Abstract
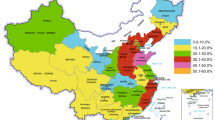
The aim of the study was to identify the Cryptosporidium parvum subtypes circulating in Polish cattle and their distribution in relation to the age and health status of tested animals. In total, 779 fecal samples were obtained from young cattle originating from 237 farms. C. parvum strains were identified at the 18 small-subunit ribosomal RNA (SSU rRNA), COWP, and LIB13 loci and were subsequently analyzed by sequencing at the 60-kDa glycoprotein (GP60) locus for subtype determination. The presence of 71 C. parvum strains belonging to IIa, IId, or IIl subtype families was shown. The strains from the IIa allele family prevailed with IIaA17G1R1, IIaA17G2R1, and IIaA15G2R1 subtypes occurring frequently. Two novel subtypes IIaA10G1R1 and IIlA19R3 were detected for the first time in a bovine host. The highest C. parvum prevalence (22.5 %, confidence interval (CI) = 2.5 %) was observed among the youngest animals up to 2 weeks of age, followed by the prevalence among those aged 2 to 4 weeks (6.6 %, CI = 2.6 %) and then among older cattle (4.9 %, CI = 2.1). The occurrence of diarrhea in animals was associated with the presence of the IIaA16G1R1b subtype, while infections caused by IIaA15G2R1 strains were more likely to be asymptomatic. The geographical distribution of subtypes revealed that strains from the IIa subtype family were detected all over the country frequently compared to the IId and IIl subtypes, the sporadic appearances of which confirmed their endemic occurrence. Subtype analysis revealed the presence of zoonotic strains indicating cattle as a reservoir for human cryptosporidiosis.




Similar content being viewed by others
References
Abe N, Makoto M, Isao K, Iseki M (2006) Subgenotype analysis of Cryptosporidium parvum isolates from humans and animals in Japan using the 60-kDa glycoprotein gene sequences. Parasitol Res 99:303–305
Abeywardena H, Jex AR, Nolan MJ, Haydon SR, Stevens MA, McAnulty RW, Gasser RB (2012) Genetic characterisation of Cryptosporidium and Giardia from dairy calves: discovery of species/genotypes consistent with those found in humans. Infect Genet Evol 12:1984–1993
Alves M, Xiao L, Sulaiman I, Lal AA, Matos O, Antunes F (2003) Subgenotype analysis of Cryptosporidium isolates from humans, cattle, and zoo ruminants in Portugal. J Clin Microbiol 41:2744–2747
Alves M, Xiao L, Antunes F, Matos O (2006) Distribution of Cryptosporidium subtypes in humans and domestic and wild ruminants in Portugal. Parasitol Res 99:287–292
Amer S, Harfoush M, He H (2010) Molecular and phylogenetic analyses of Cryptosporidium spp from dairy cattle in Egypt. J Egypt Soc Parasitol 40:349–366
Björkman C, Lindström L, Oweson C, Ahola H, Troell K, Axén C (2015) Cryptosporidium infections in suckler herd beef calves. Parasitology 142:1108–1114
Broglia A, Reckinger S, Cacció SM, Nöckler K (2008) Distribution of Cryptosporidium parvum subtypes in calves in Germany. Vet Parasitol 154:8–13
Brook EJ, Anthony Hart C, French NP, Christley RM (2009) Molecular epidemiology of Cryptosporidium subtypes in cattle in England. Vet J 179:378–382
Budu-Amoako E, Greenwood SJ, Dixon BR, Barkema HW, McClure JT (2012) Occurrence of Cryptosporidium and Giardia on beef farms and water sources within the vicinity of the farms on Prince Edward Island, Canada. Vet Parasitol 184:1–9
Cacciò SM, Thompson RC, McLauchlin J, Smith HV (2005) Unravelling Cryptosporidium and Giardia epidemiology. Trends Parasitol 21:430–437
Cama VA, Ross JM, Crawford S, Kawai V, Chavez-Valdez R, Vargas D, Vivar A, Ticona E, Navincopa M, Williamson J, Ortega Y, Gilman RH, Bern C, Xiao L (2007) Differences in clinical manifestations among Cryptosporidium species and subtypes in HIV-infected persons. J Infect Dis 196:68
Chalmers RM, Smith RP, Hadfield SJ, Elwin K, Giles M (2011) Zoonotic linkage and variation in Cryptosporidium parvum from patients in the United Kingdom. Parasitol Res 108:1321–1325
de Graaf DC, Vanopdenbosch E, Ortega-Mora LM, Abbassi H, Peeters JE (1999) A review of the importance of cryptosporidiosis in farm animals. Int J Parasitol 29:1269–1287
Díaz P, Hadfield SJ, Quílez J, Soilán M, López C, Panadero R, Díez-Baños P, Morrondo P, Chalmers RM (2012) Assessment of three methods for multilocus fragment typing of Cryptosporidium parvum from domestic ruminants in north west Spain. Vet Parasitol 186:188–195
Drumo R, Widmer G, Morrison LJ, Tait A, Grelloni V, D’Avino N, Pozio E, Cacciò SM (2012) Evidence of host-associated populations of Cryptosporidium parvum in Italy. Appl Environ Microbiol 78:3523–3529
Duranti A, Cacciò SM, Pozio E, Di Egidio A, De Curtis M, Battisti A, Scaramozzino P (2009) Risk factors associated with Cryptosporidium parvum infection in cattle. Zoonoses Public Health 56:176–182
Epe C, Coati N, Schnieder T (2004) Results of parasitological examinations of faecal samples from horses, ruminants, pigs, dogs, cats, hedgehogs and rabbits between 1998 and 2002. Dtsch Tierarztl Wochenschr 111:243–247
Gait R, Soutar RH, Hanson M, Fraser C, Chalmers R (2008) Outbreak of cryptosporidiosis among veterinary students. Vet Rec 162:843–845
Geurden T, Berkvens D, Martens C, Casaert S, Vercruysse J, Claerebout E (2007) Molecular epidemiology with subtype analysis of Cryptosporidium in calves in Belgium. Parasitology 134:1981–1987
Glaberman S, Moore JE, Lowery CJ, Chalmers RM, Sulaiman I, Elwin K, Rooney PJ, Millar BC, Dooley JS, Lal AA, Xiao L (2002) Three drinking-water-associated cryptosporidiosis outbreaks, Northern Ireland. Emerg Infect Dis 8:631–633
Hijjawi N, Ng J, Yang R, Atoum MF, Ryan U (2010) Identification of rare and novel Cryptosporidium GP60 subtypes in human isolates from Jordan. Exp Parasitol 125:161–164
Homan W, van Gorkom T, Kan YY, Hepener J (1999) Characterization of Cryptosporidium parvum in human and animal feces by single-tube nested polymerase chain reaction and restriction analysis. Parasitol Res 85:707–712
Hunter PR, Thompson RC (2005) The zoonotic transmission of Giardia and Cryptosporidium. Int J Parasitol 35:1181–1190
Imre K, Matos O, Dărăbus G, Mederle N, Oprescu I, Morariu S, Ilie MS, Hotea I, Imre M (2009) First genetic identification of Cryptosporidium spp. in cattle in Romania. Lucr Şt Med Vet Timişoara 52:26–30
Imre K, Dărăbuş G, Mederle N, Oprescu I, Morariu S, Ilie M, Hotea I, Imre M, Indre D, Balint A, Sorescu D (2010) Intraspecific characterization of some Cryptosporidium parvum isolates from calves and lambs in Western Romania using molecular techniques. Sci Parasitol 11:47–50
Insulander M, Silverlås C, Lebbad M, Karlsson L, Mattsson JG, Svenungsson B (2013) Molecular epidemiology and clinical manifestations of human cryptosporidiosis in Sweden. Epidemiol Infect 141:1009–1020
Kaupke A, Kwit E, Chalmers RM, Michalski MM, Rzeżutka A (2014) An outbreak of massive mortality among farm rabbits associated with Cryptosporidium infection. Res Vet Sci 97:85–87
Lassen B, Ståhl M, Enemark HL (2014) Cryptosporidiosis—an occupational risk and a disregarded disease in Estonia. Acta Vet Scand 5:56–36
Leoni F, Amar C, Nichols G, Pedraza-Díaz S, McLauchlin J (2006) Genetic analysis of Cryptosporidium from 2414 humans with diarrhoea in England between 1985 and 2000. J Med Microbiol 55:703–707
Liu H, Shen Y, Yin J, Yuan Z, Jiang Y, Xu Y, Pan W, Hu Y, Cao J (2014) Prevalence and genetic characterization of Cryptosporidium, Enterocytozoon, Giardia and Cyclospora in diarrheal outpatients in China. BMC Infect Dis 14:25. doi:10.1186/1471-2334-14-25
Majewska AC, Werner A, Sulima P, Luty T (1999) Survey on equine cryptosporidiosis in Poland and the possibility of zoonotic transmission. Ann Agric Environ Med 6:161–165
Majewska AC, Werner A, Sulima P, Luty T (2000) Prevalence of Cryptosporidium in sheep and goats bred on five farms in west-central region of Poland. Vet Parasitol 89:269–275
Majewska AC, Werner A, Sulima P (2001) Występowanie kryptosporidiozy u bydła hodowlanego w jednym gospodarstwie rolnym—całoroczne badania. Wiad Parazytol 47(suppl 2):31
Majewska AC, Solarczyk P, Tamang L, Graczyk TK (2004) Equine Cryptosporidium parvum infections in western Poland. Parasitol Res 93:274–278
Millar C, Moore J, Lowery C, McCorry K, Dooley J (2001) Successful PCR amplification of genomic DNA from Cryptosporidium parvum oocysts extracted from a human faecal sample: a rapid and simple method suited for outbreak analysis. Int J Hyg Environ Health 204 :191 –194
Mišic Z, Abe N (2007) Subtype analysis of Cryptosporidium parvum isolates from calves on farms around Belgrade, Serbia and Montenegro, using the 60 kDa glycoprotein gene sequences. Parasitology 134:351–358
Muhid A, Robertson I, Ng J, Ryan U (2011) Prevalence of and management factors contributing to Cryptosporidium sp. infection in pre-weaned and post-weaned calves in Johor, Malaysia. Exp Parasitol 127:534–538
O’Brien E, McInnes L, Ryan U (2008) Cryptosporidium GP60 genotypes from humans and domesticated animals in Australia, North America and Europe. Exp Parasitol 118:118–121
Olson ME, O’Handley RM, Ralston BJ, McAllister TA, Thompson RC (2004) Update on Cryptosporidium and Giardia infections in cattle. Trends Parasitol 20:185–191
Plutzer J, Karanis P (2007) Genotype and subtype analyses of Cryptosporidium isolates from cattle in Hungary. Vet Parasitol 146:357–362
Quilez J, Torres E, Chalmers RM, Robinson G, Del Cacho E, Sanchez-Acedo C (2008) Cryptosporidium species and subtype analysis from dairy calves in Spain. Parasitology 135:1613–1620
Rieux A, Chartier C, Pors I, Delafosse A, Paraud C (2013) Molecular characterization of Cryptosporidium isolates from high-excreting young dairy calves in dairy cattle herds in Western France. Parasitol Res 112:3423–3431
Ryan U, Hijjawi N (2015) New developments in Cryptosporidium research. Int J Parasitol 45:367–373
Rzeżutka A, Kaupke A (2013) Occurrence and molecular identification of Cryptosporidium species isolated from cattle in Poland. Vet Parasitol 196:301–306
Rzeżutka A, Kaupke A, Kozyra I, Pejsak Z (2014) Molecular studies on pig cryptosporidiosis in Poland. Pol J Vet Sci 17:577–582
Santín M, Trout JM (2008) Livestock. In: Fayer R, Xiao L (eds) Cryptosporidium and cryptosporidiosis. CRC, Boca Raton, pp 451–483
Silverlås C, Näslund K, Björkman C, Mattsson JG (2010) Molecular characterisation of Cryptosporidium isolates from Swedish dairy cattle in relation to age, diarrhoea and region. Vet Parasitol 169:289–295
Silverlås C, Bosaeus-Reineck H, Näslund K, Björkman C (2013) Is there a need for improved Cryptosporidium diagnostics in Swedish calves? Int J Parasitol 43:155–161
Soba B, Logar J (2008) Genetic classification of Cryptosporidium isolates from humans and calves in Slovenia. Parasitology 135:1263–1270
Stantič-Pavlinic M, Xiao L, Glaberman S, Lal AA, Orazen T, Rataj-Verglez A, Logar J, Berce I (2003) Cryptosporidiosis associated with animal contacts. Wien Klin Wochenschr 115:125–127
Sulaiman IM, Hira PR, Zhou L, Al-Ali FM, Al-Shelahi FA, Shweiki HM, Iqbal J, Khalid N, Xiao L (2005) Unique endemicity of cryptosporidiosis in children in Kuwait. J Clin Microbiol 43:2805–2809
Tanriverdi S, Arslan MO, Akiyoshi DE, Tzipori S, Widmer G (2003) Identification of genotypically mixed Cryptosporidium parvum populations in humans and calves. Mol Biochem Parasitol 130:13–22
Thompson HP, Dooley JS, Kenny J, McCoy M, Lowery CJ, Moore JE, Xiao L (2007) Genotypes and subtypes of Cryptosporidium spp. in neonatal calves in Northern Ireland. Parasitol Res 100:619–624
Waldron LS, Ferrari BC, Power ML (2009) Glycoprotein 60 diversity in C. hominis and C. parvum causing human cryptosporidiosis in NSW, Australia. Exp Parasitol 122:124–127
Wang R, Zhang L, Axén C, Bjorkman C, Jian F, Amer S, Liu A, Feng Y, Li G, Lv C, Zhao Z, Qi M, Dong H, Wang H, Ning C, Sun Y, Xiao L (2014) Cryptosporidium parvum IId family: clonal population and dispersal from Western Asia to other geographical regions. Sci Rep 27:4208
Wielinga PR, de Vries A, van der Goot TH, Mank T, Mars MH, Kortbeek LM, van der Giessen JW (2008) Molecular epidemiology of Cryptosporidium in humans and cattle in The Netherlands. Int J Parasitol 38:809–817
Xiao L (2010) Molecular epidemiology of cryptosporidiosis: an update. Exp Parasitol 124:80–89
Xiao L, Escalante L, Yang C, Sulaiman I, Escalante AA, Montali RJ, Fayer R, Lal AA (1999) Phylogenetic analysis of Cryptosporidium parasites based on the small-subunit rRNA gene locus. Appl Environ Microbiol 65:1578–1583
Zhang W, Wang R, Yang F, Zhang L, Cao J, Zhang X, Ling H, Liu A, Shen Y (2013) Distribution and genetic characterizations of Cryptosporidium spp. in pre-weaned dairy calves in Northeastern China’s Heilongjiang Province. PLoS ONE 8(1), e54857
Zintl A, Neville D, Maguire D, Fanning S, Mulcahy G, Smith HV, De Waal T (2007) Prevalence of Cryptosporidium species in intensively farmed pigs in Ireland. Parasitol 134:1575–1582
Acknowledgments
The authors thank Dr. Rachel M. Chalmers, Cryptosporidium Reference Unit in Swansea, UK, for her valuable comments on the manuscript; Prof. Mirosław M. Michalski, Department of Parasitology and Invasive Diseases, Faculty of Veterinary Medicine, University of Warmia and Mazury in Olsztyn, Poland, for the help in sampling arrangement; as well as Dr. Iwona Kozyra, Department of Food and Environmental Virology, National Veterinary Research Institute in Puławy, Poland, for the assistance in the graphical presentation of data. The study was supported by the Ministry of Science and Higher Education of Poland (research project no. S/071).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Kaupke, A., Rzeżutka, A. Emergence of novel subtypes of Cryptosporidium parvum in calves in Poland. Parasitol Res 114, 4709–4716 (2015). https://doi.org/10.1007/s00436-015-4719-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00436-015-4719-1