Abstract
Spatial transcriptomics (ST) provides novel insights into the tumor microenvironment (TME). ST allows the quantification and illustration of gene expression profiles in the spatial context of tissues, including both the cancer cells and the microenvironment in which they are found. In cancer research, ST has already provided novel insights into cancer metastasis, prognosis, and immunotherapy responsiveness. The clinical precision oncology application of next-generation sequencing (NGS) and RNA profiling of tumors relies on bulk methods that lack spatial context. The ability to preserve spatial information is now possible, as it allows us to capture tumor heterogeneity and multifocality. In this narrative review, we summarize precision oncology, discuss tumor sequencing in the clinic, and review the available ST research methods, including seqFISH, MERFISH (Vizgen), CosMx SMI (NanoString), Xenium (10x), Visium (10x), Stereo-seq (STOmics), and GeoMx DSP (NanoString). We then review the current ST literature with a focus on solid tumors organized by tumor type. Finally, we conclude by addressing an important question: how will spatial transcriptomics ultimately help patients with cancer?
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Spatial transcriptomics (ST) is the study and quantification of messenger RNA (mRNA) transcripts as a surrogate for gene expression in the spatial context of cancer and the associated microenvironment (Marx 2021; Moses and Pachter 2022). The use of ST in cancer research has the potential to have a significant impact on patients through its application in precision oncology care. This highly detailed illustration of gene expression has been made possible with the recent development of a number of spatial platforms.
This review summarizes tumor sequencing in the clinical setting and tumor sequencing in the research laboratory, including bulk and spatial methods. We then review some of the current key ST literature categorized by tumor type, with an emphasis on novel studies that directly analyzed original human samples. We conclude with a discussion on the future directions for the integration of ST into the clinical care of patients with cancer.
Tumor sequencing in the clinic
Precision oncology
Precision medicine, sometimes called personalized medicine, refers to the selection of specific therapeutics for a patient based upon the unique characteristics of that individual or the disease being treated (Jain 2002). Precision oncology is the application of this concept to cancer treatment and most commonly refers to the use of molecular tumor profiling to guide the selection of cancer therapies (Yates et al. 2018). This approach to cancer therapy has led to the rapid development of targeted therapies for most solid tumors, many of which now define the standard-of-care options. Examples include cancers such as malignant melanoma with BRAF variants, ovarian carcinomas with BRCA alterations and non-small cell lung carcinoma, where the list of targeted treatments is ever expanding but already includes EGFR, ALK, ROS-1, NTRK, MET, RET, BRAF, and KRAS (Imyanitov et al. 2021).
The advent of precision oncology has largely come about due to the increasing accessibility of comprehensive genomic profiling (CGP), which in turn has driven drug development. However, significant disparities remain globally for patients with cancer both in access to testing and affordability of targeted treatments (Mateo et al. 2022). The CGP can help guide decision-making in solid tumor management; however, prior to testing, there are many practical factors for clinicians to consider.
Tumor tissue processing
Genomic sequencing of cancer starts with the critical initial step of obtaining suitable tumor tissue for testing. Often clinicians will rely on tumor tissue that has already been collected previously during the patient’s treatment journey, so-called ‘archival’ tissue. Archival tissue can be obtained from formalin-fixed paraffin-embedded (FFPE) tumor blocks or unstained tissue on microscope slides prepared at the time of previous surgical resection or from previous tumor biopsies. The reliance upon FFPE tissue comes with several limitations, including degraded nucleic acids, mutational noise, and artifacts from fixation (Do and Dobrovic 2015). Many of these limitations can be overcome by obtaining ‘fresh’ biopsies. However, laboratory advances are continually being made to optimize the extraction of DNA from FFPE tissue (Inoue et al. 2021; Oba et al. 2022), as most clinical tissue samples are stored in this way.
Next, performing next-generation sequencing (NGS) of tumors requires tissues to be homogenized prior to sequencing; therefore, the result is an average read-out of the genomic material present, which may include both benign and malignant areas depending on the sample. This also results in the loss of any specific spatial tumor context.
Variants and the Molecular Tumor Board (MTB)
Finding variants in the context of clinical cancer CGP involves identifying genetic changes, such as mutations and alterations in a tumor’s DNA. The first step after sequencing the DNA involves ‘sequence alignment,’ whereby the DNA obtained from the tumor sample is aligned to the reference genome. Variant calling algorithms are subsequently used to detect differences in the form of mutations or other alterations (Feng et al. 2023). The next key step is the laboratory-clinic interface, where the detected variants are clinically interpreted to inform treatment decisions. This process is often carried out by a Molecular Tumor Board (MTB).
Tumor sequencing in the research laboratory
Cancer genomics and transcriptomics
Genomics and transcriptomics both play key roles in cancer research. Briefly, genomics refers to the entire set of genes in a tumor determined by analyzing DNA to identify variants such as mutations, deletions, or amplifications that may cause cancer or predict treatment benefit. On the other hand, transcriptomics focuses on the RNA transcripts produced by genes, providing an indication of the dynamic pattern of gene expression at a given timepoint.
Bulk methods
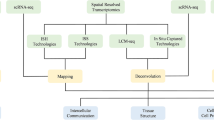
Transcriptomics utilizes various techniques to study the transcriptome, that is, the quantification of all RNA transcripts, which can be used as a surrogate for gene expression. The understanding of genomic function has increased rapidly with the widespread use of RNA-seq in molecular biology (Stark et al. 2019). Like in genomic analysis, the main limitation of this bulk transcriptomic approach is the loss of a cell type-specific understanding and spatial context of gene expression, both of which are now being explored through single-cell sequencing and ST. The complimentary nature of bulk and spatial approaches may allow integration of both methods in clinical and research settings in the near future (Fig. 1).
Single-cell sequencing
Single-cell RNA sequencing (scRNA-seq) provides insights into the heterogeneity of tissue samples and cell populations. Briefly, isolation of individual cells often involves cell sorting or microfluidic techniques. The genetic content of each cell is subsequently bound to a specific detection probe functioning as a ‘barcode’. The barcoded fragments for hundreds to tens of thousands of cells are pooled, extracted, and subsequently sequenced. Although scRNA-seq is a powerful stand-alone tool for investigating heterogeneity, it still has the limitation of tissue dissociation and therefore loss of spatial context. This issue has now been addressed with a variety of spatial methods, including ST.
Spatial transcriptomics (ST)
Overview
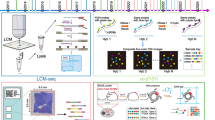
Spatial transcriptomics (ST) is the study and quantification of messenger RNA (mRNA) transcripts as a surrogate for gene expression in the spatial context of cancer cells and their associated microenvironment (Marx 2021; Moses and Pachter 2022). Multiple ST technologies have been developed, all of which result in the generation of large volumes of sequencing data for each specimen that is analyzed. Recently, advances in technology have dramatically improved the speed and quality of data acquisition and processing, which has given rise to extremely detailed resolution. Several reviews of the investigative methods employed for ST have been published in the literature in recent years (Asp et al. 2020; Liao et al. 2021; Moses and Pachter 2022; Rao et al. 2021). While the catalog of proprietary systems available for spatial transcriptomic analysis is rapidly expanding, the methods employed are generally categorized by the way in which the data are obtained and can be separated broadly into imaging-based methods and sequencing-based methods (Table 1).
Imaging-based methods: ISH and ISS
Imaging-based methods can be broadly subdivided into in situ hybridization (ISH) or in situ sequencing (ISS) methods.
In situ hybridization (ISH)
Single-molecule FISH (smFISH) is a quantitative method that uses five fluorophores per DNA or RNA molecule to image probe-labeled transcripts. However, this technique is limited by the number of detectable genes due to spectral overlap (Lewis et al. 2021). This issue has been addressed and overcome by employing multiple rounds of sequential hybridization, such as sequential fluorescence in situ hybridization (seqFISH) using color barcodes (Williams et al. 2022) or multiplexed error-robust FISH (MERFISH), which employs binary barcodes (Chen et al. 2015). While barcoding and sequential hybridization have allowed scaling and multiplexing of FISH techniques, they are limited by the increased time required for imaging, as well as the relatively small area of tissue that can be imaged.
The CosMx Spatial Molecular Imager (SMI) (NanoString) resolves RNA and protein expression at single-molecule resolution. The technology involves tissue permeabilization, probe hybridization, and slide assembly insertion in the SMI. Fluorescently labeled secondary probes containing a UV cleavable linker are then added. CosMx allows simultaneous imaging and quantification of 1000 + RNA and 64 + protein targets at subcellular resolution (He et al. 2022).
In situ sequencing (ISS)
In situ sequencing (ISS) is considered an imaging-based technique, as the method also involves visualization of mRNA directly in a section of tissue or cell sample. ISS enables in situ targeted gene expression profiling, a method commercialized as Cartana and subsequently acquired by 10x Genomics (Lewis et al. 2021). In this method padlock probes are hybridized to transcripts, probe ends are ligated, and the products are amplified using rolling circle amplification (RCA). Next, fluorescent probes are hybridized to the RCA product (Williams et al. 2022). The resulting fluorescent DNAs are read using iterative imaging technologies. This modification of Cartana technology was subsequently launched as Xenium (10x Genomics) in December 2022. Xenium is a panel-based assay that can currently be run with a maximum panel of 480 gene markers, although significantly larger panels are expected with recent announcements of plans for a 5 K (5000 gene) panel.
Sequencing-based methods
Microdissected specimens
Spatial regions of interest (ROIs) can also be isolated manually from tissues using microdissection, for example, to isolate a tumor deposit from an immune cell infiltrate. Laser capture microdissection (LCM) is the most common physical microdissection technique in which selected areas are dissected using a UV laser or by fusion of tissue with a membrane via an IR laser. These selected regions are then subjected to RNA extraction.Gene expression profiling is performed with a complementary DNA (cDNA) microarray or RNA sequencing (RNA-seq) or by dissociation into single cells for single-cell RNA-seq (scRNA-seq)(Moses and Pachter 2022). While this approach can yield detailed results for the selected ROIs, the major limitation is the impracticality of dissecting multiple discrete tissue compartments across a larger tissue sample(Asp et al. 2020).
NGS with spatial barcoding
The spatial location of transcripts can also be determined by capturing them directly from tissue sections(Moses and Pachter 2022). The samples are positioned on arrayed reverse transcription primers with unique positional barcodes via a strategy first described as “spatial transcriptomics” (ST). The mRNA captured with these spatially barcoded probes is converted to cDNA and then sequenced using previously established next-generation sequencing (NGS) platforms.
The resolution of such an approach is dependent upon the size of each capture spot. The initial ST platform had a spot diameter of 100 μm with a center-to-center distance of 200 μm(Ståhl et al. 2016). The product later commercialized by 10x Genomics, called Visium, increased the resolution by decreasing the size of each capture spot to 55 μm in a hexagonal array, in which each capture spot can contain approximately 1 to 10 cells depending on the tissue analyzed. In the latest iteration, Visium HD further increased the resolution with each slide featuring 2 × 2 μm barcoded squares, with no gaps between squares. Additional barcoded products, such as Slide-seq and Slide-seqV2, have a 10 μm diameter resolution (Stickels et al. 2021). Recently, the Stereo-seq (STOmics) platform has refined the microarray pattern to a 200 nm (0.2 μm) spot resolution (Chen et al. 2022). Traditionally, analysis of transcriptomes has been dependent upon the use of fresh frozen tissue to preserve mRNA and prevent degradation. However, newer technologies can now employ additional steps to allow formalin-fixed paraffin-embedded (FFPE) tissues to be sequenced, an approach compatible with many clinical specimens that in turn creates great opportunity for translational research.
Another commercially available sequencing-based ST product is the GeoMx Digital Spatial Profiler (DSP) by NanoString, which combines immunofluorescence techniques with optical barcoding technology (Li and Wang 2021). The platform uses gene-specific probes linked to unique barcodes for spatial mRNA analysis on tissue slides. After staining with fluorescent antibodies, regions of interest (ROIs) are selected for barcode collection, library construction, and sequencing. It accommodates more samples per slide than array-based platforms and allows for customization with custom probes. Challenges include validating antibody staining for satisfactory results (Y. Wang, B. Liu et al. 2023a).
Regardless of the specific platform used, analysis of data obtained via spatially barcoded approaches requires computational reconstruction. Given that each barcoded capture spot may contain overlapping cells, the mRNA detected will include content from more than one cell type. Therefore, a process of deconvolution must be performed to infer what cell types are present (Williams et al. 2022). Deconvolution can be achieved either through utilizing publicly available data or preferably via generation of a concordant single-cell RNA sequencing dataset from the same tissue samples undergoing spatial analysis (Kleino et al. 2022; Mañanes et al. 2024).
Spatial transcriptomics in solid tumors: what we’ve learned thus far
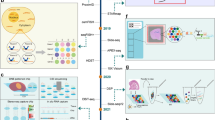
There has recently been an explosion in the use of ST for cancer research, with numerous publications across multiple cancer types. Almost all solid organ malignancies are represented in the literature. Here, we review the current key literature categorized by tumor type (Fig. 2), with an emphasis on novel studies that directly analyzed original human samples, excluding papers where the approach was purely an in silico (computational) investigation of previously available datasets (See Table 2).
Brain: Glioblastoma
Primary brain tumors, such as glioblastoma (GBM), are known to be among the most treatment-resistant solid tumors. Various spatial profiling methods have been applied in brain tumor research to investigate this highly complex, treatment-resistant TME, the early applications of which have previously been reviewed (Kalita-de Croft et al. 2021).
More recently, spatial research into the lymphoid cell population of GBM patients revealed that T-cell dysfunction was associated with an increased response to interleukin 10 (IL-10), which was spatially related to the presence of HMOX1 + myeloid cells (Ravi et al. 2022). This provided novel insights into the immune-suppressed TME of GBM. The role of immune-suppressive myeloid cells, which exhibit high expression of the epigenetic enzyme KDM6B, has also been recently reported (Goswami et al. 2023).
The integration of metabolomics and ST data revealed an association between reactive hypoxia programs and specific metabolic alterations in some regions, including enrichment of genes related to phosphoadenylate metabolism (Ravi et al. 2022b). Comparative analysis of matched primary and recurrence samples has shown that patients with GBM exhibit a shift in proneural (PN) to mesenchymal (MES) phenotypes at recurrence (Wang et al. 2022a). Additionally, matched primary-recurrence samples have also been utilized to reveal distinct tissue states; specifically, fatty acid biosynthesis enrichment in “the tissue state defined by the cohabitation of astrocyte-like/mesenchymal glioma cells, reactive astrocytes, and macrophages” is associated with recurrent GBM and shorter survival (Al-Dalahmah et al. 2023). Patient cohorts receiving neoadjuvant immunotherapy have also been analyzed, demonstrating the role of tumor-associated macrophage (TAM) subpopulations, particularly the presence of SIGLEC9 + TAMs, in immunotherapy nonresponders (Mei et al. 2023).
Head and neck cancer
Insights into the architecture of the TME are now possible with ST. Oral squamous cell carcinoma (SCC) researchers have identified unique transcriptional signatures of the cell layer at the border of the tumor (termed the leading edge) as distinct from the tumor core; additionally, unique ligand–receptor interactions have also been identified at the leading edge (Arora et al. 2023).
The model of oral SCC initiation from a precancerous lesion is well known; however, the underlying mechanisms have only recently been probed using scRNA-seq and ST. VEGFA and TGFβ signaling were found to be enriched in oral SCC initiation based on analysis of paired samples across the spectrum of normal, dysplastic, and cancerous tissues (Sun et al. 2023b).
The presence of perineural invasion (PNI) in oral SCC is a known factor increasing the risk of poor clinical outcomes. Investigators applied GeoMx DSP to eight tumor samples with PNI to show that nerves in close proximity to tumors have unique transcriptomic signatures, including changes to myelin reflecting injury and enrichment of axonogenesis and stress response genes, the gradients of which are dependent on nerve-tumor distance (Schmitd et al. 2022). The investigators proposed that nerve injury is molecular rather than physical based upon these findings.
Breast cancer
The original description of the “spatial transcriptomics” method involved analysis of the mouse brain and a human breast cancer specimen (Ståhl et al. 2016), and since this time, the use of ST in breast cancer research has accelerated rapidly. Initially, researchers focused their work on descriptive studies providing a transcriptional atlas of the cellular landscape of estrogen receptor (ER)-positive and triple-negative breast cancer (Wu et al. 2021), as well as HER2-positive breast cancer (Andersson et al. 2021). However, recently, more specific lines of investigation have been pursued.
The immune microenvironment of breast cancer has been a research area of great interest. Mao and colleagues performed an integrated analysis of patient blood, lymph node, tumor and adjacent normal tissue to illustrate the environment-dependent changes in multiple cell types. They showed that axillary lymph nodes transform into a tumor-like state with T-cell exhaustion, as well as a decrease in B cells and neutrophils (Mao et al. 2023). Additionally, based on a laser capture microdissection approach, the immune repertoires of intraepithelial T and B cells have been shown to be consistently less diverse and more clonal than those of stromal T and B cells (Romanens et al. 2023b). A comparative analysis of samples from African American women and Caucasian women revealed race-associated differences in regions with immune-rich infiltrates (Bassiouni et al. 2023).
Cancer metabolism has also been investigated in the spatial context of breast cancer, with researchers analyzing primary tumors and paired metastatic lymph nodes showing high levels of oxidative phosphorylation, particularly those located at the tumor leading edge (Liu et al. 2023b).
Thoracic cancer
The central role of immune checkpoint inhibitors (ICIs) in lung cancer therapy has understandably led researchers to focus their attention largely on the immune system and immune microenvironment. Mechanisms of resistance are now being explored by integrating clinical patient information, such as response to ICI therapy, with ST profiles. For example, enrichment of tumor-associated macrophages (TAMs) in pretreatment samples has been shown to be associated with resistance to immunotherapy treatment in a cohort of 152 patients, with spatial analysis of a subset of these implicating CD27, ITGAM, and CCL5 upregulation as drivers (Larroquette et al. 2022). CD68 + macrophages are also enriched with PD1 + and FoxP3 + cells in refractory tumors, while the expression of the IL-2 receptor alpha (CD25) is upregulated in tumor regions of responding patients (Monkman et al. 2023).
Comparative exploration of tumor and adjacent normal lung tissue has also been leveraged to determine the antigenic landscape of so-called “cold” lung cancers, with investigators finding a relatively greater frequency of neoantigens located within HLA-I presentation hotspots in CD3 + CD8 + T-cell-excluded tumors (Kraemer et al. 2023).
Occasionally, patients can present with multiple primary lung cancers, which can be clinically difficult to distinguish from intrapulmonary metastasis. Clinical approaches for differentiating these two phenomena have previously been limited to comparing biopsy samples via histopathological assessment or matching the NGS profiles of each tumor performed as part of standard care. Recently, using scRNA-seq and ST, a newly described population of epithelial cells, termed CLDN2 + alveolar type II cells, was shown to be enriched in multiple primary lung cancers (Wang et al. 2023).
Brain metastases from non-small cell lung cancer are a devastating occurrence for patients. The brain has long been known by clinicians as a ‘sanctuary site’ for metastases and is a relatively immune-privileged organ. The functional disruption of immune regulation and fibrosis has now been illustrated using spatial analysis, with T-cell- and antibody-mediated adaptive immune responses being specifically compromised in the brain metastasis microenvironment (Zhang et al. 2022).
Upper gastrointestinal cancers: gastric, esophageal, pancreatic
Upper gastrointestinal cancers such as gastric cancer and pancreatic cancer are known for being intrinsically resistant to many therapies. The mesenchymal phenotype has been linked to treatment resistance and has been explored with ST in gastric cancer, with the discovery of gastric cancer cells in a partial epithelial–mesenchymal transition state characterized by TGF-β signaling (Jang et al. 2023).
The exact location where a tumor interacts with its surroundings, known as the “tumor-normal interface”, has become an area of great interest with the advent of ST, which is uniquely tailored to this analysis compared to single-cell approaches. Using surgically resected gastric cancer specimens, researchers were able to define an immune cell-dominated tumor-normal interface with distinct immunometabolic alterations, such as disordered arginine and proline metabolism and reprogrammed lipid synthesis (Sun et al. 2023a). ST has also been used in esophageal cancer to elucidate biomarkers of progression from squamous precancerous lesions to carcinoma, with increases in TAGLN2 and decreases in CRNN suggested as candidate indicators for risk of progression (Liu et al. 2023a).
With respect to pancreatic cancer, researchers are investigating scRNA-seq and ST data, and are comparing untreated patient samples with those from patients receiving neoadjuvant therapy and finding a neural-like progenitor malignant cell program that was enriched after neoadjuvant therapy (Hwang et al. 2022). In a similarly designed cohort of treatment-naive and chemotherapy-treated patients, investigators found enrichment of inflammatory cancer-associated fibroblasts (CAFs) that upregulate metallothioneins, coordinated expression of TIGIT in exhausted and regulatory T cells, and individual tumors harboring multiple KRAS gene variant hotspots (Cui Zhou et al. 2022). More recently, insights into the immune landscape have been described, including further reports of the exhausted T-cell phenotype and the role of immunosuppressive myeloid cells (Yousuf et al. 2023).
Lower gastrointestinal cancer: colorectal, liver metastases
Colorectal cancer (CRC) is another tumor where immunotherapy has a limited role, outside of a defined subset of patients who have deficient mismatch repair (MMR) tumors (André et al. 2020). Therefore, immune microenvironment factors have been investigated using ST to identify the mechanisms and therapeutic targets that could improve immunotherapy responsiveness. The colocalization of FAP + fibroblasts and SPP1 + macrophages in CRC was demonstrated with ST and subsequently analyzed in a patient cohort in which high FAP or SPP1 expression was correlated with decreased benefit from immunotherapy (Qi et al. 2022).
A unique clinical manifestation of colorectal cancer metastasis is its preponderance for isolated spread to the liver, for which surgical resection is often recommended. Comparative analysis of primary and liver metastatic samples has allowed detailed descriptions of their transcriptional differences, which include MCAM1 + fibroblast enrichment in liver metastatic tumors (Wang et al. 2022b) as well as the identification of distinct senescent cancer cell subtypes (Garbarino et al. 2023). The invasive edge of liver metastatic tumors has also been characterized by ST, with long-term survivors demonstrating adaptive immune cell populations that are transcriptomically enriched for type II interferon signaling and MHC class II antigen presentation (Wood et al. 2023).
Fascinating insights into the intratumour microbiota have also been made using ST in colorectal cancer, with spatial illustration that bacteria tend to populate niches that are less vascular and highly immunosuppressed, providing evidence of an organized nonrandom distribution of microbiota in these tumors (Galeano Niño et al. 2022). With increasing reports of microbiota detection in other solid tumors, this field of research is likely to accelerate in the coming years.
Urological cancer: prostate, bladder, kidney
The spatial transcriptome of prostate cancer was investigated as early as 2018 using one of the initial platforms, at the time called ‘Spatial Transcriptomics’, which had a lower resolution spot size of 100 μm (Berglund et al. 2018). This initial application of ST provided an atlas-style description of the landscape of prostate tumors with respect to different tissue components. This included stroma, intraepithelial neoplasia, immune cells, and cancer cells. The heterogeneity of metastatic prostate cancer has also been a focus of spatial research, with investigators constructing tissue microarrays of diverse anatomic sites of metastasis. They found that the vast majority of metastases examined had an immune-suppressed microenvironment, as evidenced by a relative absence of inflammatory infiltrates lacking PD-1, PD-L1 and CTLA-4 (Brady et al. 2021). This immune ‘cold’ phenotype was also observed in localized prostate cancer, where ST illustrated the presence of suppressive myeloid populations, exhausted T cells, and high stromal angiogenic activity (Hirz et al. 2023). A unique feature of ST is the ability to infer ligand‒receptor interactions in tissues, with research demonstrating that primary prostate tumors from patients with metastatic disease again exhibit T-cell dysfunction, potentially impaired by nearby regulatory T cells (Salachan et al. 2023).
In bladder cancer, ST has been used to assess differences between primary and recurrent tumor samples, with the recurrent group demonstrating higher degrees of transcriptomic heterogeneity, increased expression of immune response genes and cell adhesion (Lindskrog et al. 2023). Compared with primary tumors, recurrent bladder tumors also exhibit increased cell-to-cell communication between cancer-associated fibroblasts (CAFs) and malignant cells (Shi et al. 2023). Kidney cancer is known to be responsive to immunotherapy, and spatial analysis has revealed that the underlying mechanisms may include myofibroblastic cancer-associated fibroblasts (myCAFs) (Davidson et al. 2023). Elucidating the immune phenotype of high-risk clear cell renal cell carcinoma (ccRCC) with ST has also provided insights into the immunogenic TME of tumors that have abundant exhausted/pro-tumor immune cells and provided clues to mechanisms explaining the infrequent lack of response to immunotherapy, including exhausted immune cells and lack of expression of the PD-1, PD-L1 and CTLA4 genes (Raghubar et al. 2023).
Gynecological cancer: ovarian, endometrial
Ovarian high-grade serous carcinoma most commonly presents at an advanced stage; however, research into its earlier precursor, serous tubal intraepithelial carcinoma (STIC), using spatial methods has helped identify increased protein levels of insulin-like growth factor binding protein-2 (IGFBP2) as a mechanism of proliferation (Wang et al. 2022b). High-grade serous carcinoma of the ovary is known to be an aggressive cancer, but despite this, a subset of patients can survive long-term. Spatial analysis of tumor tissue from these patients, compared to that from shorter-term survivors, revealed enrichment of immune cell types infiltrating the tumors of long-term survivors (Ferri-Borgogno et al. 2023).
In endometrial cancer, ST has been integrated into an early-phase clinical trial of a Netrin-1 antibody (NP137), in which biopsy specimens were collected and analyzed to show a treatment-related reduction in epithelial–mesenchymal transition (EMT), allowing simultaneous insights into clinical outcomes and pharmacological mechanisms of action in the spatial context (Cassier et al. 2023).
Skin cancer: melanoma, cutaneous SCC
Melanoma researchers applied ST methods very early, with initial exploratory studies of melanoma lymph node metastases showing heterogeneity in intra- and intertumoral gene expression (Thrane et al. 2018).
Melanoma brain metastases are a frequent cause of mortality in patients with advanced melanoma. ST combined with scRNA-seq has shown clusters of lymphoid aggregates dominated by plasma cells and, interestingly, has shown spatially differential expression of signatures relating to oxidative phosphorylation and glycolysis. These findings highlight the heterogeneity of these metabolic programs within the metastatic microenvironment (Biermann et al. 2022).
Analogous to cutaneous melanoma, in squamous cell carcinoma of the skin localized disease can be resected and more advanced disease is treated with immunotherapy. Investigators have shown that at the tumor leading edge, CAFs and endothelial transcripts are enriched at the juncture of tumor-specific keratinocytes and adjacent stroma, and the expression of basal tumor genes is increased(Ji et al. 2020).
Discussion and future directions
The field of spatial transcriptomic cancer research is exploding, and the initial awe-inspiring images that allowed us to visualize the spatial expression of genes in human tissue are truly remarkable breakthroughs. However, we cannot simply empirically accept that “more is better” in regard to extracting information from a biological system or indeed an individual tumor. In its infancy, spatial transcriptomics was applied in a purely investigational or discovery-science approach, as would be expected with any newly developed method.
To realize the transformative possibilities of ST, we need to leverage the unique spatial insights that are being made and bring the findings to patients via a precision oncology approach.
Cancer behavior is now being investigated at the cellular level directly in the tissue context, providing insights into the microenvironment, including different interactions at the tumor leading edge compared to tumor core (Arora et al. 2023; Ji et al. 2020; Y. M. Liu et al. 2023a), which provides clues to the mechanisms of invasion and metastasis. Understanding of the stepwise progression from precancerous to cancerous lesions is also being spatially resolved, which could pave the way forward for early intervention or cancer prevention (Liu et al. 2023a). The central role of the immune microenvironment is now better understood, revealing the various mechanisms that cancer utilizes to cloak itself from recognition, as well as highlighting the exhausted T-cell phenotype and myeloid-derived suppressor populations present in many tumors (Goswami et al. 2023; Ravi et al. 2022a).
The current landscape of literature utilizing ST thus far has been predominantly descriptive in nature but has resulted in some fascinating insights into differing populations, such as long-term survivors compared with short-term survivors (Ferri-Borgogno et al. 2023). While new prognostic factors are welcomed by clinicians, the next step is to harness this knowledge of spatial cancer behavior to inform further research and eventually develop new therapies (Cassier et al. 2023).
The issue of sampling also needs to be kept in mind when putting ST insights into context. Given that multifocal tumors and metastatic deposits are heterogeneous, marrying ST with single-cell and bulk methods, along with direct testing in established research models, will better capture broader tissue heterogeneity and help translate these new findings to patients sooner.
The thoughtful design of clinical trials to include the collection of biospecimens is one factor that may allow ST to take the next step. The future of ST research will be greatly enhanced by a sharper focus on comparative analysis of samples between groups, particularly with respect to their responsiveness to treatment. Cancer treatment selection could be greatly enhanced once we are able to start answering the kind of questions that continue to plague clinicians in the clinic: why did my patient stop responding to immunotherapy? Which combination therapy is best in this situation? Should I treat earlier or later? How did my patient develop resistance to their targeted therapy?
The field is accelerating at a rapid pace, but ultimately, how will spatial transcriptomics help patients with cancer?
As platforms continue to increase their resolution and as costs decrease, this novel method will no doubt have widespread use across future cancer research.
The labor-intensive processing steps and large datasets generated from ST, however, make it unlikely that this kind of analysis could be deployed directly in a clinical pathology laboratory today. However, one potential approach would be to utilize ST on pathology samples where deeper insights are needed from a diagnostic perspective. For example, molecular diagnostics are now being integrated into cancer pathology more frequently to identify targetable pathogenic variants (Dutta et al. 2023). One potential application could be to determine the tumor’s likelihood of being immune-responsive. While this would obviously require validation in a broader population, one could envision a situation in the future where ST could be utilized to identify the subgroup of patients who benefit from immune therapy but do not have the current standard approved biomarkers, such as PD-L1, MSI, or TMB.
Near term benefit for patients from ST research is most likely to be realized through an integrated cycle of research discovery and validation in appropriate settings (Fig. 1). Discoveries made using ST that can be validated in concordant bulk analyses, relevant preclinical models, or translational datasets from clinical trials will be more likely to provide an impact for patients sooner. This process of validation may allow useful surrogates to be identified that have the potential to be integrated into current diagnostic workflows. Emerging methods such as digital pathology analysis could also be enhanced by thoughtful integration of ST data.
Although the timeline for when patients will directly benefit from ST is not entirely clear, it is however clear, that the ST discoveries will undoubtedly be a transformational launching-pad for future precision medicine research, as well as drug discovery, and drug development.
Data availability
No datasets were generated or analysed during the current study.
References
Al-Dalahmah O, Argenziano MG, Kannan A, Mahajan A, Furnari J, Paryani F, Boyett D, Save A, Humala N, Khan F, Li J, Lu H, Sun Y, Tuddenham JF, Goldberg AR, Dovas A, Banu MA, Sudhakar T, Bush E, Canoll P (2023) Re-convolving the compositional landscape of primary and recurrent glioblastoma reveals prognostic and targetable tissue states. Nat Commun 14(1):2586. https://doi.org/10.1038/s41467-023-38186-1
Andersson A, Larsson L, Stenbeck L, Salmén F, Ehinger A, Wu SZ, Al-Eryani G, Roden D, Swarbrick A, Borg Å, Frisén J, Engblom C, Lundeberg J (2021) Spatial deconvolution of HER2-positive breast cancer delineates tumor-associated cell type interactions. Nat Commun 12(1):6012. https://doi.org/10.1038/s41467-021-26271-2
André T, Shiu KK, Kim TW, Jensen BV, Jensen LH, Punt C, Smith D, Garcia-Carbonero R, Benavides M, Gibbs P, de la Fouchardiere C, Rivera F, Elez E, Bendell J, Le DT, Yoshino T, Van Cutsem E, Yang P, Farooqui MZ, Diaz H (2020) Pembrolizumab in microsatellite-instability-high Advanced Colorectal Cancer. N Engl J Med 383(23):2207–2218. https://doi.org/10.1056/NEJMoa2017699. Jr
Arora R, Cao C, Kumar M, Sinha S, Chanda A, McNeil R, Samuel D, Arora RK, Matthews TW, Chandarana S, Hart R, Dort JC, Biernaskie J, Neri P, Hyrcza MD, Bose P (2023) Spatial transcriptomics reveals distinct and conserved tumor core and edge architectures that predict survival and targeted therapy response. Nat Commun 14(1):5029. https://doi.org/10.1038/s41467-023-40271-4
Asp M, Bergenstråhle J, Lundeberg J (2020) Spatially resolved transcriptomes-Next Generation Tools for tissue exploration. BioEssays 42(10):e1900221. https://doi.org/10.1002/bies.201900221
Bassiouni R, Idowu MO, Gibbs LD, Robila V, Grizzard PJ, Webb MG, Song J, Noriega A, Craig DW, Carpten JD (2023) Spatial transcriptomic analysis of a diverse patient cohort reveals a conserved Architecture in Triple-negative breast Cancer. Cancer Res 83(1):34–48. https://doi.org/10.1158/0008-5472.Can-22-2682
Berglund E, Maaskola J, Schultz N, Friedrich S, Marklund M, Bergenstråhle J, Tarish F, Tanoglidi A, Vickovic S, Larsson L, Salmén F, Ogris C, Wallenborg K, Lagergren J, Ståhl P, Sonnhammer E, Helleday T, Lundeberg J (2018) Spatial maps of prostate cancer transcriptomes reveal an unexplored landscape of heterogeneity. Nat Commun 9(1):2419. https://doi.org/10.1038/s41467-018-04724-5
Biermann J, Melms JC, Amin AD, Wang Y, Caprio LA, Karz A, Tagore S, Barrera I, Ibarra-Arellano MA, Andreatta M, Fullerton BT, Gretarsson KH, Sahu V, Mangipudy VS, Nguyen TTT, Nair A, Rogava M, Ho P, Koch PD, Izar B (2022) Dissecting the treatment-naive ecosystem of human melanoma brain metastasis. Cell 185(14):2591–2608e2530. https://doi.org/10.1016/j.cell.2022.06.007
Brady L, Kriner M, Coleman I, Morrissey C, Roudier M, True LD, Gulati R, Plymate SR, Zhou Z, Birditt B, Meredith R, Geiss G, Hoang M, Beechem J, Nelson PS (2021) Inter- and intra-tumor heterogeneity of metastatic prostate cancer determined by digital spatial gene expression profiling. Nat Commun 12(1):1426. https://doi.org/10.1038/s41467-021-21615-4
Cassier PA, Navaridas R, Bellina M, Rama N, Ducarouge B, Hernandez-Vargas H, Delord JP, Lengrand J, Paradisi A, Fattet L, Garin G, Gheit H, Dalban C, Pastushenko I, Neves D, Jelin R, Gadot N, Braissand N, Léon S, Mehlen P (2023) Netrin-1 blockade inhibits tumour growth and EMT features in endometrial cancer. Nature 620(7973):409–416. https://doi.org/10.1038/s41586-023-06367-z
Chen KH, Boettiger AN, Moffitt JR, Wang S, Zhuang X (2015) Spatially resolved, highly multiplexed RNA profiling in single cells. Science 348(6233):aaa6090–aaa6090. https://doi.org/10.1126/science.aaa6090
Chen A, Liao S, Cheng M, Ma K, Wu L, Lai Y, Qiu X, Yang J, Xu J, Hao S, Wang X, Lu H, Chen X, Liu X, Huang X, Li Z, Hong Y, Jiang Y, Peng J, Wang J (2022) Spatiotemporal transcriptomic atlas of mouse organogenesis using DNA nanoball-patterned arrays. Cell 185(10):1777–1792e1721. https://doi.org/10.1016/j.cell.2022.04.003
Cui Zhou D, Jayasinghe RG, Chen S, Herndon JM, Iglesia MD, Navale P, Wendl MC, Caravan W, Sato K, Storrs E, Mo CK, Liu J, Southard-Smith AN, Wu Y, Deen NA, Baer N, Fulton JM, Wyczalkowski RS, Liu MA, Ding R, L (2022) Spatially restricted drivers and transitional cell populations cooperate with the microenvironment in untreated and chemo-resistant pancreatic cancer. Nat Genet 54(9):1390–1405. https://doi.org/10.1038/s41588-022-01157-1
Davidson G, Helleux A, Vano YA, Lindner V, Fattori A, Cerciat M, Elaidi RT, Verkarre V, Sun CM, Chevreau C, Bennamoun M, Lang H, Tricard T, Fridman WH, Sautes-Fridman C, Su X, Plassard D, Keime C, Thibault-Carpentier C, Malouf GG (2023) Mesenchymal-like tumor cells and myofibroblastic cancer-associated fibroblasts are associated with progression and immunotherapy response of clear-cell renal cell carcinoma. Cancer Res. https://doi.org/10.1158/0008-5472.Can-22-3034
Do H, Dobrovic A (2015) Sequence artifacts in DNA from formalin-fixed tissues: causes and strategies for minimization. Clin Chem 61(1):64–71. https://doi.org/10.1373/clinchem.2014.223040
Dutta R, Vallurupalli M, McVeigh Q, Huang FW, Rebbeck TR (2023) Understanding inequities in precision oncology diagnostics. Nat Cancer 4(6):787–794. https://doi.org/10.1038/s43018-023-00568-1
Feng B, Lai J, Fan X, Liu Y, Wang M, Wu P, Zhou Z, Yan Q, Sun L (2023) Systematic comparison of variant calling pipelines of target genome sequencing cross multiple next-generation sequencers. Front Genet 14:1293974. https://doi.org/10.3389/fgene.2023.1293974
Ferri-Borgogno S, Zhu Y, Sheng J, Burks JK, Gomez JA, Wong KK, Wong STC, Mok SC (2023) Spatial transcriptomics depict ligand-receptor cross-talk heterogeneity at the Tumor-Stroma Interface in Long-Term Ovarian Cancer survivors. Cancer Res 83(9):1503–1516. https://doi.org/10.1158/0008-5472.Can-22-1821
Galeano Niño JL, Wu H, LaCourse KD, Kempchinsky AG, Baryiames A, Barber B, Futran N, Houlton J, Sather C, Sicinska E, Taylor A, Minot SS, Johnston CD, Bullman S (2022) Effect of the intratumoral microbiota on spatial and cellular heterogeneity in cancer. Nature 611(7937):810–817. https://doi.org/10.1038/s41586-022-05435-0
Garbarino O, Lambroia L, Basso G, Marrella V, Franceschini B, Soldani C, Pasqualini F, Giuliano D, Costa G, Peano C, Barbarossa D, Annarita D, Salvati A, Terracciano L, Torzilli G, Donadon M, Faggioli F (2023) Spatial resolution of cellular senescence dynamics in human colorectal liver metastasis. Aging Cell e13853. https://doi.org/10.1111/acel.13853
Goswami S, Raychaudhuri D, Singh P, Natarajan SM, Chen Y, Poon C, Hennessey M, Tannir AJ, Zhang J, Anandhan S, Kerrigan BP, Macaluso MD, He Z, Jindal S, Lang FF, Basu S, Sharma P (2023) Myeloid-specific KDM6B inhibition sensitizes glioblastoma to PD1 blockade. Nat Cancer. https://doi.org/10.1038/s43018-023-00620-0
He S, Bhatt R, Brown C, Brown EA, Buhr DL, Chantranuvatana K, Danaher P, Dunaway D, Garrison RG, Geiss G, Gregory MT, Hoang ML, Khafizov R, Killingbeck EE, Kim D, Kim TK, Kim Y, Klock A, Korukonda M, Beechem JM (2022) High-plex imaging of RNA and proteins at subcellular resolution in fixed tissue by spatial molecular imaging. Nat Biotechnol 40(12):1794–1806. https://doi.org/10.1038/s41587-022-01483-z
Hirz T, Mei S, Sarkar H, Kfoury Y, Wu S, Verhoeven BM, Subtelny AO, Zlatev DV, Wszolek MW, Salari K, Murray E, Chen F, Macosko EZ, Wu CL, Scadden DT, Dahl DM, Baryawno N, Saylor PJ, Kharchenko PV, Sykes DB (2023) Dissecting the immune suppressive human prostate tumor microenvironment via integrated single-cell and spatial transcriptomic analyses. Nat Commun 14(1):663. https://doi.org/10.1038/s41467-023-36325-2
Hwang WL, Jagadeesh KA, Guo JA, Hoffman HI, Yadollahpour P, Reeves JW, Mohan R, Drokhlyansky E, Van Wittenberghe N, Ashenberg O, Farhi SL, Schapiro D, Divakar P, Miller E, Zollinger DR, Eng G, Schenkel JM, Su J, Shiau C, Regev A (2022) Single-nucleus and spatial transcriptome profiling of pancreatic cancer identifies multicellular dynamics associated with neoadjuvant treatment. Nat Genet 54(8):1178–1191. https://doi.org/10.1038/s41588-022-01134-8
Imyanitov EN, Iyevleva AG, Levchenko EV (2021) Molecular testing and targeted therapy for non-small cell lung cancer: current status and perspectives. Crit Rev Oncol Hematol 157:103194. https://doi.org/10.1016/j.critrevonc.2020.103194
Inoue H, Tomida S, Horiguchi S, Kato H, Matsuoka H, Sanehira E, Matsuoka M, Yanai H, Hirasawa A, Toyooka S (2021) Best practices for the extraction of genomic DNA from formalin-fixed paraffin-embedded tumor tissue for cancer genomic profiling tests. Pathol Int 71(5):360–364. https://doi.org/10.1111/pin.13086
Jain KK (2002) Personalized medicine. Curr Opin Mol Ther 4(6):548–558
Jang E, Shin MK, Kim H, Lim JY, Lee JE, Park J, Kim J, Kim H, Shin Y, Son HY, Choi YY, Hyung WJ, Noh SH, Suh JS, Sung JY, Huh YM, Cheong JH (2023) Clinical molecular subtyping reveals intrinsic mesenchymal reprogramming in gastric cancer cells. Exp Mol Med 55(5):974–986. https://doi.org/10.1038/s12276-023-00989-z
Ji AL, Rubin AJ, Thrane K, Jiang S, Reynolds DL, Meyers RM, Guo MG, George BM, Mollbrink A, Bergenstråhle J, Larsson L, Bai Y, Zhu B, Bhaduri A, Meyers JM, Rovira-Clavé X, Hollmig ST, Aasi SZ, Nolan GP, Khavari PA (2020) Multimodal Analysis of Composition and spatial Architecture in Human squamous cell carcinoma. Cell 182(2):497–514e422. https://doi.org/10.1016/j.cell.2020.05.039
Kalita-de Croft P, Sadeghi Rad H, Gasper H, O’Byrne K, Lakhani SR, Kulasinghe A (2021) Spatial profiling technologies and applications for brain cancers. Expert Rev Mol Diagn 21(3):323–332. https://doi.org/10.1080/14737159.2021.1900735
Kleino I, Frolovaitė P, Suomi T, Elo LL (2022) Computational solutions for spatial transcriptomics. Comput Struct Biotechnol J 20:4870–4884. https://doi.org/10.1016/j.csbj.2022.08.043
Kraemer AI, Chong C, Huber F, Pak H, Stevenson BJ, Müller M, Michaux J, Altimiras ER, Rusakiewicz S, Simó-Riudalbas L, Planet E, Wiznerowicz M, Dagher J, Trono D, Coukos G, Tissot S, Bassani-Sternberg M (2023) The immunopeptidome landscape associated with T cell infiltration, inflammation and immune editing in lung cancer. Nat Cancer 4(5):608–628. https://doi.org/10.1038/s43018-023-00548-5
Larroquette M, Guegan JP, Besse B, Cousin S, Brunet M, Le Moulec S, Le Loarer F, Rey C, Soria JC, Barlesi F, Bessede A, Scoazec JY, Soubeyran I, Italiano A (2022) Spatial transcriptomics of macrophage infiltration in non-small cell lung cancer reveals determinants of sensitivity and resistance to anti-PD1/PD-L1 antibodies. J Immunother Cancer 10(5). https://doi.org/10.1136/jitc-2021-003890
Lewis SM, Asselin-Labat M-L, Nguyen Q, Berthelet J, Tan X, Wimmer VC, Merino D, Rogers KL, Naik SH (2021) Spatial omics and multiplexed imaging to explore cancer biology. Nat Methods 18(9):997–1012. https://doi.org/10.1038/s41592-021-01203-6
Li X, Wang CY (2021) From bulk, single-cell to spatial RNA sequencing. Int J Oral Sci 13(1):36. https://doi.org/10.1038/s41368-021-00146-0
Liao J, Lu X, Shao X, Zhu L, Fan X (2021) Uncovering an Organ’s Molecular Architecture at single-cell resolution by spatially resolved transcriptomics. Trends Biotechnol 39(1):43–58. https://doi.org/10.1016/j.tibtech.2020.05.006
Lindskrog SV, Schmøkel SS, Nordentoft I, Lamy P, Knudsen M, Prip F, Strandgaard T, Jensen JB, Dyrskjøt L (2023) Single-nucleus and spatially resolved Intratumor Subtype heterogeneity in bladder Cancer. Eur Urol Open Sci 51:78–88. https://doi.org/10.1016/j.euros.2023.03.006
Liu X, Zhao S, Wang K, Zhou L, Jiang M, Gao Y, Yang R, Yan S, Zhang W, Lu B, Liu F, Zhao R, Liu W, Zhang Z, Liu K, Li X, Dong Z (2023a) Spatial transcriptomics analysis of esophageal squamous precancerous lesions and their progression to esophageal cancer. Nat Commun 14(1):4779. https://doi.org/10.1038/s41467-023-40343-5
Liu YM, Ge JY, Chen YF, Liu T, Chen L, Liu CC, Ma D, Chen YY, Cai YW, Xu YY, Shao ZM, Yu KD (2023b) Combined single-cell and spatial transcriptomics reveal the metabolic evolvement of breast Cancer during early dissemination. Adv Sci (Weinh) 10(6):e2205395. https://doi.org/10.1002/advs.202205395
Mañanes D, Rivero-García I, Relaño C, Torres M, Sancho D, Jimenez-Carretero D, Torroja C, Sánchez-Cabo F (2024) SpatialDDLS: an R package to deconvolute spatial transcriptomics data using neural networks. Bioinformatics 40(2). https://doi.org/10.1093/bioinformatics/btae072
Mao X, Zhou D, Lin K, Zhang B, Gao J, Ling F, Zhu L, Yu S, Chen P, Zhang C, Zhang C, Ye G, Fong S, Chen G, Luo W (2023) Single-cell and spatial transcriptome analyses revealed cell heterogeneity and immune environment alternations in metastatic axillary lymph nodes in breast cancer. Cancer Immunol Immunother 72(3):679–695. https://doi.org/10.1007/s00262-022-03278-2
Marx V (2021) Method of the year: spatially resolved transcriptomics. Nat Methods 18(1):9–14. https://doi.org/10.1038/s41592-020-01033-y
Mateo J, Steuten L, Aftimos P, André F, Davies M, Garralda E, Geissler J, Husereau D, Martinez-Lopez I, Normanno N, Reis-Filho JS, Stefani S, Thomas DM, Westphalen CB, Voest E (2022) Delivering precision oncology to patients with cancer. Nat Med 28(4):658–665. https://doi.org/10.1038/s41591-022-01717-2
Mei Y, Wang X, Zhang J, Liu D, He J, Huang C, Liao J, Wang Y, Feng Y, Li H, Liu X, Chen L, Yi W, Chen X, Bai HM, Wang X, Li Y, Wang L, Liang Z, Jia G (2023) Siglec-9 acts as an immune-checkpoint molecule on macrophages in glioblastoma, restricting T-cell priming and immunotherapy response. Nat Cancer. https://doi.org/10.1038/s43018-023-00598-9
Monkman J, Kim H, Mayer A, Mehdi A, Matigian N, Cumberbatch M, Bhagat M, Ladwa R, Mueller SN, Adams MN, O’Byrne K, Kulasinghe A (2023) Multi-omic and spatial dissection of immunotherapy response groups in non-small cell lung cancer. Immunology. https://doi.org/10.1111/imm.13646
Moses L, Pachter L (2022) Museum of spatial transcriptomics. Nat Methods 19(5):534–546. https://doi.org/10.1038/s41592-022-01409-2
Oba U, Kohashi K, Sangatsuda Y, Oda Y, Sonoda KH, Ohga S, Yoshimoto K, Arai Y, Yachida S, Shibata T, Ito T, Miura F (2022) An efficient procedure for the recovery of DNA from formalin-fixed paraffin-embedded tissue sections. Biol Methods Protoc 7(1):bpac014. https://doi.org/10.1093/biomethods/bpac014
Qi J, Sun H, Zhang Y, Wang Z, Xun Z, Li Z, Ding X, Bao R, Hong L, Jia W, Fang F, Liu H, Chen L, Zhong J, Zou D, Liu L, Han L, Ginhoux F, Liu Y, Su B (2022) Single-cell and spatial analysis reveal interaction of FAP(+) fibroblasts and SPP1(+) macrophages in colorectal cancer. Nat Commun 13(1):1742. https://doi.org/10.1038/s41467-022-29366-6
Raghubar AM, Matigian NA, Crawford J, Francis L, Ellis R, Healy HG, Kassianos AJ, Ng MSY, Roberts MJ, Wood S, Mallett AJ (2023) High risk clear cell renal cell carcinoma microenvironments contain protumour immunophenotypes lacking specific immune checkpoints. NPJ Precis Oncol 7(1):88. https://doi.org/10.1038/s41698-023-00441-5
Rao A, Barkley D, França GS, Yanai I (2021) Exploring tissue architecture using spatial transcriptomics. Nature 596(7871):211–220. https://doi.org/10.1038/s41586-021-03634-9
Ravi VM, Neidert N, Will P, Joseph K, Maier JP, Kückelhaus J, Vollmer L, Goeldner JM, Behringer SP, Scherer F, Boerries M, Follo M, Weiss T, Delev D, Kernbach J, Franco P, Schallner N, Dierks C, Carro MS, Heiland DH (2022a) T-cell dysfunction in the glioblastoma microenvironment is mediated by myeloid cells releasing interleukin-10. Nat Commun 13(1):925. https://doi.org/10.1038/s41467-022-28523-1
Ravi VM, Will P, Kueckelhaus J, Sun N, Joseph K, Salié H, Vollmer L, Kuliesiute U, von Ehr J, Benotmane JK, Neidert N, Follo M, Scherer F, Goeldner JM, Behringer SP, Franco P, Khiat M, Zhang J, Hofmann UG, Heiland DH (2022b) Spatially resolved multi-omics deciphers bidirectional tumor-host interdependence in glioblastoma. Cancer Cell 40(6):639–655e613. https://doi.org/10.1016/j.ccell.2022.05.009
Romanens L, Chaskar P, Marcone R, Ryser S, Tille JC, Genolet R, Heimgartner-Hu K, Heimgartner K, Moore JS, Liaudet N, Kaya G, Pittet MJ, Dietrich PY, Delorenzi M, Speiser DE, Harari A, Tsantoulis P, Labidi-Galy SI (2023a) Clonal expansion of intra-epithelial T cells in breast cancer revealed by spatial transcriptomics. Int J Cancer. https://doi.org/10.1002/ijc.34620
Romanens L, Chaskar P, Marcone R, Ryser S, Tille JC, Genolet R, Heimgartner-Hu K, Heimgartner K, Moore JS, Liaudet N, Kaya G, Pittet MJ, Dietrich PY, Delorenzi M, Speiser DE, Harari A, Tsantoulis P, Labidi-Galy SI (2023b) Clonal expansion of intra-epithelial T cells in breast cancer revealed by spatial transcriptomics. Int J Cancer 153(9):1568–1578. https://doi.org/10.1002/ijc.34620
Salachan PV, Rasmussen M, Ulhøi BP, Jensen JB, Borre M, Sørensen KD (2023) Spatial whole transcriptome profiling of primary tumor from patients with metastatic prostate cancer. Int J Cancer. https://doi.org/10.1002/ijc.34708
Schmitd LB, Perez-Pacheco C, Bellile EL, Wu W, Casper K, Mierzwa M, Rozek LS, Wolf GT, Taylor JMG, D’Silva NJ (2022) Spatial and transcriptomic analysis of Perineural Invasion in oral Cancer. Clin Cancer Res 28(16):3557–3572. https://doi.org/10.1158/1078-0432.Ccr-21-4543
Shi ZD, Sun Z, Zhu ZB, Liu X, Chen JZ, Hao L, Zhu JF, Pang K, Wu D, Dong Y, Liu YF, Chen WH, Liang Q, Zhuo SC, Han CH (2023) Integrated single-cell and spatial transcriptomic profiling reveals higher intratumour heterogeneity and epithelial-fibroblast interactions in recurrent bladder cancer. Clin Transl Med 13(7):e1338. https://doi.org/10.1002/ctm2.1338
Ståhl PL, Salmén F, Vickovic S, Lundmark A, Navarro JF, Magnusson J, Giacomello S, Asp M, Westholm JO, Huss M, Mollbrink A, Linnarsson S, Codeluppi S, Borg Å, Pontén F, Costea PI, Sahlén P, Mulder J, Bergmann O, Frisén J (2016) Visualization and analysis of gene expression in tissue sections by spatial transcriptomics. Science 353(6294):78–82. https://doi.org/10.1126/science.aaf2403
Stark R, Grzelak M, Hadfield J (2019) RNA sequencing: the teenage years. Nat Rev Genet 20(11):631–656. https://doi.org/10.1038/s41576-019-0150-2
Statements & declarations
Stickels RR, Murray E, Kumar P, Li J, Marshall JL, Bella D, Arlotta DJ, Macosko P, E. Z., Chen F (2021) Highly sensitive spatial transcriptomics at near-cellular resolution with Slide-seqV2. Nat Biotechnol 39(3):313–319. https://doi.org/10.1038/s41587-020-0739-1
Sun C, Wang A, Zhou Y, Chen P, Wang X, Huang J, Gao J, Wang X, Shu L, Lu J, Dai W, Bu Z, Ji J, He J (2023a) Spatially resolved multi-omics highlights cell-specific metabolic remodeling and interactions in gastric cancer. Nat Commun 14(1):2692. https://doi.org/10.1038/s41467-023-38360-5
Sun L, Kang X, Wang C, Wang R, Yang G, Jiang W, Wu Q, Wang Y, Wu Y, Gao J, Chen L, Zhang J, Tian Z, Zhu G, Sun S (2023b) Single-cell and spatial dissection of precancerous lesions underlying the initiation process of oral squamous cell carcinoma. Cell Discov 9(1):28. https://doi.org/10.1038/s41421-023-00532-4
Thrane K, Eriksson H, Maaskola J, Hansson J, Lundeberg J (2018) Spatially resolved Transcriptomics enables dissection of genetic heterogeneity in stage III cutaneous malignant melanoma. Cancer Res 78(20):5970–5979. https://doi.org/10.1158/0008-5472.Can-18-0747
Wang L, Jung J, Babikir H, Shamardani K, Jain S, Feng X, Gupta N, Rosi S, Chang S, Raleigh D, Solomon D, Phillips JJ, Diaz AA (2022a) A single-cell atlas of glioblastoma evolution under therapy reveals cell-intrinsic and cell-extrinsic therapeutic targets. Nat Cancer 3(12):1534–1552. https://doi.org/10.1038/s43018-022-00475-x
Wang Y, Huang P, Wang BG, Murdock T, Cope L, Hsu FC, Wang TL, Shih IM (2022b) Spatial transcriptomic analysis of Ovarian Cancer precursors reveals reactivation of IGFBP2 during Pathogenesis. Cancer Res 82(24):4528–4541. https://doi.org/10.1158/0008-5472.Can-22-1620
Wang F, Long J, Li L, Wu ZX, Da TT, Wang XQ, Huang C, Jiang YH, Yao XQ, Ma HQ, Lian ZX, Zhao ZB, Cao J (2023a) Single-cell and spatial transcriptome analysis reveals the cellular heterogeneity of liver metastatic colorectal cancer. Sci Adv 9(24):eadf5464. https://doi.org/10.1126/sciadv.adf5464
Wang Y, Chen D, Liu Y, Shi D, Duan C, Li J, Shi X, Zhang Y, Yu Z, Sun N, Wang W, Ma Y, Xu X, Otkur W, Liu X, Xia T, Qi H, Piao HL, Liu HX (2023b) Multidirectional characterization of cellular composition and spatial architecture in human multiple primary lung cancers. Cell Death Dis 14(7):462. https://doi.org/10.1038/s41419-023-05992-w
Wang Y, Liu B, Zhao G, Lee Y, Buzdin A, Mu X, Zhao J, Chen H, Li X (2023c) Spatial transcriptomics: technologies, applications and experimental considerations. Genomics 115(5):110671. https://doi.org/10.1016/j.ygeno.2023.110671
Williams CG, Lee HJ, Asatsuma T, Vento-Tormo R, Haque A (2022) An introduction to spatial transcriptomics for biomedical research. Genome Med 14(1). https://doi.org/10.1186/s13073-022-01075-1
Wood CS, Pennel KAF, Leslie H, Legrini A, Cameron AJ, Melissourgou-Syka L, Quinn JA, van Wyk HC, Hay J, Roseweir AK, Nixon C, Roxburgh CSD, McMillan DC, Biankin AV, Sansom OJ, Horgan PG, Edwards J, Steele CW, Jamieson NB (2023) Spatially resolved transcriptomics deconvolutes prognostic histological subgroups in patients with Colorectal Cancer and Synchronous Liver metastases. Cancer Res 83(8):1329–1344. https://doi.org/10.1158/0008-5472.Can-22-2794
Wu SZ, Al-Eryani G, Roden DL, Junankar S, Harvey K, Andersson A, Thennavan A, Wang C, Torpy JR, Bartonicek N, Wang T, Larsson L, Kaczorowski D, Weisenfeld NI, Uytingco CR, Chew JG, Bent ZW, Chan CL, Gnanasambandapillai V, Swarbrick A (2021) A single-cell and spatially resolved atlas of human breast cancers. Nat Genet 53(9):1334–1347. https://doi.org/10.1038/s41588-021-00911-1
Yates LR, Seoane J, Le Tourneau C, Siu LL, Marais R, Michiels S, Soria JC, Campbell P, Normanno N, Scarpa A, Reis-Filho JS, Rodon J, Swanton C, Andre F (2018) The European Society for Medical Oncology (ESMO) Precision Medicine Glossary. Ann Oncol 29(1):30–35. https://doi.org/10.1093/annonc/mdx707
Yousuf S, Qiu M, Voithenberg LVV, Hulkkonen J, Macinkovic I, Schulz AR, Hartmann D, Mueller F, Mijatovic M, Ibberson D, AlHalabi KT, Hetzer J, Anders S, Brüne B, Mei HE, Imbusch CD, Brors B, Heikenwälder M, Gaida MM, Roth S (2023) Spatially resolved multi-omics single-cell analyses inform mechanisms of immune-dysfunction in pancreatic cancer. Gastroenterology. https://doi.org/10.1053/j.gastro.2023.05.036
Zhang Q, Abdo R, Iosef C, Kaneko T, Cecchini M, Han VK, Li SS (2022) The spatial transcriptomic landscape of non-small cell lung cancer brain metastasis. Nat Commun 13(1):5983. https://doi.org/10.1038/s41467-022-33365-y
Acknowledgements
Michael A Cilento is supported by a Research Training Program Scholarship from The University of Adelaide/Australian Government. Figures created with BioRender.com.
Funding
No specific funding was associated with this manuscript.
Open Access funding enabled and organized by CAUL and its Member Institutions
Author information
Authors and Affiliations
Contributions
All the authors participated in the conceptualization and design: MC, CS, and LB. The first draft of the manuscript was written by MC. Figures were created by MC. All the authors critically revised the manuscript: MC, CS, and LB. All the authors read and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
Michael A Cilento: declares no potential competing interests. Christopher J Sweeney: Consulting or Advisory Role: Sanofi, Janssen, Astellas Pharma, Bayer, Genentech, Pfizer, Lilly, MDS, Point Biopharma; Research Funding: Janssen Biotech (Inst), Astellas Pharma (Inst), Sanofi (Inst), Bayer (Inst), Patents, Royalties, Other Intellectual Property: Parthenolide (Indiana University): dimethylaminoparthenolide (Leuchemix); Exelixis: Abiraterone plus cabozantinib combination; FRAS1 SNP and tristetraprolin as biomarkers of lethal prostate cancer. Stock or Other Ownership: Leuchemix. Lisa M Butler: declares no potential competing interests.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cilento, M.A., Sweeney, C.J. & Butler, L.M. Spatial transcriptomics in cancer research and potential clinical impact: a narrative review. J Cancer Res Clin Oncol 150, 296 (2024). https://doi.org/10.1007/s00432-024-05816-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00432-024-05816-0