequencing. PLoS One 7:e40409. doi:10.1371/journal.pone.0040409
De Ron AM, Salinero MC, Vela P (2014) Variation in oil content in Camellia japonica seeds. Proce Int Camellia Cong, Pontevedra, Spain 286–291. http://hdl.handle.net/10261/126721
Dušková E, Dušek K, Indrák P, Smékalová K (2016) Postharvest changes in essential oil content and quality of lavender flowers. Ind Crops Prod 79:225–231. doi:10.1016/j.indcrop.2015.11.007
Evaristo AB, Grossi JAS, Pimentel LD, de Melo Goulart S, Martins AD, dos Santos VL, Motoike S (2016) Harvest and post-harvest conditions influencing macauba (Acrocomia aculeata) oil quality attributes. Ind Crops Prod 85:63–73. doi:10.1016/j.indcrop.2016.02.052
Fang X, Fei X, Hong S, Jin Y (2016) Aqueous enzymatic extraction and demulsification of camellia seed oil (Camellia oleifera Abel.) and the oil’s physicochemical properties. Eur J Lipid Sci Technol 118:244–251. doi:10.1002/ejlt.201400582
Fu L, Ding Z, Han B, Hu W, Li Y, Zhang J (2016) Physiological investigation and transcriptome analysis of polyethylene glycol (PEG)-induced dehydration stress in Cassava. Int J MolSci 17:283. doi:10.3390/ijms17030283
Galli V, Guzman F, Messias RS, Körbes AP, Silva SD, Margis-Pinheiro M, Margis R (2014) Transcriptome of tung tree mature seeds with an emphasis on lipid metabolism genes. Tree Genet Genomes 10:1353–1367. doi:10.1007/s11295-014-0765-6
Gao B, Zhang D, Li X, Yang H, Wood AJ (2014) De novo assembly and characterization of the transcriptome in the desiccation-tolerant moss Syntrichia caninervis. BMC Res Notes 7:490–502. doi:10.1186/1756-0500-7-490
Grabherr MG, Haas BJ, Yassour M, Levin JZ, Thompson DA, Amit I, Adiconis X, Fan L, Raychowdhury R, Zeng Q, Chen Z, Mauceli E, Hacohen N, Gnirke A, Rhind N, Palma F, Birren BW, Nusbaum C, Lindblad-Toh K, Friedman N, Regev A (2013) Trinity: reconstructing a full-length transcriptome without a genome from RNA-Seq data. Nat Biotechnol 29:644–652. doi:10.1038/nbt.1883
Haiyan Z, Bedgood DR, Bishop AG, Prenzler PD, Robards K (2007) Food chemistry endogenous biophenol, fatty acid and volatile profiles of selected oils. Food Chem 100:1544–1551
Haro Bailon AD, Obregon S, Rio Celestino MD, Mansilla P, Salinero MC (2014) Variability in seed storage components (protein, oil and fatty acids) in a camellia germplasm collection. Int Camellia Cong. http://hdl.handle.net/10261/126721
Harwood JL, Guschina IA (2013) Regulation of lipid synthesis in oil crops. FEBS Lett 587:2079–2081. doi:10.1016/j.febslet.2013.05.018
He L, Zhou G, Liu J, Xu J (2016) Population genetic analyses of the fungal pathogen colletotrichum fructicola on tea-Oil trees in China. PLoS One 11:e0156841. doi:10.1371/journal.pone.0156841
Hsieh C, Yang J, Chuang Y, Wang E, Lee Y (2013) Effects of roasting prior to pressing on the camellia oil quality. J Taiwan Agri Res 62:249–258. doi:10.6156/JTAR/2013.06203.05
Hyun TK, Kumar D, Cho Y-Y, Hyun H-N, Kim J-S (2013) Computational identification and phylogenetic analysis of the oilbody structural proteins, oleosin and caleosin, in castor bean and flax. Gene 515:454–460. doi:10.1016/j.gene.2012.11.065
Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. Multidiscip J Microb Ecol 99:138–148
Jolivet P, Acevedo F, Boulard C, d’Andréa S, Faure JD, Kohli A, Nesi N, Valot B (2013) Crop seed oil bodies: from challenges in protein identification to an emerging picture of the oil body proteome. Proteomics 13:1836–1849. doi:10.1002/pmic.201200431
Kanehisa M, Araki M, Goto S, Hattori M, Hirakawa M, Itoh M, Katayama T, Kawashima S, Okuda S, Tokimatsu T, Yamanishi Y (2008) KEGG for linking genomes to life and the environment. Nucleic Acids Res 36(suppl 1):480–484. doi:10.1093/nar/gkm882
Lee CP, Yen GC (2006) Antioxidant activity and bioactive compounds of tea seed (Camellia oleifera Abel) oil. J Agri Food Chem 54:779–784
Li H, Fang X, Zhong H, Fei X, Luo F (2014a) Variation of physicochemical properties and nutritional components of oil-tea camellia seeds during ripping. J For Res 27:86–91
Li Y, Lee S, Langleite T, Norheim F, Pourteymour S, Jensen J, Stadheim HK, Storås TH, Davanger S, Gulseth HL, Birkeland KI, Drevon CA, Holen T (2014b) Subsarcolemmal lipid droplet responses to a combined endurance and strength exercise intervention. Physiol Rep 2:e12187. doi:10.14814/phy2.12187
Li S, Wang L, Shu Q, Wu J, Chen L, Shao S, Yin D (2015) Fatty acid composition of developing tree peony (Paeonia section Moutan DC.) seeds and transcriptome analysis during seed development. BMC Genom 16:208. doi:10.1186/s12864-015-1429-0
Liu Y, Huang Z, Ao Y, Li W, Zhang Z (2013) Transcriptome analysis of yellow horn (Xanthoceras sorbifolia Bunge): a potential oil-rich seed tree for biodiesel in China. PLoS One 8:e74441. doi:10.1371/journal.pone.0074441
Liu Z, Xiang H, Hu S, Mou R, Ren J, Zhang S (2014) Drying technology and equipment of oil-tea camellia seeds. China Oils Fats 39:74–77
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25:402–408. doi:10.1006/meth.2001.1262
Lock YY, Snyder CL, Zhu W, Siloto RM, Weselake RJ, Shah S (2009) Antisense suppression of type 1 diacylglycerol acyltransferase adversely affects plant development in Brassica napus. Physiol Plant 137:61–71. doi:10.1111/j.1399-3054.2009.01258.x
Long L, Xue Y, Zhang D, Zhu L, Zhang R (2012) Investigation on main characteristic composition of Camellia oleifera seed oil. China Oils Fats 37:78–81
Mao X, Cai T, Olyarchuk JG, Wei L (2005) Automated genome annotation and pathway identification using the KEGG Orthology (KO) as a controlled vocabulary. Bioinformatics 21:3787–3793. doi:10.1093/bioinformatics/bti430
Mortazavi A, Williams BA, McCue K, Schaeffer L, Wold B (2008) Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5:621–628. doi:10.1038/nmeth.1226
Nakkaew A, Chotigeat W, Eksomtramage T, Phongdara A (2008) Cloning and expression of a plastid-encoded subunit, beta-carboxyltransferase gene (accD) and a nuclear-encoded subunit, biotin carboxylase of acetyl-CoAcarboxylase from oil palm (Elaeis guineensis Jacq.). Plant Sci 175:497–504. doi:10.1016/j.plantsci.2008.05.023
National Development and Reform Commission of China (2009) The national camellia industry development plan (2009–2020). http://www.doc88.com/p-730755963155.html. Accessed 31 July 2017
Nobusawa T, Okushima Y, Nagata N, Kojima M, Sakakibara H, Umeda M (2013) Synthesis of very-long-chain fatty acids in the epidermis controls plant organ growth by restricting cell proliferation. PLoS Biol 11:e1001531. doi:10.1371/journal.pbio.1001531
Pan L, Zhang X, Wang J, Ma X, Zhou M, Huang L, Nie G, Wang P, Yang Z, Li J (2016) Transcriptional profiles of drought-related genes in modulating metabolic processes and antioxidant defenses in Lolium multiflorum. Front Plant Sci 7:519. doi:10.3389/fpls.2016.00519
Parthibane V, Iyappan R, Vijayakumar A, Venkateshwari V, Rajasekharan R (2012) Serine/threonine/tyrosine protein kinase phosphorylates oleosin, a regulator of lipid metabolic functions. Plant Physiol 159:95–104. doi:10.1104/pp.112.197194
Rajwade AV, Kadoo NY, Borikar SP, Harsulkar AM, Ghorpade PB, Gupta VS (2014) Differential transcriptional activity of SAD, FAD2 and FAD3 desaturase genes in developing seeds of linseed contributes to varietal variation in a-linolenic acid content. Phytochem 98:41–53. doi:10.1016/j.phytochem.2013.12.002
Reddy DS, Bhatnagar-Mathur P, Cindhuri KS, Sharma KK (2013) Evaluation and validation of reference genes for normalization of quantitative real-time PCR based gene expression studies in peanut. PLoS One 8:e78555. doi:10.1371/journal.pone.0078555
Renfro DP, Mcintosh BK, Venkatraman A, Siegele DA, Hu JC (2011) GONUTS: the gene ontology normal usage tracking system. Nucleic Acids Res40 Database. doi:10.1093/nar/gkr907
Robards K, Prenzler P, Ryan D, Zhong H (2009) Camellia oil and tea oil. In: Moreau RA, Kamal-Eldin A (eds) Gourmet and health-promoting specialty oils. AOCS Press, Urbana
Rohini G, Rama S, Bijal T, Himabindu K, Lakshmanan K, Nitin M, Rajeev KV, Sabhyata B, Mukesh J (2016) Transcriptome analyses reveal genotype and developmental stage-specific molecular responses to drought and salinity stresses in chickpea. Sci Rep 6:19228. doi:10.1038/srep19228
Shi C-Y, Yang H, Wei C-L, Yu O, Zhang Z-Z, Jiang C-J, Sun J, Li Y-Y, Chen Q, Xia T, Wan X-C (2011) Deep sequencing of the Camellia sinensis transcriptome revealed candidate genes for major metabolic pathways of tea-specific compounds. BMC Genom 12:131. doi:10.1186/1471-2164-12-131
Simpson JP, Ohlrogge JB (2016) A novel pathway for triacylglycerol biosynthesis is responsible for the accumulation of massive quantities of glycerolipids in the surface wax of bayberry (Myrica pensylvanica) fruit. Plant Cell 28:248–264. doi:10.1105/tpc.15.00900
Su MH, Ming CS, Lin K (2014) Chemical composition of seed oils in native Taiwanese Camellia species. Food Chem 156:369–373
Tibaldi G, Emanuela F, Ni Silvana (2010) Cultivation practices do not change the Salvia sclarea L. essential oil but drying process does. J Food Agri Environ 8:790–794
Tibaldi G, Fontana E, Nicola S (2013) Postharvest management affects spearmint and calamint essential oils. J Sci Food Agri 93:580–586. doi:10.1002/jsfa.5836
Wang L, Feng Z, Wang X, Wang X, Zhang X (2009) DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26:136–138. doi:10.1093/bioinformatics/btp612
Wang Y, Sun D, Chen H, Qian L, Xu P (2011) Fatty acid composition and antioxidant activity of tea (Camellia sinensis L.) seed oil extracted by optimized supercritical carbon dioxide. Int J Mol Sci 12:7708–7719. doi:10.3390/ijms12117708
Wang X, Xu R, Wang R, Liu A (2012) Transcriptome analysis of Sacha Inchi (Plukenetia volubilis L.) seeds at two developmental stages. BMC Genom 13:716. doi:10.1186/1471-2164-13-716
Wang L, Yu S, Tong C, Zhao Y, Liu Y, Song C, Zhang Y, Zhang X, Wang Y, Hua W, Li D, Li F, Yu J, Xu C, Han X, Huang S, Tai S, Wang J, Xu X, Li Y, Liu S, Varshney RK, Wang J, Zhang X (2014) Genome sequencing of the high oil crop sesame provides insight into oil biosynthesis. Genome Biol 15:R39. doi:10.1186/gb-2014-15-2-r39
Weselake RJ, Shah S, Tang M, Quant PA, Snyder CL, Furukawa-Stoffer TL, Zhu W, Taylor DC, Zou J, Kumar A, Hall L, Laroche A, Rakow G, Raney P, Moloney MM, Harwood JL (2008) Metabolic control analysis is helpful for informed genetic manipulation of oilseed rape (Brassica napus) to increase seed oil content. J Exp Bot 59:3543–3549. doi:10.1093/jxb/ern206
Xia E, Jiang J, Huang H, Zhang L, Zhang H, Gao L (2014) Transcriptome analysis of the oil-rich tea plant, Camellia oleifera, reveals candidate genes related to lipid metabolism. PLoS One 9:e104150. doi:10.1371/journal.pone.0104150
Xie Y, Zhizhen LI, Yao X, Huang Y, Huang J (2013) Correlation of fruit characters and oil content and fatty acid composition of camellia meiocarpa. China Oils Fats 38:80–83
Xu R, Wang R, Liu A (2011) Expression profiles of genes involved in fatty acid and triacylglycerol synthesis in developing seeds of Jatropha (Jatropha curcas L.). Biomass Bioenergy 35:1683–1692. doi:10.1016/j.biombioe.2011.01.001
Yang C, Liu X, Chen Z, Lin Y, Wang S (2016) Comparison of oil content and fatty acid profile of ten new Camellia oleifera cultivars. J Lipids 2016:3982486. doi:10.1155/2016/3982486
Yao Q, Huang H, Tong Y, Xia E, Gao L (2016) Transcriptome analysis identifies candidate genes related to triacylglycerol and pigment biosynthesis and photoperiodic flowering in the ornamental and oil-producing plant, Camellia reticulata (Theaceae). Front Plant Sci 7:163. doi:10.3389/fpls.2016.00163
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11:R14. doi:10.1186/gb-2010-11-2-r14
Yu M, Liu F, Zhu W, Sun M, Liu J, Li X (2015) New features of triacylglycerol biosynthetic pathways of peanut seeds in early developmental stages. Funct Integr Genom 15:707–716. doi:10.1007/s10142-015-0447-y
Yuan J, Wang C, Chen H, Ye J, Zhou H (2012) Oil content and fatty acid composition analysis of different varieties of Camellia oleifera seeds. China Oils Fats 37:75–79
Zeng Y, Tan X, Zhang L, Jiang N, Cao H (2014) Identification and expression of fructose-1, 6-bisphosphate aldolase genes and their relations to oil content in developing seeds of tea oil tree (Camellia oleifera). PLoS One 9:e107422. doi:10.1371/journal.pone.0107422
Zhang H, Finiguerra M, Dam HG, Huang Y, Xu D, Liu G, Liu G, Lin S (2013) An improved method for achieving high-quality RNA for copepod transcriptomic studies. J Exp Marine Biol Ecol 446:57–66. doi:10.1016/j.jembe.2013.04.021
Zhang D, Jin Q, Wang X, Xue Y, Zhong C, Zhang D (2015) The adulterating research of identifying camellia oil based on fatty acids. J Chin Cereals Oils Assoc 30:124–128
Zhou CF, Lin P, Yao XH, Wang KL, Chang J, Han XJ (2013) Selection of reference genes for quantitative real-time PCR in six oil-tea camellia based on RNA-seq. Mol Biol 47:836–851. doi:10.1134/S0026893313060198
Acknowledgements
We thank Minhou Tongkou State Forest Farm of Fujian Province for serving as our experimental base. We also thank Professor Bruce C Larson of University of British Columbia for help and guidance on the paper.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Funding
This work was supported by grants received under the Doctoral Fund of the Ministry of Education of China (K4112020A), Fujian Province Major Science and Technology Project (2013NZ0207) and The Task 2 of the Key Service Project 5 for the Characteristic Institute of CAS (TSS-2015-014-FW-5-2).
Data archiving statement
Sequence information of all unigenes used in this study is available in the Dryad Digital Repository, doi:10.5061/dryad.2ph6s.
Additional information
Communicated by M. Buckeridge.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Feng, JL., Yang, ZJ., Bai, WW. et al. Transcriptome comparative analysis of two Camellia species reveals lipid metabolism during mature seed natural drying. Trees 31, 1827–1848 (2017). https://doi.org/10.1007/s00468-017-1588-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00468-017-1588-5