Abstract
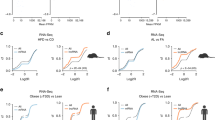
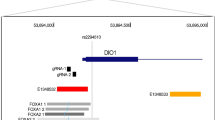
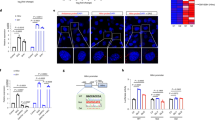
Long non-coding RNA (lncRNA) genes represent a large class of transcripts that are widely expressed across species. As most human lncRNAs are non-conserved, we recently employed a unique humanized liver mouse model to study lncRNAs expressed in human livers. We identified a human hepatocyte-specific lncRNA, hLMR1 (human lncRNA metabolic regulator 1), which is induced by feeding and promotes hepatic cholesterol synthesis. Recent genome-wide association studies (GWAS) found that several single-nucleotide polymorphisms (SNPs) from the hLMR1 gene locus are associated with blood lipids and markers of liver damage. These results suggest that dietary and genetic factors may regulate hLMR1 to affect disease progression. In this study, we first screened for nutritional/hormonal factors and found that hLMR1 was robustly induced by insulin/glucose in cultured human hepatocytes, and this induction is dependent on the transcription factor SREBP1. We then tested if GWAS SNPs genetically linked to hLMR1 could regulate hLMR1 expression. We found that DNA sequences flanking rs9653945, a SNP from the last exon of the hLMR1 gene, functions as an enhancer that can be robustly activated by SREBP1c depending on the presence of rs9653945 major allele (G). We further performed CRISPR base editing in human HepG2 cells and found that rs9653945 major (G) to minor (A) allele modification resulted in blunted insulin/glucose-induced expression of hLMR1. Finally, we performed genotyping and gene expression analyses using a published human NAFLD RNA-seq dataset and found that individuals homozygous for rs9653945-G have a higher expression of hLMR1 and risk of NAFLD. Taken together, our data support a model that rs9653945-G predisposes individuals to insulin/glucose-induced hLMR1, contributing to the development of hyperlipidemia and NAFLD.




Similar content being viewed by others
Data availability
All data that support the findings of this study are available from the corresponding author upon reasonable request.
References
Ahituv N (2016) Exonic enhancers: proceed with caution in exome and genome sequencing studies. Genome Med 8:14. https://doi.org/10.1186/s13073-016-0277-0
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Consortium EP (2012) An integrated encyclopedia of DNA elements in the human genome. Nature 489:57–74. https://doi.org/10.1038/nature11247
Consortium GT (2013) The Genotype-Tissue Expression (GTEx) project. Nat Genet 45:580–585. https://doi.org/10.1038/ng.2653
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, Batut P, Chaisson M, Gingeras TR (2013) STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29:15–21. https://doi.org/10.1093/bioinformatics/bts635
Graham SE, Clarke SL, Wu KH, Kanoni S, Zajac GJM, Ramdas S, Surakka I, Ntalla I, Vedantam S, Winkler TW et al (2021) The power of genetic diversity in genome-wide association studies of lipids. Nature 600:675–679. https://doi.org/10.1038/s41586-021-04064-3
Hasegawa M, Kawai K, Mitsui T, Taniguchi K, Monnai M, Wakui M, Ito M, Suematsu M, Peltz G, Nakamura M, Suemizu H (2011) The reconstituted “humanized liver” in TK-NOG mice is mature and functional. Biochem Biophys Res Commun 405:405–410. https://doi.org/10.1016/j.bbrc.2011.01.042
Horn CL, Morales AL, Savard C, Farrell GC, Ioannou GN (2022) Role of cholesterol-associated steatohepatitis in the development of NASH. Hepatol Commun 6:12–35. https://doi.org/10.1002/hep4.1801
Huang TP, Newby GA, Liu DR (2021) Precision genome editing using cytosine and adenine base editors in mammalian cells. Nat Protoc 16:1089–1128. https://doi.org/10.1038/s41596-020-00450-9
Johnsson P, Lipovich L, Grander D, Morris KV (2014) Evolutionary conservation of long non-coding RNAs; sequence, structure, function. Biochim Biophys Acta 1840:1063–1071. https://doi.org/10.1016/j.bbagen.2013.10.035
Klarin D, Damrauer SM, Cho K, Sun YV, Teslovich TM, Honerlaw J, Gagnon DR, DuVall SL, Li J, Peloso GM et al (2018) Genetics of blood lipids among ~300,000 multi-ethnic participants of the Million Veteran Program. Nat Genet 50:1514–1523. https://doi.org/10.1038/s41588-018-0222-9
Li H (2011) A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics 27:2987–2993. https://doi.org/10.1093/bioinformatics/btr509
Li P, Ruan X, Yang L, Kiesewetter K, Zhao Y, Luo H, Chen Y, Gucek M, Zhu J, Cao H (2015) A liver-enriched long non-coding RNA, lncLSTR, regulates systemic lipid metabolism in mice. Cell Metab 21:455–467. https://doi.org/10.1016/j.cmet.2015.02.004
Liao Y, Smyth GK, Shi W (2013) The Subread aligner: fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Res 41:e108. https://doi.org/10.1093/nar/gkt214
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
Mardinoglu A, Wu H, Bjornson E, Zhang C, Hakkarainen A, Rasanen SM, Lee S, Mancina RM, Bergentall M, Pietilainen KH et al (2018) An integrated understanding of the rapid metabolic benefits of a carbohydrate-restricted diet on hepatic steatosis in humans. Cell Metab 27(559–571):e555. https://doi.org/10.1016/j.cmet.2018.01.005
Orozco G, Schoenfelder S, Walker N, Eyre S, Fraser P (2022) 3D genome organization links non-coding disease-associated variants to genes. Front Cell Dev Biol 10:995388. https://doi.org/10.3389/fcell.2022.995388
Pasquali L, Gaulton KJ, Rodriguez-Segui SA, Mularoni L, Miguel-Escalada I, Akerman I, Tena JJ, Moran I, Gomez-Marin C, van de Bunt M et al (2014) Pancreatic islet enhancer clusters enriched in type 2 diabetes risk-associated variants. Nat Genet 46:136–143. https://doi.org/10.1038/ng.2870
Qi G, Chatterjee N (2018) Heritability informed power optimization (HIPO) leads to enhanced detection of genetic associations across multiple traits. PLoS Genet 14:e1007549. https://doi.org/10.1371/journal.pgen.1007549
Reed BD, Charos AE, Szekely AM, Weissman SM, Snyder M (2008) Genome-wide occupancy of SREBP1 and its partners NFY and SP1 reveals novel functional roles and combinatorial regulation of distinct classes of genes. PLoS Genet 4:e1000133. https://doi.org/10.1371/journal.pgen.1000133
Roadmap Epigenomics C, Kundaje A, Meuleman W, Ernst J, Bilenky M, Yen A, Heravi-Moussavi A, Kheradpour P, Zhang Z, Wang J et al (2015) Integrative analysis of 111 reference human epigenomes. Nature 518:317–330. https://doi.org/10.1038/nature14248
Ruan X, Li P, Cangelosi A, Yang L, Cao H (2016) A long non-coding RNA, lncLGR, regulates hepatic glucokinase expression and glycogen storage during fasting. Cell Rep 14:1867–1875. https://doi.org/10.1016/j.celrep.2016.01.062
Ruan X, Li P, Chen Y, Shi Y, Pirooznia M, Seifuddin F, Suemizu H, Ohnishi Y, Yoneda N, Nishiwaki M et al (2020) In vivo functional analysis of non-conserved human lncRNAs associated with cardiometabolic traits. Nat Commun 11:45. https://doi.org/10.1038/s41467-019-13688-z
Ruan X, Li P, Ma Y, Jiang CF, Chen Y, Shi Y, Gupta N, Seifuddin F, Pirooznia M, Ohnishi Y et al (2021) Identification of human long noncoding RNAs associated with nonalcoholic fatty liver disease and metabolic homeostasis. J Clin Invest. https://doi.org/10.1172/JCI136336
Sakaue S, Kanai M, Tanigawa Y, Karjalainen J, Kurki M, Koshiba S, Narita A, Konuma T, Yamamoto K, Akiyama M et al (2021) A cross-population atlas of genetic associations for 220 human phenotypes. Nat Genet 53:1415–1424. https://doi.org/10.1038/s41588-021-00931-x
Sallam T, Jones M, Thomas BJ, Wu X, Gilliland T, Qian K, Eskin A, Casero D, Zhang Z, Sandhu J et al (2018) Transcriptional regulation of macrophage cholesterol efflux and atherogenesis by a long noncoding RNA. Nat Med 24:304–312. https://doi.org/10.1038/nm.4479
Sollis E, Mosaku A, Abid A, Buniello A, Cerezo M, Gil L, Groza T, Gunes O, Hall P, Hayhurst J et al (2023) The NHGRI-EBI GWAS Catalog: knowledgebase and deposition resource. Nucleic Acids Res 51:D977–D985. https://doi.org/10.1093/nar/gkac1010
Suppli MP, Rigbolt KTG, Veidal SS, Heeboll S, Eriksen PL, Demant M, Bagger JI, Nielsen JC, Oro D, Thrane SW et al (2019) Hepatic transcriptome signatures in patients with varying degrees of nonalcoholic fatty liver disease compared with healthy normal-weight individuals. Am J Physiol Gastrointest Liver Physiol 316:G462–G472. https://doi.org/10.1152/ajpgi.00358.2018
Thuronyi BW, Koblan LW, Levy JM, Yeh WH, Zheng C, Newby GA, Wilson C, Bhaumik M, Shubina-Oleinik O, Holt JR, Liu DR (2019) Continuous evolution of base editors with expanded target compatibility and improved activity. Nat Biotechnol 37:1070–1079. https://doi.org/10.1038/s41587-019-0193-0
Toth PP, Potter D, Ming EE (2012) Prevalence of lipid abnormalities in the United States: the National Health and Nutrition Examination Survey 2003–2006. J Clin Lipidol 6:325–330. https://doi.org/10.1016/j.jacl.2012.05.002
Vosa U, Claringbould A, Westra HJ, Bonder MJ, Deelen P, Zeng B, Kirsten H, Saha A, Kreuzhuber R, Yazar S et al (2021) Large-scale cis- and trans-eQTL analyses identify thousands of genetic loci and polygenic scores that regulate blood gene expression. Nat Genet 53:1300–1310. https://doi.org/10.1038/s41588-021-00913-z
Wang Y, Viscarra J, Kim SJ, Sul HS (2015) Transcriptional regulation of hepatic lipogenesis. Nat Rev Mol Cell Biol 16:678–689. https://doi.org/10.1038/nrm4074
Yang L, Li P, Yang W, Ruan X, Kiesewetter K, Zhu J, Cao H (2016) Integrative transcriptome analyses of metabolic responses in mice define pivotal LncRNA metabolic regulators. Cell Metab 24:627–639. https://doi.org/10.1016/j.cmet.2016.08.019
Zhao XY, Li S, Wang GX, Yu Q, Lin JD (2014) A long noncoding RNA transcriptional regulatory circuit drives thermogenic adipocyte differentiation. Mol Cell 55:372–382. https://doi.org/10.1016/j.molcel.2014.06.004
Acknowledgements
We thank Dr. Timothy Osborne for providing the active SREBP1c plasmid. We are grateful to Dr. Timothy Osborne for critical reading of the manuscript and helpful comments. This study was funded by Johns Hopkins University institutional funds for Dr. Xiangbo Ruan.
Funding
This work was supported by Johns Hopkins University institutional funds for Dr. Xiangbo Ruan.
Author information
Authors and Affiliations
Contributions
M.J. and I.P. performed the experiments and analyzed the data. S.T. performed the bioinformatics analyses. M.J., S.T. and X.R. wrote the manuscript. X.R. conceived and supervised the study.
Corresponding author
Ethics declarations
Conflict of Interests
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Jaso-Vera, M.E., Takaoka, S., Patel, I. et al. Integrative regulation of hLMR1 by dietary and genetic factors in nonalcoholic fatty liver disease and hyperlipidemia. Hum. Genet. (2024). https://doi.org/10.1007/s00439-024-02654-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00439-024-02654-5