Abstract
Main conclusion
Genetic loci, particularly those with an effect in the independent panel, could be utilised to further reduce LMA expression when used with favourable combinations of genes known to affect LMA.
Abstract
Late maturity α-amylase (LMA) is a grain quality defect involving elevated α-amylase within the aleurone of wheat (Triticum aestivum L.) grains. The genes known to affect expression are the reduced height genes Rht-B1 (chromosome 4B) and Rht-D1 (chromosome 4D), and an ent-copalyl diphosphate synthase gene (LMA-1) on chromosome 7B. Other minor effect loci have been reported, but these are poorly characterised and further genetic understanding is needed. In this study, twelve F4-derived populations were created through single seed descent, genotyped and evaluated for LMA. LMA-1 haplotype C and the Rht-D1b allele substantially reduced LMA expression. The alternative dwarfing genes Rht13 and Rht18 had no significant effect on LMA expression. Additional quantitative trait loci (QTL) were mapped at 16 positions in the wheat genome. Effects on LMA expression were detected for four of these QTL in a large independent panel of Australian wheat lines. The QTL detected in mapping populations and confirmed in the large independent panel provide further opportunity for selection against LMA, especially if combined with Rht-D1b and/or favourable haplotypes of LMA-1.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Late maturity α-amylase (LMA) is a grain quality defect in wheat (Triticum spp.) (Mares and Mrva 2008; Mrva et al. 2009). This defect results in the synthesis of a high pI α-amylase during the middle to late stages of grain development (Mrva and Mares 2001a). The high-pI α-amylase is not degraded and remains within the grain through to maturity. Grain parcels with high α-amylase activity are associated with a general decline in the quality of non-leavened products (Edwards et al. 1989; Kiszonas et al. 2018; Olaerts et al. 2018; Fairlie et al. 2023). These grain parcels are frequently downgraded at grain elevators, reducing revenue for wheat producers (USDA 2017). Assessment of LMA expression is too expensive for use in early generation selection. This creates a bottleneck at the end of the breeding pipeline, reducing the rate of genetic gain for other important traits. To enable breeders to select against LMA expression earlier in the breeding program, genes associated with LMA expression should be identified and markers developed for selection.
There are few known genetic mechanisms that reduce LMA expression. Expression of LMA is known to be a result of the α-amylase produced from the TaAMY1 and TaAMY4 genes (Mieog et al. 2017). The TaAMY1 gene is also expressed during grain germination and is influenced by the gibberellic acid (GA) signalling pathway (Mrva and Mares 1999; Mieog et al. 2017). Another gene associated with LMA expression was identified by Derkx et al. (2021) and designated LMA-1. The LMA-1 gene, which is on the long arm of chromosome 7B, encodes an enzyme that is responsible for converting geranylgeranyl diphosphate to ent-copalyl diphosphate at the start of the GA signalling pathway. Several haplotypes of the LMA-1 gene were associated with low LMA expression. One of these, haplotype C, is in linkage disequilibrium with a boron tolerance allele at the Bo1 boron tolerance locus (Derkx et al. 2021). LMA expression is also reduced by the Rht-B1b, Rht-D1b and Rht-B1c alleles, which have mutations that prevent the binding of GA complexes to the DELLA protein (Mrva and Mares 1996; Middleton et al. 2012). Unlike many other dwarfing mutants, DELLA mutants are not GA responsive (Ellis et al. 2004).
The non-DELLA dwarfing genes Rht13 and Rht18 act by diverting GA out of the GA signalling pathway (Rebetzke et al. 2011; Ford et al. 2018). The non-DELLA dwarfing genes are GA-responsive; application of exogenous GA increases plant height (Ellis et al. 2004). Rht13 and Rht18 are of interest to wheat breeders as alternatives to Rht-B1b and Rht-D1b for the development of semi-dwarf wheat varieties with improved early vigour (Ellis et al. 2004). However, it is not known what effect the GA-sensitive dwarfing genes have on LMA expression.
The aim of the research reported here was to investigate the effects of LMA-1 haplotype C (LMA-1-C), Rht-D1b, Rht13 and Rht18 on LMA expression, and to discover other genetic loci that could be used to select against LMA expression. This was performed in three stages, firstly, the evaluation of known genes in seven populations, secondly, quantitative trait loci (QTL) analysis in five populations and thirdly, further investigation of QTL in a panel of Australian wheat lines.
Materials and methods
Plant material
Twelve F4-derived populations, with the number of lines tested in each population in brackets (Hal/875 (80), Hal/Esp (129), Hal/Kat (79), Hal/Sce (90), Kat/Sce (112), M13/Sce (139), M18/875 (40), M18/Esp (90), M18/Hal (40), M18/Kat (40), M18/Ken (40) and M18/Sce (132); Tables 1 and 2), were produced through crossing and four generations of single seed descent. The purpose of the M13/Sce, M18/875, M18/Esp, M18/Hal, M18/Kat, M18/Ken and M18/Sce populations (Table 1) was for the evaluation of known genes, while Hal/875, Hal/Esp, Hal/Kat, Hal/Sce and Kat/Sce (Table 2) were used to map new QTL. The use of M18 (Magenta-Rht18) across several populations was to assess its effect in different genetic backgrounds, while Hal (Halberd) was used as a common parent due to its tall genotype and low LMA expression. Lines were genotyped using markers linked with Bo1 and LMA-1-C (Pallota et al. 2014), Rht-D1b (Ellis et al. 2002), Rht13 (Borrill et al. 2022) and Rht18 (Ford et al. 2018) in the populations known to be segregating for the particular gene. The genotypic data were used to select homozygous genotypes, tall genotypes in four populations (Hal/875, Hal/Esp, Hal/Kat and Hal/Sce, Table 2), lines without LMA-1-C in one population (Kat/Sce, Table 2) and to help correct for segregation distortion at the above loci in seven populations (M13/Sce, M18/875, M18/Esp, M18/Hal, M18/Kat, M18/Ken and M18/Sce, Table 1). The purpose of selection for tall genotypes or against LMA-1-C was to avoid the large effect of Rht-D1b or LMA-1-C from masking loci of small effect in the QTL analysis. Selected lines were genotyped using the custom Axiom™ Affymetrix array containing 18,101 single nucleotide polymorphism markers as described by Norman et al. (2017). Further selection against highly related lines was applied prior to the inclusion of lines in the experiment.
Plants were grown in accordance with findings reported by Derkx and Mares (2020), which indicated that a cool shock is not essential to induce LMA expression. The plants were grown in a mild, stress-free environment, under natural light with a peak photosynthetic flux measured at approximately 1200 µmol m−2 s−1. The cooling target for the evaporative cooling was set at 25 °C. The primary culm was harvested, oven-dried at 40 °C for three days and the grain threshed out using a Wintersteiger LD180 laboratory thresher (Wintersteiger AG, Ried, Austria). Grain was then stored in cool (11 °C) conditions until required in the high-pI α-amylase enzyme-linked immunosorbent assay (ELISA).
High-pI α-amylase enzyme-linked immunosorbent assay (ELISA)
Grain from each sample was milled using a Perten 3310 laboratory mill fitted with a type-2 fine disc (PerkinElmer Co., Hägersten, Sweden). Grist of each sample (200 mg ± 4 mg) was aliquoted into 96-well 2 mL titre plates for extraction, extracted using 1 mL of 0.5% NaCl and 0.02% CaCl2 extraction buffer per well and incubated for 16 h at 37 °C at 100 motions min−1. The extraction plate was then centrifuged at 2000 g for 10 min. An ELISA specific to high-pI α-amylase (SARDI, Urrbrae, SA, Australia) was then performed (Verity et al. 1999).
Genetic maps
Linkage maps were constructed for five of the populations (Hal/875, Hal/Esp, Hal/Kat, Hal/Sce and Kat/Sce, Table 2) based on genetic data for the known loci (LMA-1-C, Rht-D1b, Rht13 and Rht18) and data from the 18,101-feature single nucleotide polymorphism array. All mapping work was carried out using the R/ASMap package (Taylor and Butler 2017), which draws on functionality from R/qtl (Broman et al. 2003). Lines with missing genotype calls for more than 2000 markers were removed. For pairs or groups of lines with genetic similarity greater than 98%, marker data were merged into a single consensus genotype for linkage mapping. Monomorphic markers, markers with greater than 20% missing calls and markers that had a minor allele frequency less than 25% were removed. The remaining high-quality markers were then distributed to linkage groups associated with each chromosome using the mstmap algorithm (Wu et al. 2008). Linkage groups with fewer than eight markers were then removed. Lines that had high rates of crossovers, double crossovers or missing marker data were also removed. The markers were then remapped within their assigned linkage groups and genetic distances estimated using the R/ASMap quickEst function. Linkage groups were then compared by chromosome, with alleles switched for linkage groups that were out of phase. Linkage groups were then merged by chromosome and the markers remapped at the chromosome level and genetic distances re-estimated.
Experiment design and analysis
Experiment design
The glasshouse experiment was designed using the R/odw package (Butler and Cullis 2018) in the R statistical computing environment (R Core Team 2023). The 1024 entries included lines from each of the twelve populations, the parents of the populations and two additional control lines (RAC655 and Seri 82). A randomised complete block design consisting of three replicate blocks of 32 rows and 32 columns was used (Bailey 2008). The design was randomised according to the treatment, Line. The observational units in the glasshouse phase of the experiment were 3072 cells spread across multiple seed-raising trays, and each observational unit (OU1) could be completely indexed by Column and Row.
The ELISA phase of the experiment was designed using R/odw with a replication level of 33%. The multiphase design accounted for the design factors of the distinct glasshouse and laboratory ELISA phases of the experiment (Smith et al. 2006). The experiment was designed such that OU1’s with multiple observational units in the ELISA phase were resolvable across blocks of 21 ELISA plates (Wblock). The design was randomised according to the treatment and treatment factor, OU1 and Block. The observational units in the ELISA phase were 4032 wells spread across 42 ELISA plates, and could be completely indexed by Plate, Wcolumn and Wrow.
Data preparation and base analysis
Data checking and fitting of linear mixed models were performed in R/ASReml (Gilmour et al. 1997; Butler et al. 2018; R Core Team 2023). Given the highly skewed distribution of the optical density (OD) data, a generalised logistic transformation (logitOD) accounting for the minimum and maximum OD values was considered appropriate.
Studentised conditional residuals were then used to help identify any outliers, with a single data point removed. The design factors for the observational units of the second phase of the experiment were drawn from the first (Block, Column, Row and OU1) and second phase (Wblock, Plate, Wcolumn and Wrow) and included as random effects. Variance for Wcolumn and Wrow was investigated at the Plate level to allow for different levels of variance in each plate, given that each plate was processed individually.
Evaluation of the effects of known genes
The effect of LMA-1 (haplotypes C and F), Rht-D1 (Rht-D1a and Rht-D1b), Rht13 and Rht18 on LMA expression was assessed in each population in which the gene was segregating (Table 1). The genes were evaluated in a joint analysis across seven populations (M13/Sce, M18/875, M18/Esp, M18/Hal, M18/Kat, M18/Ken and M18/Sce) using R/ASReml (Henderson 1953; Butler et al. 2018). The genes and gene interactions included as fixed effects were LMA-1, Rht-D1, Rht13, Rht18, LMA-1:Rht-D1, LMA-1:Rht13, LMA-1:Rht18, Rht-D1:Rht13, Rht-D1:Rht18 and LMA-1:Rht-D1:Rht18. Genetic data for markers within 20 cM exclusion windows for the LMA-1, Rht-D1, Rht13 and Rht18 genes was removed prior to derivation of the k-matrix (Taylor and Verbyla 2011). The random model contained the k-matrix and Line to partition the additive and residual genetic effects, respectively (Forni et al. 2011). Design factors relevant to the first (Block, Column, Row, OU1) and second phase (Wblock, Plate, Wcolumn and Wrow) of the experiment were also included in the random model (Bailey 2008). In the final model only the significant genes LMA-1 and Rht-D1 and the significant interaction LMA-1:Rht-D1 were included, along with a k-matrix derived with 20 cM exclusion windows for LMA-1 and Rht-D1.
QTL analysis
The package R/wgaim (Taylor and Verbyla 2011) was used in conjunction with the baseline linear mixed model to detect significant QTL intervals in five populations (Hal/875, Hal/Esp, Hal/Kat, Hal/Sce and Kat/Sce, Table 2). For the baseline linear mixed model using R/ASReml, Population was included as a fixed effect. The random model contained Line (partitioned by population to allow the QTL search to be performed by population), in addition to the design factors relevant to the first (Block, Column, Row and OU1) and second phase (Wblock, Plate, Wcolumn and Wrow) of the experiment (Bailey 2008). The QTL search was performed for each population using R/wgaim, where the familywise α-level of significance was set at 0.05, the two flanking markers of the QTL are identified rather than the most likely marker, and an exclusion window of 20 cM applied to the genetic data near the flanking markers of each QTL. The exclusion window was applied after a QTL was identified and applied for the remainder of the analysis. The physical positions of the flanking markers for each QTL were obtained through a BLAST search of the Chinese Spring International Wheat Genome Sequencing Consortium RefSeq v2.1 genome assembly (Zhu et al. 2021) via https://wheat.pw.usda.gov/blast/ and are reported for ease of comparison with previously identified QTL.
Investigation of QTL in a breeding panel
A panel comprising of 614 Australian wheat varieties and breeding lines that had previously been genotyped with the 18,101-feature single nucleotide polymorphism array and assessed for LMA expression (2019–2021, best linear unbiased predictors for OD from each year) (Butler et al. 2009) using the glasshouse-based cool shock method of Mrva and Mares (2001a) was used for investigation of identified QTL. Haplotypes were assigned based on the data for the two flanking markers of each QTL interval, with the potential for two parental and two recombinant haplotypes to be assigned for each QTL. The two parental haplotypes were used for QTL investigation. An initial k-matrix was developed using markers that were not within 20 cM of any QTL interval (Taylor and Verbyla 2011). The random model contained the treatment Line, in conjunction with a k-matrix to partition the additive and residual genetic effects (Forni et al. 2011), and the treatment factor Year. The fixed model consisted of all identified QTL. QTL with no significant effect in the panel were then removed from the fixed model, replaced by the interactions of the four remaining (significant) QTL and the analysis re-run with an updated k-matrix based on markers that were not within 20 cM of the remaining QTL.
Results
Effects of known genes
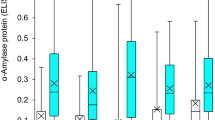
Among the lines used for evaluating the effects of known genes, LMA expression ranged from -5.5 to 2.0 logitOD. The broad-sense heritability of the analysis was 0.87 and the narrow-sense heritability 0.69, as calculated from Suppl. Table S1. The Rht-D1b allele of the reduced height gene was associated with a reduction in LMA expression of 1.86 logitOD units when compared to Rht-D1a (Table 3). LMA-1-C, which is carried by Espada, Halberd and Scepter, was associated with a reduction in LMA expression of 1.90 logitOD units, compared to LMA-1 haplotype F, which is carried by AGT Katana, Kennedy, Magenta Rht13, Magenta Rht18 and RAC875. The effects of LMA-1 and Rht-D1 were only partially additive, with the combination of Rht-D1b and LMA-1-C having a genotypic effect of 1.13 logitOD, contributing to an overall reduction in LMA expression of 2.63 logitOD units when compared to the combination of Rht-D1a and LMA-1 haplotype F. The alternative dwarfing genes Rht13 and Rht18 had no significant effect on LMA expression, either alone or in combination with LMA-1 or Rht-D1.
QTL analysis for mapping populations
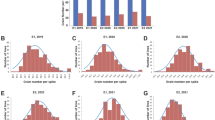
The QTL interval analysis performed in five mapping populations (Hal/875, Hal/Esp, Hal/Kat, Hal/Sce and Kat/Sce, Table 2) detected 17 QTL across 13 of the 21 wheat chromosomes (Table 4), where the two markers flanking the QTL and their RefSeq v2.1 physical position is presented. These QTL accounted for between 2.9 and 77% of the genetic variance, with absolute effects ranging from 0.22 to 1.07 logitOD units. Of the 17 QTL, only two (QLMA.agt-7B.1 in Hal/Kat and QLMA.agt-7B.2 in Hal/875) collocated with each other, near LMA-1 on chromosome 7B. Each of these 7B QTL explained more than 70% of the genetic variance, with the low LMA haplotype inherited from Halberd, which carries LMA-1-C. The 7B locus was not identified in other populations being analysed for QTL as they were homozygous for LMA-1 haplotype F (Table 2). At the QTL on other chromosomes, the low LMA haplotypes were inherited from AGT Katana (QLMA.agt-1B, QLMA.agt-3D, QLMA.agt-4B.1 and QLMA.agt-7A.1), Espada (QLMA.agt-2B), Halberd (QLMA.agt-2A.1, QLMA.agt-3A, QLMA.agt-3B.2, QLMA.agt-4B.2, QLMA.agt-5A, QLMA.agt-5B and QLMA.agt-7D), RAC875 (QLMA.agt-2A.2 QLMA.agt-2D QLMA.agt-3B.1) and Scepter (QLMA.agt-7A.2).
Investigation of QTL in a breeding panel
Data for LMA expression from the Australian wheat industry LMA screen was used to further investigate effects of the 17 QTL positions listed in Table 4. Only four of these QTL had significant effects in the panel of Australian wheat lines, as shown in Table 5. The QTL on chromosome 7B had the largest effect, with the Halberd haplotype contributing a large reduction in LMA expression (0.36 optical density, OD). QLMA.agt-7A.1 had a moderate effect, with the Halberd haplotype resulting in a slight decrease (0.25 OD) in LMA expression. QLMA.agt-3A also had a moderate effect, with the Halberd haplotype decreasing LMA expression by 0.20 OD. QLMA.agt-2B had a small effect, with the Halberd haplotype decreasing LMA expression by 0.11 OD relative to the Espada haplotype. The combination of the QLMA.agt-2B and QLMA.agt-7A.1 QTL had a small effect, with the interaction of the AGT Katana and Espada low-LMA donor haplotypes for each respective QTL reducing LMA expression by 0.24 OD relative to the Halberd haplotype. However, an overall increase in LMA expression of 0.12 OD was observed for the interaction after accounting for the individual effect of each QTL haplotype.
Discussion
The reduction in LMA expression associated with LMA-1-C and Rht-D1b (Table 3) is consistent with previous observations (Mrva and Mares 1996, 2002; Derkx et al. 2021). The ent-copalyl diphosphate synthase gene at the LMA-1 locus may affect LMA via the GA biosynthesis pathway (Derkx et al. 2021). Meanwhile, the Rht-D1b allele affects the functionality of DELLA in the GA-signalling pathway (Peng et al. 1999; Hartweck and Olszewski 2006), ultimately reducing both plant height and LMA expression (Mrva and Mares 1996; Ellis et al. 2004). In contrast, the alternative dwarfing genes Rht13 and Rht18, which divert GA out of the GA biosynthesis pathway (Rebetzke et al. 2011; Ford et al. 2018), did not reduce LMA expression despite producing a similar reduction in height to that of Rht-D1b (Ellis et al. 2004). The alternative dwarfing genes Rht13 and Rht18 have been proposed as potential replacements for Rht-B1b and Rht-D1b, which are widely used to reduce plant height, but also reduce early vigour (Ellis et al. 2004). Given that LMA expression can be reduced by Rht-B1b and Rht-D1b (Mrva and Mares 1996), wheat breeders will have to balance increasing early vigour against the risk of increasing LMA expression in deciding whether to use the Rht13 and Rht18 dwarfing genes as an alternative source of height reduction over Rht-B1b or Rht-D1b.
Several QTL were found to affect LMA expression and are of potential interest for genetic selection against LMA. However, these QTL had much smaller effect than the QTL associated with the LMA-1 gene located on chromosome 7B (Table 4). The lack of QTL in common between populations could be due to monomorphic regions, relatively small sample size or error variance. Based on its estimated physical position on chromosome 1B (335.3 Mb), QLMA.agt-1B, which had a moderate effect on LMA expression, seems to differ from the 1B/1R translocation (at 25.5 Mb) which affects falling number and α-amylase content (Farrell et al. 2013; Mohler et al. 2014). QLMA.agt-1B is also different to QTL for α-amylase content reported by Yang et al. (2014) (460.0 Mb) and Mares et al. (2023) (123.7 Mb).
QLMA.agt-2A.1 and QLMA.agt-2A.2 both have small effects on LMA expression. The estimated physical position of QLMA.agt-2A.1 (63.4 Mb) is close to the estimated physical position of one of two LMA QTL (76.1 Mb and 130.5 Mb) reported by Mares et al. (2023). QLMA.agt-2A.2 (at 771.3 Mb) is not near any previously reported QTL associated with LMA. QLMA.agt-2B has a small effect on LMA expression. At an estimated position of 769.9 Mb, it is closer to a QTL reported by Mares et al. (2023) (726.1 Mb) than to one reported by Zhang et al. (2014) (677.4 Mb). QLMA.agt-2D has a small effect on LMA expression. A QTL has previously been identified on chromosome 2D for α-amylase content (Tan et al. 2010). However, this QTL (647.1 Mb) is not close to the QLMA.agt-2D QTL (532.5 Mb).
QLMA.agt-3A has a small effect on LMA expression. QTL have previously been identified on chromosome 3A for α-amylase content (Verbyla and Cullis 2012; Liu et al. 2021; Mares et al. 2023). However, the QTL (7.5 Mb, 24.5 Mb and 654.1 Mb) are not close to QLMA.agt-3A (552.8 Mb). QLMA.agt-3B.1 and QLMA.agt-3B.2 have moderate effects on LMA expression. Their estimated physical positions are 86.6 Mb and 780.9 Mb, respectively. While QLMA.agt-3B.1 could correspond with a QTL that Mares et al. (2023) detected at 115.6 Mb, neither of these loci are near QTL reported by Tan et al. (2010) (12.5 Mb), Mares et al. (2023) (506.9 Mb) or Mrva and Mares (2001b) (516.2 Mb).
QLMA.agt-4B.1 and QLMA.agt-4B.2 have moderate effects on LMA expression. While QLMA.agt-4B.1 (22.9 Mb) could correspond with Rht-B1, which is located at 33.6 Mb, the fact that Scepter carries the Rht-B1a allele indicates that the loci are not coincident.
QLMA.agt-5A and QLMA.agt-5B both have a moderate effect on LMA expression. Based on the physical position of QLMA.agt-5A (575.6 Mb), it is not near any previously reported QTL for FN (Zhang et al. 2014; Börner et al. 2018; Martinez et al. 2018) or the TaAMY3 and TaAMY4 genes (645.7 Mb and 455.3 Mb, respectively) (Mieog et al. 2017). Meanwhile, QLMA.agt-5B (660.2 Mb) could correspond with a QTL that Tan et al. (2010) and Verbyla and Cullis (2012) detected at 623.8 Mb, or the TaAMY3 gene (Mieog et al. 2017) located at 648.6 Mb. However, none of these loci are near the TaAMY4 gene located at 420.7 Mb.
QLMA.agt-7A.1 and QLMA.agt-7A.2 both have a moderate effect on LMA expression. Their estimated physical positions are 18.2 Mb and 82.2 Mb, respectively. While QLMA.agt-7A.1 could correspond with a QTL identified by Martinez et al. (2018) (0.5 Mb), neither of these loci are close to a QTL reported by Liu et al. (2021) (602.7 Mb). QLMA.agt-7B.1 and QLMA.agt-7B.2 are collocating and have a large effect on LMA expression. The locus has previously been designated LMA-1 (Derkx et al. 2021) and has had a large effect in several studies investigating α-amylase content or FN (Mrva and Mares 2001b; Mrva et al. 2009; Emebiri et al. 2010; Mohler et al. 2014; Zhang et al. 2014; Börner et al. 2018; Martinez et al. 2018; Liu et al. 2021). QLMA.agt-7D has a small effect on LMA expression. QTL have been identified on chromosome 7D for α-amylase content (Liu et al. 2021; Mares et al. 2023). While QLMA.agt-7D (47.4 Mb) could correspond with the QTL identified by Liu et al. (2021) (56.6 Mb), neither of these loci are close to the QTL identified by Mares et al. (2023) located at 14.2 Mb.
The LMA-1 region of chromosome 7B, which has previously been reported to have a large effect on LMA expression and FN (Mrva and Mares 2001b; Mrva et al. 2009; Emebiri et al. 2010; Mohler et al. 2014; Zhang et al. 2014; Börner et al. 2018; Martinez et al. 2018; Liu et al. 2021), had consistently large effects on LMA expression in four of six populations that segregate for LMA-1-C (Tables 3 and 4) and had a large effect in the panel of Australian wheat lines (Table 5). QLMA.agt-2B, QLMA.agt-3A and QLMA.agt-7A.1 all had a moderate effect on LMA expression in mapping populations (Table 4) and were found to have a similar effect on LMA expression in an Australian wheat panel (Table 5). The combination of QLMA.agt-2B and QLMA.agt-7A.1 was found to be effective and could be pyramided to assist in reducing LMA. The four loci (QLMA.agt-2B, QLMA.agt-3A, QLMA.agt-7A.1 and QLMA.agt-7B.1) identified in Table 5 warrant further investigation to assess their potential for early selection against LMA in wheat breeding programs.
The greatest opportunities for wheat breeders to select against LMA are in marker-assisted selection, given the costly and destructive nature of LMA phenotyping which limits screening to the late stages of the breeding pipeline. Wheat breeders could also implement genomic selection provided a large enough training dataset is available. The evaluation of LMA-1-C and Rht-D1b confirms their potential for widespread implementation of marker-assisted selection against LMA expression. Unfortunately, the Rht13 and Rht18 dwarfing genes do not reduce LMA expression. This is consistent with their differential effect on the GA-signalling pathway compared to Rht-B1b and Rht-D1b. Therefore, wheat breeders will have to consider the implications on LMA expression when considering using these alternative dwarfing genes in place of Rht-B1b or Rht-D1b for height reduction. The QLMA.agt-2B, QLMA.agt-3A and QLMA.agt-7A.1 QTL detected here in mapping populations and confirmed in a large panel of Australian wheat lines provide a further opportunity for selection against LMA, especially if pyramided and/or combined with Rht-D1b and/or favourable haplotypes of LMA-1.
Data availability
The datasets generated during and/or analysed during the current study are available in the Figshare repository, [https://doi.org/10.25909/23160164].
Abbreviations
- GA:
-
Gibberellic acid
- LMA:
-
Late maturity α-amylase
- LogitOD:
-
Log-transformed optical density
- OD:
-
Optical density
- QTL:
-
Quantitative trait loci
References
Bailey RA (2008) Design of comparative experiments. Cambridge University Press, Cambridge. https://doi.org/10.1017/CBO9780511611483
Börner A, Nagel M, Agacka-Moldoch M, Gierke PU, Oberforster M, Albrecht T, Mohler V (2018) QTL analysis of falling number and seed longevity in wheat (Triticum aestivum L.). J Appl Genet 59:35–42. https://doi.org/10.1007/s13353-017-0422-5
Borrill P, Mago R, Xu T, Ford B, Williams SJ, Derkx A, Bovill WD, Hyles J, Bhatt D, Xia X, MacMillan C, White R, Buss W, Molnár I, Walkowiak S, Olsen O-A, Doležel J, Pozniak CJ, Spielmeyer W (2022) An autoactive NB-LRR gene causes Rht13 dwarfism in wheat. Proc Natl Acad Sci USA 119(48):e2209875119. https://doi.org/10.1073/pnas.2209875119
Broman KW, Wu H, Sen Ś, Churchill GA (2003) R/qtl: QTL mapping in experimental crosses. Bioinformatics 19:889–890. https://doi.org/10.1093/bioinformatics/btg112
Butler DG, Tan MK, Cullis BR (2009) Improving the accuracy of selection for late maturity α-amylase in wheat using multi-phase designs. Crop Pasture Sci 60:1202–1208. https://doi.org/10.1071/CP09124
Butler DG, Cullis BR (2018) NIASRA working paper 04–18: Optimal design under the linear mixed model
Butler DG, Cullis BR, Gilmour AR, Gogel BJ, Thompson R (2018) ASReml-R reference manual version 4. VSN International Ltd
Derkx AP, Mares DJ (2020) Late-maturity α-amylase expression in wheat is influenced by genotype, temperature and stage of grain development. Planta 251:51. https://doi.org/10.1007/s00425-020-03341-1
Derkx A, Baumann U, Cheong J, Mrva K, Sharma N, Pallota M, Mares D (2021) A major locus on wheat chromosome 7B associated with late-maturity α-amylase encodes a putative ent-copalyl diphosphate synthase. Front Plant Sci 12:637685. https://doi.org/10.3389/fpls.2021.637685
Edwards RA, Ross AS, Mares DJ, Ellison FW, Tomlinson JD (1989) Enzymes from rain-damaged and laboratory-germinated wheat I. Effects on product quality. J Cereal Sci 10(2):157–167. https://doi.org/10.1016/S0733-5210(89)80044-X
Ellis MH, Spielmeyer W, Gale KR, Rebetzke GJ, Richards RA (2002) Perfect markers for the Rht-B1b and Rht-D1b dwarfing genes in wheat. Theor Appl Genet 105:1038–1042. https://doi.org/10.1007/s00122-002-1048-4
Ellis MH, Rebetzke GJ, Chandler PM, Bonnett DG, Spielmeyer W, Richards RA (2004) The effect of different height reducing genes on the early growth of wheat. Funct Plant Biol 31:583–589. https://doi.org/10.1071/FP03207
Emebiri LC, Oliver JR, Mrva K, Mares DJ (2010) Association mapping of late maturity α-amylase (LMA) activity in a collection of synthetic hexaploid wheat. Mol Breeding 26:39–49. https://doi.org/10.1007/s11032-009-9375-7
Fairlie W, Edwards J, Mather D, Kuchel H (2023) End-use quality of wheat affected by late maturity α-amylase. J Cereal Sci 109:103610. https://doi.org/10.1016/j.jcs.2022.103610
Farrell AD, Kettlewell PS, Simmonds J, Flintham JE, Snape JW, Werner P, Jack PL (2013) Control of late maturity α-amylase in wheat by the dwarfing gene Rht-D1b and genes on the 1B/1R translocation. Mol Breeding 32:425–436. https://doi.org/10.1007/s11032-013-9881-5
Ford BA, Foo E, Sharwood R, Karafiatova M, Vrana J, MacMillan C, Nichols DS, Steuernagel B, Uauy C, Dolezel J, Chandler PM, Spielmeyer W (2018) Rht18 semidwarfism in wheat is due to increased GA2-oxidaseA9 expression and reduced GA content. Plant Physiol 177:168–180. https://doi.org/10.1104/pp.18.00023
Forni S, Aguilar I, Misztal I (2011) Different genomic relationship matrices for single-step analysis using phenotypic, pedigree and genomic information. Genet Selection Evol. https://doi.org/10.1186/1297-9686-43-1
Gilmour AR, Cullis BR, Verbyla AP (1997) Accounting for natural variation in the analysis of field experiments. J Agric Biol Environ Stat 2(3):269–293. https://doi.org/10.2307/1400446
Hartweck LM, Olszewski NE (2006) Rice GIBBERELLIN INSENSITIVE DWAR1 is a gibberellin receptor that illuminates and raises questions about GA signaling. Plant Cell 18:278–282. https://doi.org/10.1105/tpc.105.039958
Henderson CR (1953) Estimation of variance and covariance components. Biometrics 9:226–252. https://doi.org/10.2307/3001853
Kiszonas AM, Engle DA, Pierantoni LA, Morris CF (2018) Relationships between falling number, α-amylase activity, milling, cookie, and sponge cake quality of soft white wheat. Cereal Chem 95:373–385. https://doi.org/10.1002/cche.10041
Liu C, Parveen RS, Revolinski SR, Garland Campbell KA, Pumphrey MO, Steber CM (2021) The genetics of late maturity α-amylase (LMA) in North American spring wheat (Triticum aestivum L.). Seed Sci Res 31:159–168. https://doi.org/10.1017/S0960258521000064
Mares DJ, Mrva K (2008) Genetic variation for quality traits in synthetic wheat germplasm. Aust J Agric Res 59:406–412. https://doi.org/10.1071/AR07224
Mares D, Derkx A, Mather DE, Cheong J, Mrva K (2023) Multiple loci with cumulative effects on late maturity α-amylase (LMA) in wheat. Planta 257:96. https://doi.org/10.1007/s00425-023-04131-1
Martinez SA, Godoy J, Huang M, Zhang Z, Carter AH, Garland Campbell KA, Steber CM (2018) Genome-wide association mapping for tolerance to preharvest sprouting and low falling numbers in wheat. Front Plant Sci 9:141. https://doi.org/10.3389/fpls.2018.00141
Middleton AM, Ubeda-Tomas S, Griffiths J, Holman T, Hedden P, Thomas SG, Phillips AL, Holdsworth MJ, Bennett MJ, King JR, Owen MR (2012) Mathematical modeling elucidates the role of transcriptional feedback in gibberellin signaling. Proc Natl Acad Sci USA 109(19):7571–7576. https://doi.org/10.1073/pnas.1113666109
Mieog JC, Janecek S, Ral JP (2017) New insight in cereal starch degradation: identification and structural characterisation of four α-amylases in bread wheat. Amylase 1:35–49. https://doi.org/10.1515/amylase-2017-0004
Mohler V, Albrecht T, Mrva K, Schweizer G, Hartl L (2014) Genetic analysis of falling number in three bi-parental common winter wheat populations. Plant Breeding 133:448–453. https://doi.org/10.1111/pbr.12177
Mrva K, Mares DJ (1996) Expression of late maturity α-amylase in wheat containing gibberellic acid insensitivity genes. Euphytica 88(1):69–76. https://doi.org/10.1007/BF00029267
Mrva K, Mares DJ (1999) Regulation of high pI α-amylase synthesis in wheat aleurone by a gene(s) located on chromosome 6B. Euphytica 109(1):17–23. https://doi.org/10.1023/A:1003696026410
Mrva K, Mares DJ (2001a) Induction of late maturity α-amylase in wheat by cool temperature. Aust J Agric Res 52:477–484. https://doi.org/10.1071/AR00097
Mrva K, Mares DJ (2001b) Quantitative trait locus analysis of late maturity α-amylase in wheat using the doubled haploid population Cranbrook × Halberd. Aust J Agric Res 52:1267–1273. https://doi.org/10.1071/AR01047
Mrva K, Mares DJ (2002) Screening methods and identification of QTLs associated with late maturity α-amylase in wheat. Euphytica 126(1):55–59. https://doi.org/10.1023/A:1019667521448
Mrva K, Cheong J, Yu B, Law HY, Mares DJ (2009) Late maturity α-amylase in synthetic hexaploid wheat. Euphytica 168:403–411. https://doi.org/10.1007/s10681-009-9931-6
Norman A, Taylor J, Tanaka E, Telfer P, Edwards J, Martinant JP, Kuchel H (2017) Increased genomic prediction accuracy in wheat breeding using a large Australian panel. Theor Appl Genet 130:2543–2555. https://doi.org/10.1007/s00122-017-2975-4
Olaerts H, Vandekerckhove L, Courtin CM (2018) A closer look at the bread making process and the quality of bread as a function of the degree of preharvest sprouting of wheat (Triticum aestivum). J Cereal Sci 80:188–197. https://doi.org/10.1016/j.jcs.2018.03.004
Pallota M, Schnurbusch T, Hayes J, Hay A, Baumann U, Paull JG, Langridge P, Sutton T (2014) Molecular basis of adaptation to high soil boron in wheat landraces and elite cultivars. Nature 514:88–91. https://doi.org/10.1038/nature13538
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) ‘Green revolution’ genes encode mutant gibberellin response modulators. Nature 400:256–261. https://doi.org/10.1038/22307
R Core Team (2023) R: a language and environment for statistical computing. R Foundation for Statistical Computing. https://www.r-project.org. 2023
Rebetzke GJ, Ellis MH, Bonnett DG, Condon AG, Falk D, Richards RA (2011) The Rht13 dwarfing gene reduces peduncle length and plant height to increase grain number and yield of wheat. Field Crop Res 124:323–331. https://doi.org/10.1016/j.fcr.2011.06.022
Smith AB, Lim P, Cullis BR (2006) The design and analysis of multi-phase plant breeding experiments. J Agric Sci 144(5):393–409. https://doi.org/10.1017/S0021859606006319
Tan MK, Verbyla AP, Cullis BR, Martin P, Milgate AW, Oliver JR (2010) Genetics of late maturity α-amylase in a doubled haploid wheat population. Crop Pasture Sci 61(2):153–161. https://doi.org/10.1071/CP09239
Taylor J, Butler DG (2017) R package ASMap: efficient genetic linkage map construction and diagnosis. J Stat Softw 79(6):1–29. https://doi.org/10.18637/jss.v079.i06
Taylor J, Verbyla A (2011) R package wgaim: QTL analysis in bi-parental populations using linear mixed models. J Stat Softw 40(7):1–18. https://doi.org/10.18637/jss.v040.i07
USDA (2017) Commodity data specific to wheat, feed grains and oilseeds. USDA Farm Service Agency, Washington, DC
Verbyla AP, Cullis BR (2012) Multivariate whole genome average interval mapping: QTL analysis for multiple traits and/or environments. Theor Appl Genet 125(5):933–953. https://doi.org/10.1007/s00122-012-1884-9
Verity JCK, Hac L, Skerritt JH (1999) Development of a field enzyme-linked immunosorbent assay (ELISA) for detection of α-amylase in preharvest-sprouted wheat. Cereal Chem 76(5):673–681. https://doi.org/10.1094/CCHEM.1999.76.5.673
Wu Y, Bhat PR, Close TJ, Lonardi S (2008) Efficient and accurate construction of genetic linkage maps from the minimum spanning tree of a graph. PLoS Genet 4:10. https://doi.org/10.1371/journal.pgen.1000212
Yang J, Liu Y, Pu Z, Zhang L, Yuan Z, Chen G, Wei Y, Zheng Y, Liu D, Wang J (2014) Molecular characterization of high pI α-amylase and its expression QTL analysis in synthetic wheat RILs. Mol Breeding 34(3):1075–1085. https://doi.org/10.1007/s11032-014-0098-z
Zhang J, Chen J, Bowman BC, O’Brien K, Marshall JM, Bonman JM (2014) Association mapping of Hagberg falling number in hard white spring wheat. Crop Sci 54(3):1243–1252. https://doi.org/10.2135/cropsci2013.08.0551
Zhu T, Wang L, Rimbert H, Rodriguez JC, Deal KR, DeOliveira R, Choulet F, Keeble-Gagnère G, Tibbits J, Rogers J, Eversole K, Appels R, Gu YQ, Mascher M, Dvorak J, Luo M-C (2021) Optical maps refine the bread wheat Triticum aestivum cv. Chinese Spring Genome Assembly. Plant J 107:303–314. https://doi.org/10.1111/tpj.15289
Acknowledgements
The authors would like to acknowledge the plot management and laboratory support provided by the technicians at Australian Grain Technologies and the South Australian Research and Development Institute. The authors would like to thank Adinda Derkx and Daryl Mares who provided initial experimental guidance, and Alison Smith for guidance on the design and analysis of multiphase experiments.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions. WF was supported by an Australian Government Research Training Program Scholarship (F2016L01602) and a South Australian Grains Industry Trust supplementary scholarship (UA618).
Author information
Authors and Affiliations
Contributions
WF: Conceptualization, Data curation, Formal analysis, Investigation, Methodology, Project administration, Resources, Visualization, Writing – original draft. AN: Data curation, Formal analysis, Writing – review & editing. JE: Conceptualization, Funding acquisition, Methodology, Resources, Supervision, Writing – review & editing. DM: Supervision, Writing – review & editing. HK: Conceptualization, Project administration, Methodology, Resources, Supervision, Writing – review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fairlie, W., Norman, A., Edwards, J. et al. Genetic analysis of late-maturity α-amylase in twelve wheat populations. Planta 259, 40 (2024). https://doi.org/10.1007/s00425-023-04319-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-023-04319-5