Abstract
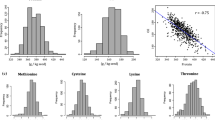
As a globally important legume crop, soybean provides excellent sources of protein and oil for human and livestock nutrition. Improving seed protein and oil contents has always been an important objective in soybean breeding. Water-soluble protein plays a significant role in the processing and efficacy of soybean protein. Here, a genome-wide association study (GWAS) of seed compositions (protein, oil, and water-soluble protein contents) was conducted using 211 diverse soybean accessions genotyped with a 355 K SoySNP array. Three, four, and five QTLs were identified related to the protein, oil, and water-soluble protein contents, respectively. Furthermore, five QTLs (qPC-15-1, qOC-8-1, qOC-12-1, qOC-20-1 and qWSPC-8-1) were detected in multiple environments. Analysis of the favorable alleles for oil and water-soluble protein contents showed that qOC-8-1 (qWSPC-8-1) exerted inverse effects on oil and water-soluble protein synthesis. Relative expression analysis suggested that Glyma.15G049200 in qPC-15-1 affects protein synthesis and Glyma.08G107800 in qOC-8-1 and qWSPC-8-1 might be involved in oil and water-soluble protein synthesis, producing opposite effects. The candidate genes and significant SNPs detected in the present study will allow a deeper understanding of the genetic basis for the regulation of protein, oil and water-soluble protein contents and provide important information that could be utilized in marker-assisted selection for soybean quality improvement.




Similar content being viewed by others
References
Bachlava E, Dewey RE, Burton JW, Cardinal AJ (2009) Mapping and comparison of quantitative trait loci for oleic acid seed content in two segregating soybean populations. Crop Sci 49:433–442
Barrett JC, Fry B, Maller J, Daly MJ (2004) Haploview: analysis and visualization of LD and haplotype maps. Bioinformatics 21:263–265
Chaudhary J, Patil G, Sonah H, Deshmukh R, Vuong T, Valliyodan B, Nguyen H (2015) Expanding omics resources for improvement of soybean seed composition traits. Front Plant Sci 6:1021
Chen LQ, Qu XQ, Hou BH, Sosso D, Osorio S, Fernie AR, Frommer WB (2012) Sucrose efflux mediated by SWEET proteins as a key step for phloem transport. Science 335:207–211
Cheng L, Yuan HY, Ren R, Zhao SQ, Han YP, Zhou QY, Ke DX, Wang YX, Wang L (2016) Genome-wide identification, classification, and expression analysis of amino acid transporter gene family in Glycine Max. Front Plant Sci 7:515
Chung J, Babka HL, Graef GL, Staswick PE, Lee DJ, Cregan PB, Shoemaker RC, Specht JE (2003) The seed protein, oil, and yield QTL on soybean linkage group I. Crop Sci 43:1053–1067
Clemente TE, Cahoon EB (2009) Soybean oil: genetic approaches for modification of functionality and total content. Plant Physiol 151:1030–1040
de Kraker JW, Gershenzon J (2011) From amino acid to glucosinolate biosynthesis: protein sequence changes in the evolution of methylthioalkylmalate synthase in Arabidopsis. Plant Cell 23:38–53
Diers BW, Keim P, Fehr W, Shoemaker R (1992) RFLP analysis of soybean seed protein and oil content. Theor Appl Genet 83:608–612
Eskandari M, Cober ER, Rajcan I (2013) Genetic control of soybean seed oil: II. QTL and genes that increase oil concentration without decreasing protein or with increased seed yield. Theor Appl Genet 126:1677–1687
Fait A, Nesi AN, Angelovici R, Lehmann M, Pham PA, Song L, Haslam RP, Napier JA, Galili G, Fernie AR (2011) Targeted enhancement of glutamate-to-γ-aminobutyrate conversion in arabidopsis seeds affects carbon-nitrogen balance and storage reserves in a development-dependent manner. Plant Physiol 157:1026–1042
Gruis D, Selinger DA, Curran JM, Jung R (2002) Redundant proteolytic mechanisms process seed storage proteins in the bbsence of seed-type members of the vacuolar processing enzyme family of cysteine proteases. Plant Cell 14:2863–2882
Guo B, Sleper DA, Nguyen HT, Arelli PR, Shannon JG (2006) Quantitative trait loci underlying resistance to three soybean cyst nematode populations in soybean PI 404198A. Crop Sci 46:224–233
Hu D, Kan G, Hu W, Li Y, Hao D, Li X, Yang H, Yang Z, He X, Huang F, Yu D (2019) Identification of loci and candidate genes responsible for pod dehiscence in soybean via genome-wide association analysis across multiple environments. Front Plant Sci 10:811
Hyten DL, Pantalone VR, Sams CE, Saxton AM, Landau-Ellis D, Stefaniak TR, Schmidt ME (2004) Seed quality QTL in a prominent soybean population. Theor Appl Genet 109:552–561
Kim M, Schultz S, Nelson RL, Diers BW (2016) Identification and fine mapping of a soybean seed protein QTL from PI 407788A on chromosome 15. Crop Sci 56:219–225
Lam HM, Peng SS, Coruzzi GM (1994) Metabolic regulation of the gene encoding glutamine-dependent asparagine synthetase in Arabidopsis thaliana. Plant Physiol 106:1347–1357
Li C, Zhang YM (2011) Molecular evolution of glycinin and beta-conglycinin gene families in soybean (Glycine max L. Merr.). Heredity 106:633–641
Li T, Ma X, Li N, Zhou L, Liu Z, Han H, Gui Y, Bao Y, Chen J, Dai X (2017) Genome-wide association study discovered candidate genes of Verticillium wilt resistance in upland cotton (Gossypium hirsutum L.). Plant Biotechnol J 15:1520–1532
Li YH, Reif JC, Hong HL, Li HH, Liu ZX, Ma YS, Li J, Tian Y, Li YF, Li WB, Qiu LJ (2018) Genome-wide association mapping of QTL underlying seed oil and protein contents of a diverse panel of soybean accessions. Plant Sci 266:95–101
Li S, Xu H, Yang J, Zhao T (2019) Dissecting the genetic architecture of seed protein and oil content in soybean from the Yangtze and Huaihe River Valleys using multi-locus genome-wide association studies. Int J Mol Sci 20:3041
Liang HZ, Yu YL, Wang SF, Lian Y, Wang TF, Wei YL, Gong PT, Liu XY, Fang XJ, Zhang MC (2010) QTL mapping of isoflavone, oil and protein contents in soybean (Glycine max L. Merr.). Ag Sci China 9:1108–1116
Lipka AE, Tian F, Wang Q, Peiffer J, Li M, Bradbury PJ, Gore MA, Buckler ES, Zhang Z (2012) GAPIT: genome association and prediction integrated tool. Bioinformatics 28:2397–2399
Liu YF, Li QT, Lu X, Song QX, Lam SM, Zhang WK, Ma B, Lin Q, Man WQ, Du WG, Shui GH, Chen SY, Zhang JS (2014) Soybean GmMYB73 promotes lipid accumulation in transgenic plants. BMC Plant Biol 14:73
Lu W, Wen Z, Li H, Yuan D, Li J, Zhang H, Huang Z, Cui S, Du W (2013) Identification of the quantitative trait loci (QTL) underlying water soluble protein content in soybean. Theor Appl Genet 126:425–433
Mansur L, Lark K, Kross H, Oliveira A (1993) Interval mapping of quantitative trait loci for reproductive, morphological, and seed traits of soybean (Glycine max L.). Theor Appl Genet 86:907–913
Mao T, Jiang Z, Han Y, Teng W, Zhao X, Li W (2013) Identification of quantitative trait loci underlying seed protein and oil contents of soybean across multi-genetic backgrounds and environments. Plant Breed 132:630–641
Meyer K, Stecca KL, Ewell-Hicks K, Allen SM, Everard JD (2012) Oil and protein accumulation in developing seeds is influenced by the expression of a cytosolic pyrophosphatase in Arabidopsis. Plant Physiol 159:1221–1234
Ning H, Zhang C, Yao Y, Yu D (2010) Overexpression of a soybean O-acetylserine (thiol) lyase-encoding gene GmOASTL4 in tobacco increases cysteine levels and enhances tolerance to cadmium stress. Biotechnol Lett 32:557–564
Oyoo ME, Benitez ER, Kurosaki H, Ohnishi S, Miyoshi T, Kiribuchi-Otobe C, Horigane A, Takahashi R (2011) QTL analysis of soybean seed coat discoloration associated with II TT genotype. Crop Sci 51:464–469
Panthee DR, Pantalone VR, Saxton AM, West DR, Sams CE (2006) Genomic regions associated with amino acid composition in soybean. Mol Breed 17:79–89
Pathan SM, Vuong T, Clark K, Lee J-D, Shannon JG, Roberts CA, Ellersieck MR, Burton JW, Cregan PB, Hyten DL, Nguyen HT, Sleper DA (2013) Genetic mapping and confirmation of quantitative trait loci for seed protein and oil contents and seed weight in soybean. Crop Sci 53:765–774
Patil G, Valliyodan B, Deshmukh R, Prince S, Nicander B, Zhao M, Sonah H, Song L, Lin L, Chaudhary J, Liu Y, Joshi T, Xu D, Nguyen HT (2015) Soybean (Glycine max) SWEET gene family: insights through comparative genomics, transcriptome profiling and whole genome re-sequence analysis. BMC Genom 16:520
Patil G, Mian R, Vuong T, Pantalone V, Song Q, Chen P, Shannon GJ, Carter TC, Nguyen HT (2017) Molecular mapping and genomics of soybean seed protein: a review and perspective for the future. Theor Appl Genet 130:1975–1991
Phansak P, Soonsuwon W, Hyten D, Song Q, Cregan P, Graef G, Specht J (2016) Multi-population selective genotyping to identify soybean [Glycine max (L.) Merr.] seed protein and oil QTLs. G3 (Bethesda) 6:1635–1648
Powell JE, Henders AK, McRae AF, Caracella A, Smith S, Wright MJ, Whitfield JB, Dermitzakis ET, Martin NG, Visscher PM, Montgomery GW (2012) The brisbane systems genetics study: genetical genomics meets complex trait genetics. PLoS ONE 7:e35430
Reinprecht Y, Poysa VW, Yu K, Rajcan I, Ablett GR, Pauls KP (2006) Seed and agronomic QTL in low linolenic acid, lipoxygenase-free soybean (Glycine max (L.) Merrill) germplasm. Genome 49:1510–1527
Si L, Chen J, Huang X, Gong H, Luo J, Hou Q, Zhou T, Lu T, Zhu J, Shangguan Y, Chen E, Gong C, Zhao Q, Jing Y, Zhao Y, Li Y, Cui L, Fan D, Lu Y, Weng Q, Wang Y, Zhan Q, Liu K, Wei X, An K, An G, Han B (2016) OsSPL13 controls grain size in cultivated rice. Nat Genet 48:447–456
Singh RJ, Hymowitz T (1999) Soybean genetic resources and crop improvement. Genome 42:605–616
Smith AJ, Rinne RW, Seif RD (1989) Phosphoenolpyruvate carboxylase and pyruvate kinase involvement in protein and oil biosynthesis during soybean seed development. Crop Sci 29:349–353
Song QX, Li QT, Liu YF, Zhang FX, Ma B, Zhang WK, Man WQ, Du WG, Wang GD, Chen SY, Zhang JS (2013) Soybean GmbZIP123 gene enhances lipid content in the seeds of transgenic Arabidopsis plants. J Exp Bot 64:4329–4341
Specht J, Chase K, Macrander M, Graef G, Chung J, Markwell J, Germann M, Orf J, Lark K (2001) Soybean response to water: a QTL analysis of drought tolerance. Crop Sci 41:493–509
Tajuddin T, Watanabe S, Yamanaka N, Harada K (2003) Analysis of quantitative trait loci for protein and lipid contents in soybean seeds using recombinant inbred lines. Breed Sci 53:133–140
Van K, McHale LK (2017) Meta-analyses of QTLs associated with protein and oil contents and compositions in soybean [Glycine max (L.) Merr.] seed. Int J Mol Sci 18:1180
Wang HW, Zhang B, Hao YJ, Huang J, Tian AG, Liao Y, Zhang JS, Chen SY (2007) The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J 52:716–729
Wang J, Chu S, Zhang H, Zhu Y, Cheng H, Yu D (2016) Development and application of a novel genome-wide SNP array reveals domestication history in soybean. Sci Rep 6:20728
Wang L, Yang Y, Zhang S, Che Z, Yuan W, Yu D (2020) GWAS reveals two novel loci for photosynthesis-related traits in soybean. Mol Genet Genom. https://doi.org/10.1007/s00438-020-01661-1
Warrington CV, Abdel-Haleem H, Hyten DL, Cregan PB, Orf JH, Killam AS, Bajjalieh N, Li Z, Boerma HR (2015) QTL for seed protein and amino acids in the Benning × Danbaekkong soybean population. Theor Appl Genet 128:839–850
Wu X, Blake S, Sleper D, Shannon J, Cregan P, Nguyen H (2009) QTL, additive and epistatic effects for SCN resistance in PI 437654. Theor Appl Genet 118:1093–1105
Xuan L, Zhang C, Yan T, Wu D, Hussain N, Li Z, Chen M, Pan J, Jiang L (2018) TRANSPARENT TESTA 4-mediated flavonoids negatively affect embryonic fatty acid biosynthesis in Arabidopsis. Plant Cell Environ 41:2773–2790
Zhang C, Meng Q, Gai J, Yu D (2008) Cloning and functional characterization of an O-acetylserine(thiol)lyase-encoding gene in wild soybean (Glycine soja). Mol Biol Rep 35:527–534
Zhang D, Kan G, Hu Z, Cheng H, Zhang Y, Wang Q, Wang H, Yang Y, Li H, Hao D, Yu D (2014) Use of single nucleotide polymorphisms and haplotypes to identify genomic regions associated with protein content and water-soluble protein content in soybean. Theor Appl Genet 127:1905–1915
Zhang D, Lu H, Chu S, Zhang H, Zhang H, Yang Y, Li H, Yu D (2017) The genetic architecture of water-soluble protein content and its genetic relationship to total protein content in soybean. Sci Rep 7:5053
Zhang D, Zhang H, Hu Z, Chu S, Yu K, Lv L, Yang Y, Zhang X, Chen X, Kan G, Tang Y, An YC, Yu D (2019) Artificial selection on GmOLEO1 contributes to the increase in seed oil during soybean domestication. PLoS Genet 15:e1008267
Zhao X, Dong H, Chang H, Zhao J, Teng W, Qiu L, Li W, Han Y (2019) Genome wide association mapping and candidate gene analysis for hundred seed weight in soybean [Glycine max (L.) Merrill]. BMC Genom 20:648
Zhou Z, Jiang Y, Wang Z, Gou Z, Lyu J, Li W, Yu Y, Shu L, Zhao Y, Ma Y, Fang C, Shen Y, Liu T, Li C, Li Q, Wu M, Wang M, Wu Y, Dong Y, Wan W, Wang X, Ding Z, Gao Y, Xiang H, Zhu B, Lee S-H, Wang W, Tian Z (2015) Resequencing 302 wild and cultivated accessions identifies genes related to domestication and improvement in soybean. Nat Biotechnol 33:408–414
Zhu-Shimoni JX, Galili G (1998) Expression of an arabidopsis aspartate kinase/homoserine dehydrogenase gene is metabolically regulated by photosynthesis-related signals but not by nitrogenous compounds. Plant Physiol 116:1023–1028
Acknowledgements
This work was supported in part by the Ministry of Science and Technology (2017YFE0111000), the Fundamental Research Funds for the Central Universities (KYZ201705), the National Natural Science Foundation of China (31571688, 31871649, 31671715), and the Ministry of Agriculture (2016ZX08004-003, 2016ZX08009003-004).
Author information
Authors and Affiliations
Contributions
GK, DY, and SZ designed the research, and SZ, DH, SZ, and DZ performed the research. SZ, HW, and HD analyzed the data, and SZ wrote the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies involving human participants or animals performed by any of the authors.
Additional information
Communicated by Stefan Hohmann.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, S., Hao, D., Zhang, S. et al. Genome-wide association mapping for protein, oil and water-soluble protein contents in soybean. Mol Genet Genomics 296, 91–102 (2021). https://doi.org/10.1007/s00438-020-01704-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00438-020-01704-7