Abstract
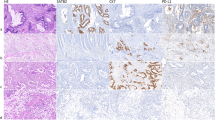
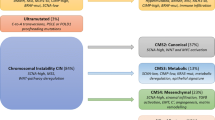
Caudal-type homeobox 2 (CDX2), special AT-rich sequence-binding protein 2 (SATB2), and keratin 20 (KRT20) are frequently used as intestinal epithelium-specific markers in immunohistochemical studies. However, subsets of colorectal carcinomas (CRCs) show loss of these markers. We analyzed The Cancer Genome Atlas data to explore molecular correlates of CDX2, SATB2, and KRT20 genes in 390 CRCs. The decreased mRNA expression of each of the three genes commonly correlated with microsatellite instability-high (MSI-H), CpG island methylator phenotype-high (CIMP-H), BRAF/RNF43 mutations, consensus molecular subtype 1, and high tumor mutational burden. The downregulation of CDX2 or SATB2 was dependent on both MSI-H and CIMP-H, whereas that of KRT20 was more dependent on MSI-H than on CIMP-H. Next, we evaluated the immunohistochemical expression of CDX2, SATB2, and KRT20 in 436 primary CRCs. In contrast to RNA-level expression, decreased expression of CDX2 and SATB2 was more dependent on CIMP-H than on MSI-H. However, consistent with RNA-level expression, decreased expression of KRT20 was more dependent on MSI-H than on CIMP-H. CIMP-H and lymphatic invasion were consistently associated with both CDX2 loss and SATB2 loss in CRCs, regardless of MSI status. In microsatellite stable CRCs, CDX2 loss correlated with BRAF mutation, whereas SATB2 loss was associated with KRAS mutations and decreased T-cell infiltration. Cases with concurrent loss of all three markers were found exclusively in MLH1-methylated MSI-H/CIMP-H CRCs. In conclusion, MSI-H and/or CIMP-H are major common correlates of decreased CDX2/SATB2/KRT20 expression in CRCs, but the specific features associated with the loss of each marker are different in CRCs.




Similar content being viewed by others
Data Availability
TCGA-COAD and TCGA-READ datasets are publicly available (https://portal.gdc.cancer.gov/). The other datasets generated and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Code availability
Not applicable.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. CA Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
van der Geest LG, Lam-Boer J, Koopman M, Verhoef C, Elferink MA, de Wilt JH (2015) Nationwide trends in incidence, treatment and survival of colorectal cancer patients with synchronous metastases. Clin Exp Metastasis 32:457–465. https://doi.org/10.1007/s10585-015-9719-0
van Gestel YR, de Hingh IH, van Herk-Sukel MP, van Erning FN, Beerepoot LV, Wijsman JH, Slooter GD, Rutten HJ, Creemers GJ, Lemmens VE (2014) Patterns of metachronous metastases after curative treatment of colorectal cancer. Cancer Epidemiol 38:448–454. https://doi.org/10.1016/j.canep.2014.04.004
Moskaluk CA, Zhang H, Powell SM, Cerilli LA, Hampton GM, Frierson HF Jr (2003) Cdx2 protein expression in normal and malignant human tissues: an immunohistochemical survey using tissue microarrays. Mod Pathol 16:913–919. https://doi.org/10.1097/01.MP.0000086073.92773.55
Werling RW, Yaziji H, Bacchi CE, Gown AM (2003) CDX2, a highly sensitive and specific marker of adenocarcinomas of intestinal origin: an immunohistochemical survey of 476 primary and metastatic carcinomas. Am J Surg Pathol 27:303–310. https://doi.org/10.1097/00000478-200303000-00003
Kaimaktchiev V, Terracciano L, Tornillo L, Spichtin H, Stoios D, Bundi M, Korcheva V, Mirlacher M, Loda M, Sauter G, Corless CL (2004) The homeobox intestinal differentiation factor CDX2 is selectively expressed in gastrointestinal adenocarcinomas. Mod Pathol 17:1392–1399. https://doi.org/10.1038/modpathol.3800205
De Lott LB, Morrison C, Suster S, Cohn DE, Frankel WL (2005) CDX2 is a useful marker of intestinal-type differentiation-a tissue microarray-based study of 629 tumors from various sites. Arch Pathol Lab Med 129:1100–1105
Dennis JL, Hvidsten TR, Wit EC, Komorowski J, Bell AK, Downie I, Mooney J, Verbeke C, Bellamy C, Keith WN, Olien KA (2005) Markers of adenocarcinoma characteristic of the site of origin: development of a diagnostic algorithm. Clin Cancer Res 11:3766–3772. https://doi.org/10.1158/1078-0432.Ccr-04-2236
Grainger S, Savory JG, Lohnes D (2010) Cdx2 regulates patterning of the intestinal epithelium. Dev Biol 339:155–165. https://doi.org/10.1016/j.ydbio.2009.12.025
Silberg DG, Swain GP, Suh ER, Traber PG (2000) Cdx1 and cdx2 expression during intestinal development. Gastroenterology 119:961–971. https://doi.org/10.1053/gast.2000.18142
Zhou Q, Toivola DM, Feng N, Greenberg HB, Franke WW, Omary MB (2003) Keratin 20 helps maintain intermediate filament organization in intestinal epithelia. Mol Biol Cell 14:2959–2971. https://doi.org/10.1091/mbc.e03-02-0059
Bae JM, Lee TH, Cho NY, Kim TY, Kang GH (2015) Loss of CDX2 expression is associated with poor prognosis in colorectal cancer patients. World J Gastroenterol 21:1457–1467. https://doi.org/10.3748/wjg.v21.i5.1457
Dalerba P, Sahoo D, Paik S, Guo XQ, Yothers G, Song N, Wilcox-Fogel N, Forgo E, Rajendran PS, Miranda SP, Hisamori S, Hutchison J, Kalisky T, Qian DL, Wolmark N, Fisher GA, van de Rijn M, Clarke MF (2016) CDX2 as a prognostic biomarker in stage II and stage III colon cancer. New Engl J Med 374:211–222. https://doi.org/10.1056/NEJMoa1506597
Hansen TF, Kjaer-Frifeldt S, Eriksen AC, Lindebjerg J, Jensen LH, Sorensen FB, Jakobsen A (2018) Prognostic impact of CDX2 in stage II colon cancer: results from two nationwide cohorts. Br J Cancer 119:1367–1373. https://doi.org/10.1038/s41416-018-0285-5
Slik K, Turkki R, Carpen O, Kurki S, Korkeila E, Sundstrom J, Pellinen T (2019) CDX2 loss with microsatellite stable phenotype predicts poor clinical outcome in stage II colorectal carcinoma. Am J Surg Pathol 43:1473–1482. https://doi.org/10.1097/PAS.0000000000001356
Graule J, Uth K, Fischer E, Centeno I, Galvan JA, Eichmann M, Rau TT, Langer R, Dawson H, Nitsche U, Traeger P, Berger MD, Schnuriger B, Hadrich M, Studer P, Inderbitzin D, Lugli A, Tschan MP, Zlobec I (2018) CDX2 in colorectal cancer is an independent prognostic factor and regulated by promoter methylation and histone deacetylation in tumors of the serrated pathway. Clin Epigenetics 10:120. https://doi.org/10.1186/s13148-018-0548-2
Kim JH, Rhee YY, Bae JM, Cho NY, Kang GH (2013) Loss of CDX2/CK20 expression is associated with poorly differentiated carcinoma, the CpG island methylator phenotype, and adverse prognosis in microsatellite-unstable colorectal cancer. Am J Surg Pathol 37:1532–1541. https://doi.org/10.1097/PAS.0b013e31829ab1c1
Berg KB, Schaeffer DF (2017) SATB2 as an immunohistochemical marker for colorectal adenocarcinoma: a concise review of benefits and pitfalls. Arch Pathol Lab Med 141:1428–1433. https://doi.org/10.5858/arpa.2016-0243-RS
Magnusson K, de Wit M, Brennan DJ, Johnson LB, McGee SF, Lundberg E, Naicker K, Klinger R, Kampf C, Asplund A, Wester K, Gry M, Bjartell A, Gallagher WM, Rexhepaj E, Kilpinen S, Kallioniemi OP, Belt E, Goos J, Meijer G, Birgisson H, Glimelius B, Borrebaeck CA, Navani S, Uhlen M, O’Connor DP, Jirstrom K, Ponten F (2011) SATB2 in combination with cytokeratin 20 identifies over 95% of all colorectal carcinomas. Am J Surg Pathol 35:937–948. https://doi.org/10.1097/PAS.0b013e31821c3dae
Dragomir A, de Wit M, Johansson C, Uhlen M, Ponten F (2014) The role of SATB2 as a diagnostic marker for tumors of colorectal origin: results of a pathology-based clinical prospective study. Am J Clin Pathol 141:630–638. https://doi.org/10.1309/AJCPWW2URZ9JKQJU
Dabir PD, Svanholm H, Christiansen JJ (2018) SATB2 is a supplementary immunohistochemical marker to CDX2 in the diagnosis of colorectal carcinoma metastasis in an unknown primary. APMIS 126:494–500. https://doi.org/10.1111/apm.12854
Eberhard J, Gaber A, Wangefjord S, Nodin B, Uhlen M, Ericson Lindquist K, Jirstrom K (2012) A cohort study of the prognostic and treatment predictive value of SATB2 expression in colorectal cancer. Br J Cancer 106:931–938. https://doi.org/10.1038/bjc.2012.34
Ma C, Olevian D, Miller C, Herbst C, Jayachandran P, Kozak MM, Chang DT, Pai RK (2019) SATB2 and CDX2 are prognostic biomarkers in DNA mismatch repair protein deficient colon cancer. Mod Pathol 32:1217–1231. https://doi.org/10.1038/s41379-019-0265-1
Ma C, Olevian DC, Lowenthal BM, Jayachandran P, Kozak MM, Chang DT, Pai RK (2018) Loss of SATB2 expression in colorectal carcinoma is associated with DNA mismatch repair protein deficiency and BRAF mutation. Am J Surg Pathol 42:1409–1417. https://doi.org/10.1097/PAS.0000000000001116
Baba Y, Nosho K, Shima K, Freed E, Irahara N, Philips J, Meyerhardt JA, Hornick JL, Shivdasani RA, Fuchs CS, Ogino S (2009) Relationship of CDX2 loss with molecular features and prognosis in colorectal cancer. Clin Cancer Res 15:4665–4673. https://doi.org/10.1158/1078-0432.CCR-09-0401
Dawson H, Koelzer VH, Lukesch AC, Mallaev M, Inderbitzin D, Lugli A, Zlobec I (2013) Loss of Cdx2 expression in primary tumors and lymph node metastases is specific for mismatch repair-deficiency in colorectal cancer. Front Oncol 3:265. https://doi.org/10.3389/fonc.2013.00265
Dawson H, Galvan JA, Helbling M, Muller DE, Karamitopoulou E, Koelzer VH, Economou M, Hammer C, Lugli A, Zlobec I (2014) Possible role of Cdx2 in the serrated pathway of colorectal cancer characterized by BRAF mutation, high-level CpG Island methylator phenotype and mismatch repair-deficiency. Int J Cancer 134:2342–2351. https://doi.org/10.1002/ijc.28564
Boland CR, Thibodeau SN, Hamilton SR, Sidransky D, Eshleman JR, Burt RW, Meltzer SJ, Rodriguez-Bigas MA, Fodde R, Ranzani GN, Srivastava S (1998) A National Cancer Institute Workshop on Microsatellite Instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res 58:5248–5257
Kim JH, Kang GH (2014) Molecular and prognostic heterogeneity of microsatellite-unstable colorectal cancer. World J Gastroenterol 20:4230–4243. https://doi.org/10.3748/wjg.v20.i15.4230
Weisenberger DJ, Siegmund KD, Campan M, Young J, Long TI, Faasse MA, Kang GH, Widschwendter M, Weener D, Buchanan D, Koh H, Simms L, Barker M, Leggett B, Levine J, Kim M, French AJ, Thibodeau SN, Jass J, Haile R, Laird PW (2006) CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet 38:787–793. https://doi.org/10.1038/ng1834
Yan HHN, Lai JCW, Ho SL, Leung WK, Law WL, Lee JFY, Chan AKW, Tsui WY, Chan ASY, Lee BCH, Yue SSK, Man AHY, Clevers H, Yuen ST, Leung SY (2017) RNF43 germline and somatic mutation in serrated neoplasia pathway and its association with BRAF mutation. Gut 66:1645–1656. https://doi.org/10.1136/gutjnl-2016-311849
Guinney J, Dienstmann R, Wang X, de Reynies A, Schlicker A, Soneson C, Marisa L, Roepman P, Nyamundanda G, Angelino P, Bot BM, Morris JS, Simon IM, Gerster S, Fessler E, De Sousa EMF, Missiaglia E, Ramay H, Barras D, Homicsko K, Maru D, Manyam GC, Broom B, Boige V, Perez-Villamil B, Laderas T, Salazar R, Gray JW, Hanahan D, Tabernero J, Bernards R, Friend SH, Laurent-Puig P, Medema JP, Sadanandam A, Wessels L, Delorenzi M, Kopetz S, Vermeulen L, Tejpar S (2015) The consensus molecular subtypes of colorectal cancer. Nat Med 21:1350–1356. https://doi.org/10.1038/nm.3967
Cancer Genome Atlas N (2012) Comprehensive molecular characterization of human colon and rectal cancer. Nature 487:330–337. https://doi.org/10.1038/nature11252
Kim JH, Kang GH (2020) Evolving pathologic concepts of serrated lesions of the colorectum. J Pathol Transl Med 54:276–289. https://doi.org/10.4132/jptm.2020.04.15
Kumar N, Tsai YH, Chen L, Zhou A, Banerjee KK, Saxena M, Huang S, Toke NH, Xing J, Shivdasani RA, Spence JR, Verzi MP (2019) The lineage-specific transcription factor CDX2 navigates dynamic chromatin to control distinct stages of intestine development. Development 146https://doi.org/10.1242/dev.172189
Brocato J, Costa M (2015) SATB1 and 2 in colorectal cancer. Carcinogenesis 36:186–191. https://doi.org/10.1093/carcin/bgu322
Ehrlich M (2019) DNA hypermethylation in disease: mechanisms and clinical relevance. Epigenetics 14:1141–1163. https://doi.org/10.1080/15592294.2019.1638701
Xu M, Xu X, Pan B, Chen X, Lin K, Zeng K, Liu X, Xu T, Sun L, Qin J, He B, Pan Y, Sun H, Wang S (2019) LncRNA SATB2-AS1 inhibits tumor metastasis and affects the tumor immune cell microenvironment in colorectal cancer by regulating SATB2. Mol Cancer 18:135. https://doi.org/10.1186/s12943-019-1063-6
Wang YQ, Jiang DM, Hu SS, Zhao L, Wang L, Yang MH, Ai ML, Jiang HJ, Han Y, Ding YQ, Wang S (2019) SATB2-AS1 suppresses colorectal carcinoma aggressiveness by inhibiting SATB2-dependent snail transcription and epithelial-mesenchymal transition. Cancer Res 79:3542–3556. https://doi.org/10.1158/0008-5472.CAN-18-2900
Chen QY, Des Marais T, Costa M (2019) Deregulation of SATB2 in carcinogenesis with emphasis on miRNA-mediated control. Carcinogenesis 40:393–402. https://doi.org/10.1093/carcin/bgz020
Verzi MP, Shin H, He HH, Sulahian R, Meyer CA, Montgomery RK, Fleet JC, Brown M, Liu XS, Shivdasani RA (2010) Differentiation-specific histone modifications reveal dynamic chromatin interactions and partners for the intestinal transcription factor CDX2. Dev Cell 19:713–726. https://doi.org/10.1016/j.devcel.2010.10.006
Kawai H, Tomii K, Toyooka S, Yano M, Murakami M, Tsukuda K, Shimizu N (2005) Promoter methylation downregulates CDX2 expression in colorectal carcinomas. Oncol Rep 13:547–551
Liao W, Overman MJ, Boutin AT, Shang X, Zhao D, Dey P, Li J, Wang G, Lan Z, Li J, Tang M, Jiang S, Ma X, Chen P, Katkhuda R, Korphaisarn K, Chakravarti D, Chang A, Spring DJ, Chang Q, Zhang J, Maru DM, Maeda DY, Zebala JA, Kopetz S, Wang YA, DePinho RA (2019) KRAS-IRF2 axis drives immune suppression and immune therapy resistance in colorectal cancer. Cancer Cell 35(559–572):e557. https://doi.org/10.1016/j.ccell.2019.02.008
Lal N, White BS, Goussous G, Pickles O, Mason MJ, Beggs AD, Taniere P, Willcox BE, Guinney J, Middleton GW (2018) KRAS mutation and consensus molecular subtypes 2 and 3 are independently associated with reduced immune infiltration and reactivity in colorectal cancer. Clin Cancer Res 24:224–233. https://doi.org/10.1158/1078-0432.CCR-17-1090
Acknowledgements
The results published here are in part based upon data generated by the TCGA Research Network: https://www.cancer.gov/tcga.
Funding
This study was supported by a grant from the Seoul National University Hospital Research Fund (04–2020-0550), a grant from the Seoul National University (800–20210387), and the National Research Foundation of Korea grants funded by the Korea government (Ministry of Science and ICT) (NRF-2016R1C1B2010627; NRF-2019R1F1A1059535).
Author information
Authors and Affiliations
Contributions
JHK designed the study. JAL, YK, KL, JHK, and GHK collected tissue samples and clinical data and performed histologic and immunohistochemical examination. JAL, M-KS, S-YY, and JHK analyzed the data. JAL and N-YC conducted the experiments. JAL, M-KS, JHK, and GHK wrote the draft. JHK and GHK revised the manuscript. All authors read and approved the final version of the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was conducted in compliance with the ethical guidelines of the 2013 Declaration of Helsinki and was approved by the Institutional Review Board of Seoul National University Hospital (IRB No. 1804–036-935).
Consent to participate and consent for publication
All tissue samples used in this study were previously registered in the Cancer Tissue Bank of Seoul National University Hospital with informed consent obtained from all patients about the research use of their tissues.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Ji Ae Lee and Mi-Kyoung Seo contributed equally to this work.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Lee, J.A., Seo, MK., Yoo, SY. et al. Comprehensive clinicopathologic, molecular, and immunologic characterization of colorectal carcinomas with loss of three intestinal markers, CDX2, SATB2, and KRT20. Virchows Arch 480, 543–555 (2022). https://doi.org/10.1007/s00428-021-03260-w
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00428-021-03260-w