Abstract
Purpose
We aimed to analyze the expression of ZWINT, NUSAP1, DLGAP5, and PRC1 in tumor tissues and adjacent tissues with public data.
Methods
The expression patterns of four genes were detected in cancer tissues and adjacent tissues by qRT-PCR. The overall survival analysis was used to explore these genes in lung adenocarcinoma and squamous cell carcinoma patients. Knockdown assays were used to select the most suitable gene among these four genes. Cell function assays with the knockdown gene were conducted in A549 and NCL H226 cells. The role of the knockdown gene in lung cancer was dissected in a mice tumor model. Transcriptome sequencing analyses with the knockdown gene were analyzed.
Results
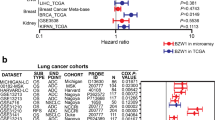
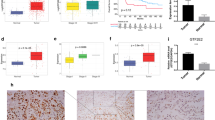
Overexpression of these genes was significantly detected in cancer tissues (P < 0.01). Overall survival revealed that high expression of these genes is closely related with poor prognosis of lung adenocarcinoma patients (P < 0.05). Knockdown of ZWINT reduced proliferation in NCI H226 and A549 cells (P < 0.05). Knockdown also inhibited cell migration, invasion, apoptosis, and colony formation (P < 0.05). ZWINT knockdown reduced tumor volume (P < 0.05). Transcriptome sequencing of ZWINT knockdown-treated A549 and NCI H226 cells indicated that 100 and 426 differentially expressed genes were obtained, respectively. Gene ontology analysis suggested that binding, biological regulation, and multicellular organismal processes were the most enriched. KEGG analysis revealed that TNF, P53, and PI3K signal networks would be the most potential ZWINT-related pathways and were identified by Western blot analysis.
Conclusions
ZWINT may be a novel target for lung cancer therapy.







Similar content being viewed by others
References
Bartkova J et al (2005) DNA damage response as a candidate anti-cancer barrier in early human tumorigenesis. Nature 434:864–870. https://doi.org/10.1038/nature03482
Bass AJ et al (2009) SOX2 is an amplified lineage-survival oncogene in lung and esophageal squamous cell carcinomas. Nat Genet 41:1238–1242. https://doi.org/10.1038/ng.465
Brendle A et al (2009) Single nucleotide polymorphisms in chromosomal instability genes and risk and clinical outcome of breast cancer: a Swedish prospective case–control study. Eur J Cancer 45:435–442. https://doi.org/10.1016/j.ejca.2008.10.001
Chen L et al (2015a) High levels of nucleolar spindle-associated protein and reduced levels of BRCA1 expression predict poor prognosis in triple-negative breast cancer. PLoS One 10:e0140572. https://doi.org/10.1371/journal.pone.0140572
Chen L, Zhuo D, Chen J, Yuan H (2015b) Screening feature genes of lung carcinoma with DNA microarray analysis. Int J Clin Exp Med 8:12161–12171
Chou HY, Wang TH, Lee SC, Hsu PH, Tsai MD, Chang CL, Jeng YM (2011) Phosphorylation of NuSAP by Cdk1 regulates its interaction with microtubules in mitosis. Cell Cycle 10:4083–4089. https://doi.org/10.4161/cc.10.23.18200
Cooper WA, Lam DC, O’Toole SA, Minna JD (2013) Molecular biology of lung cancer. J Thorac Dis 5(Suppl 5):S479–S490 https://doi.org/10.3978/j.issn.2072-1439.2013.08.03
de Bruin EC et al (2014) Spatial and temporal diversity in genomic instability processes defines lung cancer evolution. Science 346:251–256. https://doi.org/10.1126/science.1253462
Ding L et al (2008) Somatic mutations affect key pathways in lung adenocarcinoma. Nature 455:1069–1075. https://doi.org/10.1038/nature07423
Dolly SO, Collins DC, Sundar R, Popat S, Yap TA (2017) Advances in the development of molecularly targeted agents in non-small-cell lung cancer. Drugs 77:813–827. https://doi.org/10.1007/s40265-017-0732-2
Engeland K (2018) Cell cycle arrest through indirect transcriptional repression by p53: I have a DREAM. Cell Death Differ 25:114–132. https://doi.org/10.1038/cdd.2017.172
Espinosa AM et al (2013) Mitosis is a source of potential markers for screening and survival and therapeutic targets in cervical cancer. PLoS One 8:e55975. https://doi.org/10.1371/journal.pone.0055975
Gomez CR et al (2013) Prognostic value of discs large homolog 7 transcript levels in prostate cancer. PLoS One 8:e82833. https://doi.org/10.1371/journal.pone.0082833
Gordon CA, Gong X, Ganesh D, Brooks JD (2017) NUSAP1 promotes invasion and metastasis of prostate cancer. Oncotarget 8:29935–29950. https://doi.org/10.18632/oncotarget.15604
Gorgoulis VG et al (2005) Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 434:907–913. https://doi.org/10.1038/nature03485
Hanibuchi M, Kim SJ, Fidler IJ, Nishioka Y (2014) The molecular biology of lung cancer brain metastasis: an overview of current comprehensions and future perspectives. J Med Investig 61:241–253
Hassan M, Watari H, AbuAlmaaty A, Ohba Y, Sakuragi N (2014) Apoptosis and molecular targeting therapy in cancer. Biomed Res Int 2014:150845. https://doi.org/10.1155/2014/150845
Hata A et al (2013) Rebiopsy of non-small cell lung cancer patients with acquired resistance to epidermal growth factor receptor-tyrosine kinase inhibitor: comparison between T790M mutation-positive and mutation-negative populations. Cancer 119:4325–4332. https://doi.org/10.1002/cncr.28364
Hu CK, Ozlu N, Coughlin M, Steen JJ, Mitchison TJ (2012) Plk1 negatively regulates PRC1 to prevent premature midzone formation before cytokinesis. Mol Biol Cell 23:2702–2711. https://doi.org/10.1091/mbc.E12-01-0058
Huang X, Zhang X, Farahvash B, Olumi AF (2007) Novel targeted pro-apoptotic agents for the treatment of prostate cancer. J Urol 178:1846–1854. https://doi.org/10.1016/j.juro.2007.06.039
Imielinski M et al (2012) Mapping the hallmarks of lung adenocarcinoma with massively parallel sequencing. Cell 150:1107–1120. https://doi.org/10.1016/j.cell.2012.08.029
Kanehira M et al (2007) Oncogenic role of MPHOSPH1, a cancer-testis antigen specific to human bladder cancer. Cancer Res 67:3276–3285. https://doi.org/10.1158/0008-5472.can-06-3748
Kikuchi T et al (2003) Expression profiles of non-small cell lung cancers on cDNA microarrays: identification of genes for prediction of lymph-node metastasis and sensitivity to anti-cancer drugs. Oncogene 22:2192–2205. https://doi.org/10.1038/sj.onc.1206288
Kops GJ, Weaver BA, Cleveland DW (2005) On the road to cancer: aneuploidy and the mitotic checkpoint. Nat Rev Cancer 5:773–785. https://doi.org/10.1038/nrc1714
Kwak EL et al (2010) Anaplastic lymphoma kinase inhibition in non-small-cell lung cancer. N Engl J Med 363:1693–1703. https://doi.org/10.1056/NEJMoa1006448
Lin YM, Furukawa Y, Tsunoda T, Yue CT, Yang KC, Nakamura Y (2002) Molecular diagnosis of colorectal tumors by expression profiles of 50 genes expressed differentially in adenomas and carcinomas. Oncogene 21:4120–4128. https://doi.org/10.1038/sj.onc.1205518
Lin KH et al (2018) RNA-seq transcriptome analysis of breast cancer cell lines under shikonin treatment. Sci Rep 8:2672. https://doi.org/10.1038/s41598-018-21065-x
Luo HW et al (2016) Protein regulator of cytokinesis 1 overexpression predicts biochemical recurrence in men with prostate cancer. Biomed Pharmacother 78:116–120. https://doi.org/10.1016/j.biopha.2016.01.004
Maemondo M et al (2010) Gefitinib or chemotherapy for non-small-cell lung cancer with mutated EGFR N. Engl J Med 362:2380–2388. https://doi.org/10.1056/NEJMoa0909530
McGuire S (2016) World Cancer Report 2014. Geneva, Switzerland: World Health Organization, International Agency for Research on Cancer, WHO Press, 2015. Adv Nutr 7:418–419. https://doi.org/10.3945/an.116.012211
Nakamura T et al (2004) Genome-wide cDNA microarray analysis of gene expression profiles in pancreatic cancers using populations of tumor cells and normal ductal epithelial cells selected for purity by laser microdissection. Oncogene 23:2385–2400. https://doi.org/10.1038/sj.onc.1207392
Obama K, Ura K, Satoh S, Nakamura Y, Furukawa Y (2005) Up-regulation of PSF2, a member of the GINS multiprotein complex in intrahepatic cholangiocarcinoma. Oncol Rep 14:701–706
Pao W et al (2004) EGF receptor gene mutations are common in lung cancers from “never smokers” and are associated with sensitivity of tumors to gefitinib and erlotinib. Proc Natl Acad Sci USA 101:13306–13311. https://doi.org/10.1073/pnas.0405220101
Ramalingam SS, Owonikoko TK, Khuri FR (2011) Lung cancer: new biological insights and recent therapeutic advances. CA Cancer J Clin 61:91–112. https://doi.org/10.3322/caac.20102
Remon J, Moran T, Majem M, Reguart N, Dalmau E, Marquez-Medina D, Lianes P (2014) Acquired resistance to epidermal growth factor receptor tyrosine kinase inhibitors in EGFR-mutant non-small cell lung cancer: a new era begins. Cancer Treat Rev 40:93–101. https://doi.org/10.1016/j.ctrv.2013.06.002
Rooney M, Devarakonda S, Govindan R (2013) Genomics of squamous cell lung cancer Oncologist 18:707–716. https://doi.org/10.1634/theoncologist.2013-0063
Satow R et al (2010) Combined functional genome survey of therapeutic targets for hepatocellular carcinoma. Clin Cancer Res 16:2518–2528. https://doi.org/10.1158/1078-0432.ccr-09-2214
Scagliotti GV et al (2008) Phase III study comparing cisplatin plus gemcitabine with cisplatin plus pemetrexed in chemotherapy-naive patients with advanced-stage non-small-cell lung cancer. J Clin Oncol 26:3543–3551. https://doi.org/10.1200/jco.2007.15.0375
Schneider MA et al (2017) AURKA, DLGAP5, TPX2, KIF11 and CKAP5: Five specific mitosis-associated genes correlate with poor prognosis for non-small cell lung cancer patients. Int J Oncol 50:365–372. https://doi.org/10.3892/ijo.2017.3834
Sharma SV, Bell DW, Settleman J, Haber DA (2007) Epidermal growth factor receptor mutations in lung cancer. Nat Rev Cancer 7:169–181. https://doi.org/10.1038/nrc2088
Shi YX, Yin JY, Shen Y, Zhang W, Zhou HH, Liu ZQ (2017) Genome-scale analysis identifies NEK2, DLGAP5 and ECT2 as promising diagnostic and prognostic biomarkers in human lung cancer. Sci Rep 7:8072. https://doi.org/10.1038/s41598-017-08615-5
Shimo A, Nishidate T, Ohta T, Fukuda M, Nakamura Y, Katagiri T (2007) Elevated expression of protein regulator of cytokinesis 1, involved in the growth of breast cancer cells. Cancer Sci 98:174–181. https://doi.org/10.1111/j.1349-7006.2006.00381.x
Siegel R et al (2012) Cancer treatment and survivorship statistics, 2012. CA Cancer J Clin 62:220–241. https://doi.org/10.3322/caac.21149
Spigel DR et al (2011) Randomized, double-blind, placebo-controlled, phase II trial of sorafenib and erlotinib or erlotinib alone in previously treated advanced non-small-cell lung cancer. J Clin Oncol 29:2582–2589. https://doi.org/10.1200/jco.2010.30.7678
Sundar R, Soong R, Cho BC, Brahmer JR, Soo RA (2014) Immunotherapy in the treatment of non-small cell lung cancer. Lung Cancer 85:101–109. https://doi.org/10.1016/j.lungcan.2014.05.005
Wang H et al (2004) Human Zwint-1 specifies localization of Zeste White 10 to kinetochores and is essential for mitotic checkpoint signaling. J Biol Chem 279:54590–54598. https://doi.org/10.1074/jbc.M407588200
Woo Seo D, Yeop You S, Chung WJ, Cho DH, Kim JS, Su Oh J (2015) Zwint-1 is required for spindle assembly checkpoint function and kinetochore-microtubule attachment during oocyte meiosis. Sci Rep 5:15431. https://doi.org/10.1038/srep15431
Wu X et al (2017) Nucleolar and spindle associated protein 1 promotes the aggressiveness of astrocytoma by activating the Hedgehog signaling pathway. J Exp Clin Cancer Res 36:127. https://doi.org/10.1186/s13046-017-0597-y
Xie C et al (2011) KOBAS 2.0: a web server for annotation and identification of enriched pathways and diseases. Nucleic Acids Res 39:W316–W322. https://doi.org/10.1093/nar/gkr483
Xu Z, Zhou Y, Cao Y, Dinh TL, Wan J, Zhao M (2016) Identification of candidate biomarkers and analysis of prognostic values in ovarian cancer by integrated bioinformatics analysis. Med Oncol 33:130. https://doi.org/10.1007/s12032-016-0840-y
Zarogoulidis K et al (2013) Immunomodifiers in combination with conventional chemotherapy in small cell lung cancer: a phase II randomized study. Drug Des Dev Ther 7:611–617. https://doi.org/10.2147/dddt.s43184
Funding
This work was supported in part by the National Natural Science Foundation of China (no. 81602661), the Natural Science Foundation of Guangdong Province, China (no. 2016A030310164). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
FP, QL, and S-QN: bioinformatics analysis and writing of the manuscript. G-PS and YL: the discussion. MC and YB: discussion and comments on an earlier version of the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The author(s) declare that they have no competing interests.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards The Research Ethics Committee of Sun Yat-Sen University approved the collection of tissue samples for research.
Informed consent
None.
Electronic supplementary material
Below is the link to the electronic supplementary material.
432_2018_2823_MOESM1_ESM.tif
Supplementary material 1 Figure S1. mRNA abundance of ZWINT, NUSAP1, DLGAP5, and PRC1 in small cell lung cancer (SCLC), lung adenocarcinoma, and the paired adjacent normal tissues. (a) Gene expression in SCLC and the paired adjacent normal tissues. (b) Gene expression in lung adenocarcinoma and the paired adjacent normal tissues (TIF 204 KB)
Rights and permissions
About this article
Cite this article
Peng, F., Li, Q., Niu, SQ. et al. ZWINT is the next potential target for lung cancer therapy. J Cancer Res Clin Oncol 145, 661–673 (2019). https://doi.org/10.1007/s00432-018-2823-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-018-2823-1