Abstract
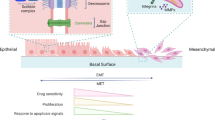
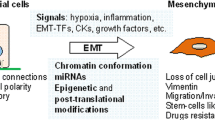
The epithelial-to-mesenchymal transition (EMT) is a reversible change in cell phenotype that plays a crucial role during normal development and cancer metastasis. EMT imparts embryonic epithelial cells with the ability to migrate and to give rise to organs or tissues at distant sites. During cancer progression, the same developmental process is utilized in an analogous manner to enable cancer cells to move to distant organs and form metastases. The reversion of EMT via the mesenchymal-to-epithelial transition (MET) appears to be required for the formation of secondary tumors at distal sites. The plasticity of epigenomic modifications that control the transcriptional program of cells enables cells to switch back and forth from epithelial and mesenchymal phenotypes during these transitions. Here, we review the interplay between complex epigenomic regulatory mechanisms and various transcription factors involved in EMT leading to changes in gene expression and cell phenotype. We also discuss the way that a deeper understanding of the epigenomic regulation of EMT might shed light onto the process of cancer progression and reveal new targets for novel and more specific anticancer epigenomic therapies.



Similar content being viewed by others
References
Agger K, Cloos PAC, Christensen J et al (2007) UTX and JMJD3 are histone H3K27 demethylases involved in HOX gene regulation and development. Nature 449:731–734. doi:10.1038/nature06145
Aghdassi A, Sendler M, Guenther A et al (2012) Recruitment of histone deacetylases HDAC1 and HDAC2 by the transcriptional repressor ZEB1 downregulates E-cadherin expression in pancreatic cancer. Gut 61:439–448. doi:10.1136/gutjnl-2011-300060
Al-Hajj M, Wicha MS, Benito-Hernandez A et al (2003) Prospective identification of tumorigenic breast cancer cells. Proc Natl Acad Sci 100:3983–3988. doi:10.1073/pnas.0530291100
Alsarraj J, Walker RC, Webster JD et al (2011) Deletion of the proline-rich region of the murine metastasis susceptibility gene Brd4 promotes epithelial-to-mesenchymal transition- and stem cell-like conversion. Cancer Res 71:3121–3131. doi:10.1158/0008-5472.CAN-10-4417
Ateeq B, Unterberger A, Szyf M, Rabbani SA (2008) Pharmacological inhibition of DNA methylation induces proinvasive and prometastatic genes in vitro and in vivo. Neoplasia 10:266–278
Bannister AJ, Zegerman P, Partridge JF et al (2001) Selective recognition of methylated lysine 9 on histone H3 by the HP1 chromo domain. Nature 410:120–124. doi:10.1038/35065138
Bannister AJ, Schneider R, Kouzarides T (2002) Histone methylation: dynamic or static? Cell 109:801–806. doi:10.1016/S0092-8674(02)00798-5
Barneda-Zahonero B, Parra M (2012) Histone deacetylases and cancer. Mol Oncol 6:579–589. doi:10.1016/j.molonc.2012.07.003
Batlle E, Sancho E, Francí C et al (2000) The transcription factor Snail is a repressor of E-cadherin gene expression in epithelial tumour cells. Nat Cell Biol 2:84–89. doi:10.1038/35000034
Baylin SB, Jones PA (2011) A decade of exploring the cancer epigenome—biological and translational implications. Nat Rev Cancer 11:726–734. doi:10.1038/nrc3130
Bender CM, Pao MM, Jones PA (1998) Inhibition of DNA methylation by 5-aza-2′-deoxycytidine suppresses the growth of human tumor cell lines. Cancer Res 58:95–101
Berger SL (2007) The complex language of chromatin regulation during transcription. Nature 447:407–412. doi:10.1038/nature05915
Bernstein BE, Mikkelsen TS, Xie X et al (2006) A bivalent chromatin structure marks key developmental genes in embryonic stem cells. Cell 125:315–326. doi:10.1016/j.cell.2006.02.041
Bird A (2002) DNA methylation patterns and epigenetic memory. Genes Dev 16:6–21. doi:10.1101/gad.947102
Bloushtain-Qimron N, Yao J, Snyder EL et al (2008) Cell type-specific DNA methylation patterns in the human breast. Proc Natl Acad Sci U S A 105:14076–14081. doi:10.1073/pnas.0805206105
Bonnet D, Dick JE (1997) Human acute myeloid leukemia is organized as a hierarchy that originates from a primitive hematopoietic cell. Nat Med 3:730–737. doi:10.1038/nm0797-730
Byles V, Zhu L, Lovaas JD et al (2012) SIRT1 induces EMT by cooperating with EMT transcription factors and enhances prostate cancer cell migration and metastasis. Oncogene 31:4619–4629. doi:10.1038/onc.2011.612
Cano A, Pérez-Moreno MA, Rodrigo I et al (2000) The transcription factor Snail controls epithelial–mesenchymal transitions by repressing E-cadherin expression. Nat Cell Biol 2:76–83. doi:10.1038/35000025
Cedar H, Bergman Y (2009) Linking DNA methylation and histone modification: patterns and paradigms. Nat Rev Genet 10:295–304. doi:10.1038/nrg2540
Chang HW, Chow V, Lam KY et al (2002) Loss of E-cadherin expression resulting from promoter hypermethylation in oral tongue carcinoma and its prognostic significance. Cancer 94:386–392. doi:10.1002/cncr.10211
Chen C-L, Liu SS, Ip S-M et al (2003) E-cadherin expression is silenced by DNA methylation in cervical cancer cell lines and tumours. Eur J Cancer 39:517–523. doi:10.1016/S0959-8049(02)00175-2
Chen J, Chan AWH, To K-F et al (2013a) SIRT2 overexpression in hepatocellular carcinoma mediates epithelial to mesenchymal transition by protein kinase B/glycogen synthase kinase-3β/β-catenin signaling. Hepatology 57:2287–2298. doi:10.1002/hep.26278
Chen X, Xiao W, Chen W et al (2013b) The epigenetic modifier trichostatin A, a histone deacetylase inhibitor, suppresses proliferation and epithelial–mesenchymal transition of lens epithelial cells. Cell Death Dis 4:e884. doi:10.1038/cddis.2013.416
Cheung P (2004) Epigenetic regulation by histone methylation and histone variants. Mol Endocrinol 19:563–573. doi:10.1210/me.2004-0496
Craene BD, Berx G (2013) Regulatory networks defining EMT during cancer initiation and progression. Nat Rev Cancer 13:97–110. doi:10.1038/nrc3447
Daigle SR, Olhava EJ, Therkelsen CA et al (2011) Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 20:53–65. doi:10.1016/j.ccr.2011.06.009
Dawson MA, Kouzarides T (2012) Cancer epigenetics: from mechanism to therapy. Cell 150:12–27. doi:10.1016/j.cell.2012.06.013
Dong C, Wu Y, Yao J et al (2012) G9a interacts with Snail and is critical for Snail-mediated E-cadherin repression in human breast cancer. J Clin Invest 122:1469–1486. doi:10.1172/JCI57349
Dong C, Wu Y, Wang Y et al (2013) Interaction with Suv39H1 is critical for Snail-mediated E-cadherin repression in breast cancer. Oncogene 32:1351–1362. doi:10.1038/onc.2012.169
Espada J, Peinado H, Lopez-Serra L et al (2011) Regulation of SNAIL1 and E-cadherin function by DNMT1 in a DNA methylation-independent context. Nucleic Acids Res 39:9194–9205. doi:10.1093/nar/gkr658
Feinberg AP (2007) Phenotypic plasticity and the epigenetics of human disease. Nature 447:433–440. doi:10.1038/nature05919
Feinberg AP, Tycko B (2004) The history of cancer epigenetics. Nat Rev Cancer 4:143–153. doi:10.1038/nrc1279
Fidler IJ (2003) The pathogenesis of cancer metastasis: the “seed and soil” hypothesis revisited. Nat Rev Cancer 3:453–458. doi:10.1038/nrc1098
Filippakopoulos P, Qi J, Picaud S et al (2010) Selective inhibition of BET bromodomains. Nature 468:1067–1073. doi:10.1038/nature09504
Fuks F, Burgers WA, Brehm A et al (2000) DNA methyltransferase Dnmt1 associates with histone deacetylase activity. Nat Genet 24:88–91. doi:10.1038/71750
Glozak MA, Seto E (2007) Histone deacetylases and cancer. Oncogene 26:5420–5432. doi:10.1038/sj.onc.1210610
Gnyszka A, Jastrzebski Z, Flis S (2013) DNA methyltransferase inhibitors and their emerging role in epigenetic therapy of cancer. Anticancer Res 33:2989–2996
Graff JR, Gabrielson E, Fujii H et al (2000) Methylation patterns of the E-cadherin 5′ CpG island are unstable and reflect the dynamic, heterogeneous loss of E-cadherin expression during metastatic progression. J Biol Chem 275:2727–2732
Gupta GP, Massagué J (2006) Cancer metastasis: building a framework. Cell 127:679–695. doi:10.1016/j.cell.2006.11.001
Harris WJ, Huang X, Lynch JT et al (2012) The histone demethylase KDM1A sustains the oncogenic potential of MLL-AF9 leukemia stem cells. Cancer Cell 21:473–487. doi:10.1016/j.ccr.2012.03.014
Herranz N, Pasini D, Díaz VM et al (2008) Polycomb complex 2 is required for E-cadherin repression by the Snail1 transcription factor. Mol Cell Biol 28:4772–4781. doi:10.1128/MCB.00323-08
Hugo H, Ackland ML, Blick T et al (2007) Epithelial—mesenchymal and mesenchymal—epithelial transitions in carcinoma progression. J Cell Physiol 213:374–383. doi:10.1002/jcp.21223
Jacobs SA, Khorasanizadeh S (2002) Structure of HP1 chromodomain bound to a Lysine 9-methylated histone H3 tail. Science 295:2080–2083. doi:10.1126/science.1069473
Jenuwein T, Allis CD (2001) Translating the histone code. Science 293:1074–1080. doi:10.1126/science.1063127
Jin B, Li Y, Robertson KD (2011) DNA methylation: superior or subordinate in the epigenetic hierarchy? Genes Cancer 2:607–617. doi:10.1177/1947601910393957
Jones PA, Baylin SB (2007) The epigenomics of cancer. Cell 128:683–692. doi:10.1016/j.cell.2007.01.029
Kaimori A, Potter JJ, Choti M et al (2010) Histone deacetylase inhibition suppresses the transforming growth factor β1–induced epithelial-to-mesenchymal transition in hepatocytes. Hepatology 52:1033–1045. doi:10.1002/hep.23765
Kang Y, Massagué J (2004) Epithelial-mesenchymal transitions: Twist in development and metastasis. Cell 118:277–279. doi:10.1016/j.cell.2004.07.011
Kirschbaum M, Frankel P, Popplewell L, Zain J, Delioukina M, Pullarkat V, Matsuoka D, Pulone B, Rotter AJ, Espinoza-Delgado I, Nademanee A, Forman SJ, Gandara D, Newman E (2011) Phase II study of vorinostat for treatment of relapsed or refractory indolent non-Hodgkin's lymphoma and mantle cell lymphoma.J Clin Oncol29:1198-1203. doi: 10.1200/JCO.2010.32.1398
Klose RJ, Bird AP (2006) Genomic DNA methylation: the mark and its mediators. Trends Biochem Sci 31:89–97. doi:10.1016/j.tibs.2005.12.008
Klose RJ, Kallin EM, Zhang Y (2006) JmjC-domain-containing proteins and histone demethylation. Nat Rev Genet 7:715–727. doi:10.1038/nrg1945
Ko H, So Y, Jeon H et al (2013) TGF-β1-induced epithelial–mesenchymal transition and acetylation of Smad2 and Smad3 are negatively regulated by EGCG in human A549 lung cancer cells. Cancer Lett 335:205–213. doi:10.1016/j.canlet.2013.02.018
Koizume S, Tachibana K, Sekiya T et al (2002) Heterogeneity in the modification and involvement of chromatin components of the CpG island of the silenced human CDH1 gene in cancer cells. Nucleic Acids Res 30:4770–4780
Kong D, Ahmad A, Bao B et al (2012) Histone deacetylase inhibitors induce epithelial-to-mesenchymal transition in prostate cancer cells. PLoS One 7:e45045. doi:10.1371/journal.pone.0045045
Kouzarides T (1999) Histone acetylases and deacetylases in cell proliferation. Curr Opin Genet Dev 9:40–48
Kouzarides T (2002) Histone methylation in transcriptional control. Curr Opin Genet Dev 12:198–209. doi:10.1016/S0959-437X(02)00287-3
Kouzarides T (2007) Chromatin modifications and their function. Cell 128:693–705. doi:10.1016/j.cell.2007.02.005
Kruidenier L, Chung C, Cheng Z et al (2012) A selective jumonji H3K27 demethylase inhibitor modulates the proinflammatory macrophage response. Nature 488:404–408. doi:10.1038/nature11262
Kummar S, Gutierrez M, Gardner ER et al (2007) Phase I trial of MS-275, a histone deacetylase inhibitor, administered weekly in refractory solid tumors and lymphoid malignancies. Clin Cancer Res 13:5411–5417. doi:10.1158/1078-0432.CCR-07-0791
Kuo MH, Allis CD (1998) Roles of histone acetyltransferases and deacetylases in gene regulation. BioEssays 20:615–626. doi:10.1002/(SICI)1521-1878(199808)20:8<615::AID-BIES4>3.0.CO;2-H
Lachner M, O’Carroll D, Rea S et al (2001) Methylation of histone H3 lysine 9 creates a binding site for HP1 proteins. Nature 410:116–120. doi:10.1038/35065132
Lei W, Zhang K, Pan X et al (2010) Histone deacetylase 1 is required for transforming growth factor-β1-induced epithelial–mesenchymal transition. Int J Biochem Cell Biol 42:1489–1497. doi:10.1016/j.biocel.2010.05.006
Li E (2002) Chromatin modification and epigenetic reprogramming in mammalian development. Nat Rev Genet 3:662–673. doi:10.1038/nrg887
Lim S, Janzer A, Becker A et al (2010) Lysine-specific demethylase 1 (LSD1) is highly expressed in ER-negative breast cancers and a biomarker predicting aggressive biology. Carcinogenesis 31:512–520. doi:10.1093/carcin/bgp324
Lin T, Ponn A, Hu X et al (2010) Requirement of the histone demethylase LSD1 in Snai1-mediated transcriptional repression during epithelial-mesenchymal transition. Oncogene 29:4896–4904. doi:10.1038/onc.2010.234
Lombaerts M, Wezel T van, Philippo K et al (2006) E-cadherin transcriptional downregulation by promoter methylation but not mutation is related to epithelial-to-mesenchymal transition in breast cancer cell lines. Br J Cancer 94:661–671. doi:10.1038/sj.bjc.6602996
Lyko F, Brown R (2005) DNA methyltransferase inhibitors and the development of epigenetic cancer therapies. J Natl Cancer Inst 97:1498–1506. doi:10.1093/jnci/dji311
Mani SA, Guo W, Liao M-J et al (2008) The epithelial-mesenchymal transition generates cells with properties of stem cells. Cell 133:704–715. doi:10.1016/j.cell.2008.03.027
Mansour AA, Gafni O, Weinberger L et al (2012) The H3K27 demethylase Utx regulates somatic and germ cell epigenetic reprogramming. Nature 488:409–413. doi:10.1038/nature11272
Marks PA, Rifkind RA, Richon VM et al (2001) Histone deacetylases and cancer: causes and therapies. Nat Rev Cancer 1:194–202. doi:10.1038/35106079
Marmorstein R, Roth SY (2001) Histone acetyltransferases: function, structure, and catalysis. Curr Opin Genet Dev 11:155–161
Martin A, Cano A (2010) Tumorigenesis: Twist1 links EMT to self-renewal. Nat Cell Biol 12:924–925. doi:10.1038/ncb1010-924
McCabe MT, Brandes JC, Vertino PM (2009) Cancer DNA methylation: molecular mechanisms and clinical implications. Clin Cancer Res 15:3927–3937. doi:10.1158/1078-0432.CCR-08-2784
McCabe MT, Ott HM, Ganji G et al (2012) EZH2 inhibition as a therapeutic strategy for lymphoma with EZH2-activating mutations. Nature 492:108–112. doi:10.1038/nature11606
McDonald OG, Wu H, Timp W et al (2011) Genome-scale epigenetic reprogramming during epithelial-to-mesenchymal transition. Nat Struct Mol Biol 18:867–874. doi:10.1038/nsmb.2084
Mertz JA, Conery AR, Bryant BM et al (2011) Targeting MYC dependence in cancer by inhibiting BET bromodomains. Proc Natl Acad Sci U S A 108:16669–16674. doi:10.1073/pnas.1108190108
Metzger E, Wissmann M, Yin N et al (2005) LSD1 demethylates repressive histone marks to promote androgen-receptor-dependent transcription. Nature 437:436–439. doi:10.1038/nature04020
Moreno-Bueno G, Portillo F, Cano A (2008) Transcriptional regulation of cell polarity in EMT and cancer. Oncogene 27:6958–6969. doi:10.1038/onc.2008.346
Nieto MA (2013) Epithelial plasticity: a common theme in embryonic and cancer cells. Science 342:1234850. doi:10.1126/science.1234850
O’Connor OA, Heaney ML, Schwartz L et al (2006) Clinical experience with intravenous and oral formulations of the novel histone deacetylase inhibitor suberoylanilide hydroxamic acid in patients with advanced hematologic malignancies. J Clin Oncol 24:166–173. doi:10.1200/JCO.2005.01.9679
Patel DJ, Wang Z (2013) Readout of epigenetic modifications. Annu Rev Biochem 82:81–118. doi:10.1146/annurev-biochem-072711-165700
Peinado H, Portillo F, Cano A (2004) Transcriptional regulation of cadherins during development and carcinogenesis. Int J Dev Biol 48:365–375. doi:10.1387/ijdb.041794hp
Peinado H, Olmeda D, Cano A (2007) Snail, Zeb and bHLH factors in tumour progression: an alliance against the epithelial phenotype? Nat Rev Cancer 7:415–428. doi:10.1038/nrc2131
Pérez-Moreno MA, Locascio A, Rodrigo I et al (2001) A new role for E12/E47 in the repression of E-cadherin expression and epithelial-mesenchymal transitions. J Biol Chem 276:27424–27431. doi:10.1074/jbc.M100827200
Polyak K, Weinberg RA (2009) Transitions between epithelial and mesenchymal states: acquisition of malignant and stem cell traits. Nat Rev Cancer 9:265–273. doi:10.1038/nrc2620
Ramadoss S, Chen X, Wang C-Y (2012) Histone demethylase KDM6B promotes epithelial-mesenchymal transition. J Biol Chem 287:44508–44517. doi:10.1074/jbc.M112.424903
Rice JC, Allis CD (2001) Histone methylation versus histone acetylation: new insights into epigenetic regulation. Curr Opin Cell Biol 13:263–273. doi:10.1016/S0955-0674(00)00208-8
Richly H, Aloia L, Di Croce L (2011) Roles of the Polycomb group proteins in stem cells and cancer. Cell Death Dis 2:e204. doi:10.1038/cddis.2011.84
Robertson KD (2002) DNA methylation and chromatin—unraveling the tangled web. Oncogene 21:5361–5379. doi:10.1038/sj.onc.1205609
Robertson KD (2005) DNA methylation and human disease. Nat Rev Genet 6:597–610. doi:10.1038/nrg1655
Saijo K, Katoh T, Shimodaira H et al (2012) Romidepsin (FK228) and its analogs directly inhibit phosphatidylinositol 3-kinase activity and potently induce apoptosis as histone deacetylase/phosphatidylinositol 3-kinase dual inhibitors. Cancer Sci 103:1994–2001. doi:10.1111/cas.12002
Scheel C, Weinberg RA (2012) Cancer stem cells and epithelial-mesenchymal transition: concepts and molecular links. Semin Cancer Biol 22:396–403. doi:10.1016/j.semcancer.2012.04.001
Schenk T, Chen WC, Göllner S et al (2012) Inhibition of the LSD1 (KDM1A) demethylase reactivates the all-trans-retinoic acid differentiation pathway in acute myeloid leukemia. Nat Med 18:605–611. doi:10.1038/nm.2661
Schildhaus H-U, Riegel R, Hartmann W et al (2011) Lysine-specific demethylase 1 is highly expressed in solitary fibrous tumors, synovial sarcomas, rhabdomyosarcomas, desmoplastic small round cell tumors, and malignant peripheral nerve sheath tumors. Hum Pathol 42:1667–1675. doi:10.1016/j.humpath.2010.12.025
Schulte JH, Lim S, Schramm A et al (2009) Lysine-specific demethylase 1 is strongly expressed in poorly differentiated neuroblastoma: implications for therapy. Cancer Res 69:2065–2071. doi:10.1158/0008-5472.CAN-08-1735
Senisterra G, Wu H, Allali-Hassani A et al (2013) Small-molecule inhibition of MLL activity by disruption of its interaction with WDR5. Biochem J 449:151–159. doi:10.1042/BJ20121280
Serce N, Gnatzy A, Steiner S et al (2012) Elevated expression of LSD1 (lysine-specific demethylase 1) during tumour progression from pre-invasive to invasive ductal carcinoma of the breast. BMC Clin Pathol 12:13. doi:10.1186/1472-6890-12-13
Seward DJ, Cubberley G, Kim S et al (2007) Demethylation of trimethylated histone H3 Lys4 in vivo by JARID1 JmjC proteins. Nat Struct Mol Biol 14:240–242. doi:10.1038/nsmb1200
Sharma S, Kelly TK, Jones PA (2010) Epigenetics in cancer. Carcinogenesis 31:27–36. doi:10.1093/carcin/bgp220
Shi Y (2007) Histone lysine demethylases: emerging roles in development, physiology and disease. Nat Rev Genet 8:829–833. doi:10.1038/nrg2218
Shi Y, Whetstine JR (2007) Dynamic regulation of histone lysine methylation by demethylases. Mol Cell 25:1–14. doi:10.1016/j.molcel.2006.12.010
Shi Y, Lan F, Matson C et al (2004) Histone demethylation mediated by the nuclear amine oxidase homolog LSD1. Cell 119:941–953. doi:10.1016/j.cell.2004.12.012
Shogren-Knaak M, Ishii H, Sun J-M et al (2006) Histone H4-K16 acetylation controls chromatin structure and protein interactions. Science 311:844–847. doi:10.1126/science.1124000
Sparmann A, Lohuizen M van (2006) Polycomb silencers control cell fate, development and cancer. Nat Rev Cancer 6:846–856. doi:10.1038/nrc1991
Sterner DE, Berger SL (2000) Acetylation of histones and transcription-related factors. Microbiol Mol Biol Rev 64:435–459. doi:10.1128/MMBR.64.2.435-459.2000
Strahl BD, Allis CD (2000) The language of covalent histone modifications. Nature 403:41–45. doi:10.1038/47412
Struhl K (1998) Histone acetylation and transcriptional regulatory mechanisms. Genes Dev 12:599–606
Tam WL, Weinberg RA (2013) The epigenetics of epithelial-mesenchymal plasticity in cancer. Nat Med 19:1438–1449. doi:10.1038/nm.3336
Thieme S, Gyárfás T, Richter C et al (2013) The histone demethylase UTX regulates stem cell migration and hematopoiesis. Blood 121:2462–2473. doi:10.1182/blood-2012-08-452003
Thiery JP (2002) Epithelial–mesenchymal transitions in tumour progression. Nat Rev Cancer 2:442–454. doi:10.1038/nrc822
Thiery JP, Sleeman JP (2006) Complex networks orchestrate epithelial–mesenchymal transitions. Nat Rev Mol Cell Biol 7:131–142. doi:10.1038/nrm1835
Thiery JP, Acloque H, Huang RYJ, Nieto MA (2009) Epithelial-mesenchymal transitions in development and disease. Cell 139:871–890. doi:10.1016/j.cell.2009.11.007
Tiwari N, Gheldof A, Tatari M, Christofori G (2012) EMT as the ultimate survival mechanism of cancer cells. Semin Cancer Biol 22:194–207. doi:10.1016/j.semcancer.2012.02.013
Valastyan S, Weinberg RA (2011) Tumor metastasis: molecular insights and evolving paradigms. Cell 147:275–292. doi:10.1016/j.cell.2011.09.024
Vesuna F, Diest P van, Chen JH, Raman V (2008) Twist is a transcriptional repressor of E-cadherin gene expression in breast cancer. Biochem Biophys Res Commun 367:235–241. doi:10.1016/j.bbrc.2007.11.151
Viré E, Brenner C, Deplus R et al (2006) The Polycomb group protein EZH2 directly controls DNA methylation. Nature 439:871–874. doi:10.1038/nature04431
Wang Y, Shang Y (2013) Epigenetic control of epithelial-to-mesenchymal transition and cancer metastasis. Exp Cell Res 319:160–169. doi:10.1016/j.yexcr.2012.07.019
Wang Y, Zhang H, Chen Y et al (2009) LSD1 Is a subunit of the NuRD complex and targets the metastasis programs in breast cancer. Cell 138:660–672. doi:10.1016/j.cell.2009.05.050
Watanabe T, Kato H, Kobayashi Y, Yamasaki S, Morita-Hoshi Y, Yokoyama H, Morishima Y, Ricker JL, Otsuki T, Miyagi-Maesima A, Matsuno Y, Tobinai K (2010) Potential efficacy of the oral histone deacetylase inhibitor vorinostat in a phase I trial in follicular and mantle cell lymphoma. Cancer Sci 101:196-200. doi: 10.1111/j.1349-7006.2009.01360.x
Witta SE, Gemmill RM, Hirsch FR et al (2006) Restoring E-Cadherin expression increases sensitivity to epidermal growth factor receptor inhibitors in lung cancer cell lines. Cancer Res 66:944–950. doi:10.1158/0008-5472.CAN-05-1988
Wolffe AP, Matzke MA (1999) Epigenetics: regulation through repression. Science 286:481–486. doi:10.1126/science.286.5439.481
Wu M-Z, Tsai Y-P, Yang M-H et al (2011) Interplay between HDAC3 and WDR5 is essential for hypoxia-induced epithelial-mesenchymal transition. Mol Cell 43:811–822. doi:10.1016/j.molcel.2011.07.012
Wysocka J, Swigut T, Milne TA et al (2005) WDR5 associates with histone H3 methylated at K4 and is essential for H3 K4 methylation and vertebrate development. Cell 121:859–872. doi:10.1016/j.cell.2005.03.036
Yang F, Sun L, Li Q et al (2012) SET8 promotes epithelial–mesenchymal transition and confers TWIST dual transcriptional activities. EMBO J 31:110–123. doi:10.1038/emboj.2011.364
Yang M-H, Hsu DS-S, Wang H-W et al (2010) Bmi1 is essential in Twist1-induced epithelial-mesenchymal transition. Nat Cell Biol 12:982–992. doi:10.1038/ncb2099
Yoshikawa M, Hishikawa K, Marumo T, Fujita T (2007) Inhibition of histone deacetylase activity suppresses epithelial-to-mesenchymal transition induced by TGF-β1 in human renal epithelial cells. J Am Soc Nephrol 18:58–65. doi:10.1681/ASN.2005111187
Yoshiura K, Kanai Y, Ochiai A et al (1995) Silencing of the E-cadherin invasion-suppressor gene by CpG methylation in human carcinomas. Proc Natl Acad Sci U S A 92:7416–7419
You JS, Jones PA (2012) Cancer genetics and epigenetics: two sides of the same coin? Cancer Cell 22:9–20. doi:10.1016/j.ccr.2012.06.008
Yuan Y, Wang Q, Paulk J et al (2012) A small-molecule probe of the histone methyltransferase G9a induces cellular senescence in pancreatic adenocarcinoma. ACS Chem Biol 7:1152–1157. doi:10.1021/cb300139y
Zhang Y, Reinberg D (2001) Transcription regulation by histone methylation: interplay between different covalent modifications of the core histone tails. Genes Dev 15:2343–2360. doi:10.1101/gad.927301
Zhao L, Li W, Zang W et al (2013) JMJD2B promotes epithelial–mesenchymal transition by cooperating with β-catenin and enhances gastric cancer metastasis. Clin Cancer Res 19:6419–6429. doi:10.1158/1078-0432.CCR-13-0254
Zuber J, Shi J, Wang E et al (2011) RNAi screen identifies Brd4 as a therapeutic target in acute myeloid leukaemia. Nature 478:524–528. doi:10.1038/nature10334
Acknowledgments
Because of the large body of work surrounding the topics of epigenomic regulation, tumor metastasis and EMT, the authors were unable to cite all the relevant work related to these topics. Thus, the authors wish to apologize and acknowledge the work of many other scientists who are diligently working in these fields but whose work was not cited here. The authors further thank U. Bedi, F. Wegwitz, M. Dobbelstein, U. Keitel and C. Scheel for insightful conversations related to the topic of this review.
Author information
Authors and Affiliations
Corresponding author
Additional information
Work in the Johnsen group is supported by grants from the Deutsche Krebshilfe (109088) and the Deutsche Forschungsgemeinschaft (JO 815/3) to S.A.J.
Rights and permissions
About this article
Cite this article
Mishra, V.K., Johnsen, S.A. Targeted therapy of epigenomic regulatory mechanisms controlling the epithelial to mesenchymal transition during tumor progression. Cell Tissue Res 356, 617–630 (2014). https://doi.org/10.1007/s00441-014-1912-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00441-014-1912-y