Abstract
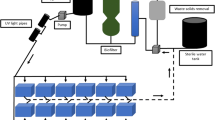
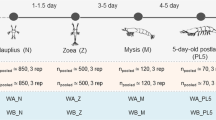
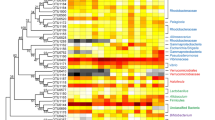
Increasing studies have revealed strong links among gut microbiota, health status, and shrimp development, but they mainly focus on the microbiota of Pacific white shrimp, Penaeus vannamei, during life stages from juveniles to adults. Little is known about shrimp microbiota dynamics at early developmental stages. In this study, with an aim to profile shrimp microbiota and its dynamics at stages nauplius, zoea, mysis, and early postlarva, we conducted a survey for the successful breeding processes in a commercial hatchery in China, sampled 33 samples including larval/postlarval shrimp, suspended substance in rearing water (SSRW), and nutrition supplements (i.e., algae and brine shrimp larvae) at stages N5, Z2, M2, and P2. The associated bacterial communities were sequenced and comparatively analyzed using high-throughput sequencing of bacterial 16S rRNA genes. Our case study results showed that bacterial community structures and compositions were strikingly different at stages N5, Z2, and P2, indicating the shift of microbiota at the three stages. Many taxa within Gamma-, Alphaproteobacteria, and Flavobacteriia classes were observed to be stage-specifically abundant and identified as taxonomic biomarkers potentially used to differentiate among shrimp at different early developmental stages. Summing up, these results shed light on larval/postlarval microbiota and its dynamics at different early developmental stages, highlighting the potential roles of shrimp development in microbiota formation and shifting.





Similar content being viewed by others
References
Avendano-Herrera R, Toranzo AE, Magarinos B (2006) Tenacibaculosis infection in marine fish caused by Tenacibaculum maritimum: a review. Dis Aquat Org 71:255–266. https://doi.org/10.3354/dao071255
Bjornsdottir R, Karadottir EG, Johannsdottir J, Thorarinsdottir EE, Smaradottir H, Sigurgisladottir S, Gudmundsdottir BK (2010) Selection of bacteria and the effects of bacterial treatment of Atlantic halibut (Hippoglossus hippoglossus L.) eggs and larvae. Aquaculture 302:219–227. https://doi.org/10.1016/j.aquaculture.2010.02.026
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59. https://doi.org/10.1038/nmeth.2276
Burns AR, Stephens WZ, Stagaman K, Wong S, Rawls JF, Guillemin K, Bohannan BJ (2016) Contribution of neutral processes to the assembly of gut microbial communities in the zebrafish over host development. ISME J 10:655–664. https://doi.org/10.1038/ismej.2015.142
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci USA 108:4516–4522. https://doi.org/10.1073/pnas.1000080107
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2014) Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642. https://doi.org/10.1093/nar/gkt1244
Doyle JJ, Doyle JL (1990) Isolation of plant DNA from fresh tissue. Focus 12:11–15
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. https://doi.org/10.1038/nmeth.2604
FAO (2018) FAO Yearbook: Fishery and Aquaculture Statistics 2016. Food and Agriculture Organization of the United Nations, Rome, 104pp. Retrieved from www.fao.org/fishery/static/Yearbook/YB2016_USBcard/index.htm
Greer CW (2010) Bacterial diversity in hydrocarbon-polluted rivers, estuaries and sediments. Springer, Heidelberg
Hou D, Huang Z, Zeng S, Liu J, Wei D, Deng X, Weng S, Yan Q, He J (2018) Intestinal bacterial signatures of white feces syndrome in shrimp. Appl Microbiol Biotechnol 102:3701–3709. https://doi.org/10.1007/s00253-018-8855-2
Huang Z, Li X, Wang L, Shao Z (2016) Changes in the intestinal bacterial community during the growth of white shrimp, Litopenaeus vannamei. Aquac Res 47:1737–1746. https://doi.org/10.1111/are.12628
Jaroenlak P, Sanguanrut P, Williams BA, Stentiford GD, Flegel TW, Sritunyalucksana K, Itsathitphaisarn O (2016) A nested PCR assay to avoid false positive detection of the microsporidian Enterocytozoon hepatopenaei (EHP) in environmental samples in shrimp farms. PLoS One 11:e0166320. https://doi.org/10.1371/journal.pone.0166320
Liu H, Wang L, Liu M, Wang B, Jiang K, Ma S, Li Q (2011) The intestinal microbial diversity in Chinese shrimp (Fenneropenaeus chinensis) as determined by PCR–DGGE and clone library analyses. Aquaculture 317:32–36. https://doi.org/10.1016/j.aquaculture.2011.04.008
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Moriarty DJW (1997) The role of microorganisms in aquaculture ponds. Aquaculture 151:333–349. https://doi.org/10.1016/s0044-8486(96)01487-1
OIE (2017) Manual of diagnostic tests for aquatic animals. World Organisation for Animal Health, Paris. Retrieved from https://www.oie.int/international-standard-setting/aquatic-code/access-online/
Qiu L, Chen MM, Wan XY, Li C, Zhang QL, Wang RY, Cheng DY, Dong X, Yang B, Wang XH, Xiang JH, Huang J (2017) Characterization of a new member of Iridoviridae, Shrimp hemocyte iridescent virus (SHIV), found in white leg shrimp ( Litopenaeus vannamei). Sci Rep 7:11834. https://doi.org/10.1038/s41598-017-10738-8
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glockner FO (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:D590–D596. https://doi.org/10.1093/nar/gks1219
Round JL, Mazmanian SK (2009) The gut microbiota shapes intestinal immune responses during health and disease. Nat Rev Immunol 9:313–323. https://doi.org/10.1038/nri2515
Rungrassamee W, Klanchui A, Chaiyapechara S, Maibunkaew S, Tangphatsornruang S, Jiravanichpaisal P, Karoonuthaisiri N (2013) Bacterial population in intestines of the black tiger shrimp (Penaeus monodon) under different growth stages. PLoS One 8:e60802. https://doi.org/10.1371/journal.pone.0060802
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. https://doi.org/10.1186/gb-2011-12-6-r60
Sriurairatana S, Boonyawiwat V, Gangnonngiw W, Laosutthipong C, Hiranchan J, Flegel TW (2014) White feces syndrome of shrimp arises from transformation, sloughing and aggregation of hepatopancreatic microvilli into vermiform bodies superficially resembling gregarines. PLoS One 9:e99170. https://doi.org/10.1371/journal.pone.0099170
Sung HH, Hsu SF, Chen CK, Ting YY, Chao WL (2001) Relationships between disease outbreak in cultured tiger shrimp (Penaeus monodon) and the composition of Vibrio communities in pond water and shrimp hepatopancreas during cultivation. Aquaculture 192:101–110. https://doi.org/10.1016/s0044-8486(00)00458-0
Tarnecki AM, Rhody NR (2017) Microbiota of common snook Centropomus undecimalis larvae exhibiting high mortality. Aquac Res. https://doi.org/10.1111/are.13377
Tran L, Nunan L, Redman RM, Mohney LL, Pantoja CR, Fitzsimmons K, Lightner DV (2013) Determination of the infectious nature of the agent of acute hepatopancreatic necrosis syndrome affecting penaeid shrimp. Dis Aquat Org 105:45–55. https://doi.org/10.3354/dao02621
Wang HL, Wan XY, Xie GS, Dong X, Wang XH, Huang J (2020) Insights into the histopathology and microbiome of Pacific white shrimp, Penaeus vannamei, suffering from white feces syndrome. Aquaculture 527:735447. https://doi.org/10.1016/j.aquaculture.2020.735447
Xin X, Qin JW (2019) Rapid start-up of partial nitritation in aerobic granular sludge bioreactor and the analysis of bacterial community dynamics. Bioprocess Biosyst Eng 42:1973–1981. https://doi.org/10.1007/s00449-019-02190-x
Xiong J, Zhu J, Zhang D (2014) The application of bacterial indicator phylotypes to predict shrimp health status. Appl Microbiol Biotechnol 98:8291–8299. https://doi.org/10.1007/s00253-014-5941-y
Xiong J, Wang K, Wu J, Qiuqian L, Yang K, Qian Y, Zhang D (2015) Changes in intestinal bacterial communities are closely associated with shrimp disease severity. Appl Microbiol Biotechnol 99:6911–6919. https://doi.org/10.1007/s00253-015-6632-z
Xiong J, Zhu J, Dai W, Dong C, Qiu Q, Li C (2017) Integrating gut microbiota immaturity and disease-discriminatory taxa to diagnose the initiation and severity of shrimp disease. Environ Microbiol 19:1490–1501. https://doi.org/10.1111/1462-2920.13701
Xue M, Wu LY, He YY, Liang HF, Wen CQ (2018) Biases during DNA extraction affect characterization of the microbiota associated with larvae of the Pacific white shrimp, Litopenaeus vannamei. PeerJ 6:e5257. https://doi.org/10.7717/peerj.5257
Yu W, Cao J, Dai W, Qiu Q, Xiong J (2018) Quantitative PCR analysis of gut disease-discriminatory phyla for determining shrimp disease incidence. Appl Environ Microbiol 84:e01387–18. https://doi.org/10.1128/AEM.01387-18
Zeng S, Huang Z, Hou D, Liu J, Weng S, He J (2017) Composition, diversity and function of intestinal microbiota in Pacific white shrimp (Litopenaeus vannamei) at different culture stages. PeerJ 5:e3986. https://doi.org/10.7717/peerj.3986
Zhang BC, Liu F, Bian HH, Liu J, Pan LQ, Huang J (2012) Isolation, identification, and pathogenicity analysis of a Vibrio parahaemolyticus strain from Litopenaeus vannamei. Prog Fish Sci 33:56–62 (in Chinese)
Zhang D, Wang X, Xiong J, Zhu J, Wang Y, Zhao Q, Chen H, Guo A, Wu J, Dai H (2014) Bacterioplankton assemblages as biological indicators of shrimp health status. Ecol Indic 38:218–224. https://doi.org/10.1016/j.ecolind.2013.11.002
Zheng Y, Yu M, Liu J, Qiao Y, Wang L, Li Z, Zhang XH, Yu M (2017) Bacterial community associated with healthy and diseased Pacific white shrimp (Litopenaeus vannamei) larvae and rearing water across different growth stages. Front Microbiol 8:1362. https://doi.org/10.3389/fmicb.2017.01362
Zokaeifar H, Babaei N, Saad CR, Kamarudin MS, Sijam K, Balcazar JL (2014) Administration of Bacillus subtilis strains in the rearing water enhances the water quality, growth performance, immune response, and resistance against Vibrio harveyi infection in juvenile white shrimp, Litopenaeus vannamei. Fish Shellfish Immunol 36:68–74. https://doi.org/10.1016/j.fsi.2013.10.007
Funding
This research was supported by the grants from Pilot National Laboratory for Marine Science and Technology (Qingdao) (No. QNLM201706), China Agriculture Research System (CARS-48), Central Public-interest Scientific Institution Basal Research Fund, CAFS (NO. 2020TD39), China Postdoctoral Science Foundation (2018M640666), and Applied Research Project for Postdoctoral Researchers in Qingdao.
Author information
Authors and Affiliations
Contributions
HW and JH designed the study, performed data analysis, and wrote the manuscript. HW conducted the experiments and collected samples. PW and TL provided help when collecting samples.
Corresponding author
Ethics declarations
Conflict of interest
Authors PW and TL are employed by the company Hainan Zhongzheng Aquatic Science and Technology CO. LTD. The remaining authors declare that the research was conducted in the absence of any commercial or financial relationships that could be construed as a potential conflict of interest.
Ethical approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wang, H., Huang, J., Wang, P. et al. Insights into the microbiota of larval and postlarval Pacific white shrimp (Penaeus vannamei) along early developmental stages: a case in pond level. Mol Genet Genomics 295, 1517–1528 (2020). https://doi.org/10.1007/s00438-020-01717-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01717-2