Abstract
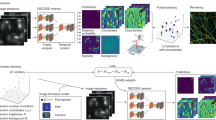
The microscope is the quintessential tool for discovery in cell biology. From its earliest incarnation as a tool to make the unseen visible, microscopes have been at the center of most revolutionizing developments in cell biology, histology and pathology. Major quantum leaps in imaging involved the dramatic improvements in resolution to see increasingly smaller structures, methods to visualize specific molecules inside of cells and tissues, and the ability to peer into living cells to study dynamics of molecules and cellular structures. The latest revolution in microscopy is Deep Imaging—the ability to look at very large numbers of samples by high-throughput microscopy at high spatial and temporal resolution. This approach is rooted in the development of fully automated high-resolution microscopes and the application of advanced computational image analysis and mining methods. Deep Imaging is enabling two novel, powerful approaches in cell biology: the ability to image thousands of samples with high optical precision allows every discernible morphological pattern to be used as a read-out in large-scale imaging-based screens, particularly in conjunction with RNAi-based screening technology; in addition, the capacity to capture large numbers of images, combined with advanced computational image analysis methods, has also opened the door to detect and analyze very rare cellular events. These two applications of Deep Imaging are revolutionizing cell biology.


Similar content being viewed by others
References
Chalfie M, Tu Y, Euskirchen G, Ward WW, Prasher DC (1994) Green fluorescent protein as a marker for gene expression. Science (New York, NY) 263:802–805
Conrad C, Gerlich DW (2010) Automated microscopy for high-content RNAi screening. J Cell Biol 188:453–461. doi:10.1083/jcb.200910105
Cremer C, Cremer T (1978) Considerations on a laser-scanning-microscope with high resolution and depth of field. Microsc Acta 81:31–44
Fong KW et al (2013) Whole-genome screening identifies proteins localized to distinct nuclear bodies. J Cell Biol 203:149–164. doi:10.1083/jcb.201303145
Hakim O et al (2012) DNA damage defines sites of recurrent chromosomal translocations in B lymphocytes. Nature 484:69–74. doi:10.1038/nature10909
Hasson SA et al (2013) High-content genome-wide RNAi screens identify regulators of parkin upstream of mitophagy. Nature 504:291–295. doi:10.1038/nature12748
Heun P, Laroche T, Shimada K, Furrer P, Gasser SM (2001) Chromosome dynamics in the yeast interphase nucleus. Science (New York, NY) 294:2181–2186. doi:10.1126/science.1065366
Krawczyk PM et al (2012) Chromatin mobility is increased at sites of DNA double-strand breaks. J Cell Sci. doi:10.1242/jcs.089847
Lazarides E, Weber K (1974) Actin antibody: the specific visualization of actin filaments in non-muscle cells. Proc Natl Acad Sci USA 71:2268–2272
Mao Z, Bozzella M, Seluanov A, Gorbunova V (2008) DNA repair by nonhomologous end joining and homologous recombination during cell cycle in human cells. Cell Cycle 7:2902–2906
Mazzarello P (1999) A unifying concept: the history of cell theory. Nat Cell Biol 1:E13–E15. doi:10.1038/8964
Meaburn KJ, Misteli T, Soutoglou E (2007) Spatial genome organization in the formation of chromosomal translocations. Semin Cancer Biol 17:80–90. doi:10.1016/j.semcancer.2006.10.008
Misteli T (2007) Beyond the sequence: cellular organization of genome function. Cell 128:787–800. doi:10.1016/j.cell.2007.01.028
Mitelman F, Johansson B, Mertens F (2007) The impact of translocations and gene fusions on cancer causation. Nat Rev Cancer 7:233–245. doi:10.1038/nrc2091
Neumann B et al (2010) Phenotypic profiling of the human genome by time-lapse microscopy reveals cell division genes. Nature 464:721–727. doi:10.1038/nature08869
Nieland TJ et al (2014) High content image analysis identifies novel regulators of synaptogenesis in a high-throughput RNAi screen of primary neurons. PloS One 9:e91744. doi:10.1371/journal.pone.0091744
Patterson G, Davidson M, Manley S, Lippincott-Schwartz J (2010) Superresolution imaging using single-molecule localization. Annu Rev Phys Chem 61:345–367. doi:10.1146/annurev.physchem.012809.103444
Pegoraro G, Voss TC, Martin SE, Tuzmen P, Guha R, Misteli T (2012) Identification of mammalian protein quality control factors by high-throughput cellular imaging. PloS One 7:e31684. doi:10.1371/journal.pone.0031684
Pepperkok R, Ellenberg J (2006) High-throughput fluorescence microscopy for systems biology. Nat Rev Mol Cell Biol 7:690–696. doi:10.1038/nrm1979
Roix JJ, McQueen PG, Munson PJ, Parada LA, Misteli T (2003) Spatial proximity of translocation-prone gene loci in human lymphomas. Nat Genet 34:287–291. doi:10.1038/ng1177
Roth J, Bendayan M, Orci L (1978) Ultrastructural localization of intracellular antigens by the use of protein A-gold complex. J Histochem Cytochem 26:1074–1081
Roukos V, Misteli T (2014) The biogenesis of chromosome translocations. Nat Cell Biol 16:293–300. doi:10.1038/ncb2941
Roukos V, Voss TC, Schmidt CK, Lee S, Wangsa D, Misteli T (2013) Spatial dynamics of chromosome translocations in living cells. Science (New York, NY) 341:660–664. doi:10.1126/science.1237150
Roukos V, Burgess RC, Misteli T (2014) Generation of cell-based systems to visualize chromosome damage and translocations in living cells. Nat Protoc (in press)
Shen J, Cookson MR (2004) Mitochondria and dopamine: new insights into recessive parkinsonism. Neuron 43:301–304. doi:10.1016/j.neuron.2004.07.012
Simpson JC, Neubrand VE, Wiemann S, Pepperkok R (2001) Illuminating the human genome. Histochem Cell Biol 115:23–29
Starkuviene V, Liebel U, Simpson JC, Erfle H, Poustka A, Wiemann S, Pepperkok R (2004) High-content screening microscopy identifies novel proteins with a putative role in secretory membrane traffic. Genome Res 14:1948–1956. doi:10.1101/gr.2658304
Symington LS, Gautier J (2011) Double-strand break end resection and repair pathway choice. Annu Rev Genet 45:247–271. doi:10.1146/annurev-genet-110410-132435
Vizeacoumar FJ et al (2010) Integrating high-throughput genetic interaction mapping and high-content screening to explore yeast spindle morphogenesis. J Cell Biol 188:69–81. doi:10.1083/jcb.200909013
Walter J, Schermelleh L, Cremer M, Tashiro S, Cremer T (2003) Chromosome order in HeLa cells changes during mitosis and early G1, but is stably maintained during subsequent interphase stages. J Cell Biol 160:685–697. doi:10.1083/jcb.200211103
Weinstock DM, Richardson CA, Elliott B, Jasin M (2006) Modeling oncogenic translocations: distinct roles for double-strand break repair pathways in translocation formation in mammalian cells. DNA Repair 5:1065–1074. doi:10.1016/j.dnarep.2006.05.028
Zhang Y et al (2012) Spatial organization of the mouse genome and its role in recurrent chromosomal translocations. Cell 148:908–921. doi:10.1016/j.cell.2012.02.002
Acknowledgments
We thank Gianluca Pegoraro and Sigal Shachar for Fig. 1. Work in the Misteli laboratory is supported by the Intramural Research Program of the National Institutes of Health (NIH), NCI, Center for Cancer Research.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Roukos, V., Misteli, T. Deep Imaging: the next frontier in microscopy. Histochem Cell Biol 142, 125–131 (2014). https://doi.org/10.1007/s00418-014-1239-5
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00418-014-1239-5