Abstract
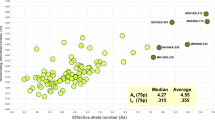
Microhaplotypes (MHs) are a promising new type of forensic markers that are defined by the combinations of two- or more single-nucleotide polymorphisms (SNPs) within 200 bp. Their advantages, such as low mutation rates, lack of stutter artifacts, and short amplicons, have improved human identification, kinship analysis, ancestry prediction, and mixture deconvolution capabilities. Information on published MHs, e.g., allele frequencies, is available in widely used public databases, ALlele FREquency Database, and MicroHapDB. However, there are abundant non-published MHs spread over the whole genome, and those databases do not incorporate other databases (e.g., the SNP Database) to provide users with more integrated information. Therefore, it is essential to establish a robust, responsive, and comprehensive MHs database. In this study, we thoroughly screened for SNP-SNP MHs among 26 populations from the 1000 Genomes Project (Phase 3). All genotype data of SNPs in each MH were converted to PHASE input files, and allele frequencies were estimated using PHASE. We compiled a detailed summary of SNP-SNPs at the global, continental, and population levels focused on haplotypes and the Ae value and supplemented our database using dbSNP data (last updated in 2015). We have successfully established a dual-SNP MH database (D-SNPsDB) of MHs within 50 bp for 26 populations in the integration of basic data such as physical positions in the human genome, mapping of variant identifiers (rsIDs), allele frequencies, and basic variant information. For public database queries, the D-SNPsDB web app was developed with the R Shiny package to get integrated information.









Similar content being viewed by others
Data availability
The SNP-SNP MH Database (D-SNPsDB) in simple text format (.txt) is available at https://1drv.ms/u/s!AnbHu1NQuWvTgQz4wN1W5HxWsgce?e=Ar0E6n. The D-SNPsDB application is available at https://1drv.ms/u/s!AnbHu1NQuWvTgRH8sKll_CCIfy8J?e=ddhhsj. The publicly available datasets analyzed in this study can be found here: http://ftp.1000genomes.ebi.ac.uk/vol1/ftp/release/20130502/ and ftp://ftp.ncbi.nih.gov/snp/organisms/human_9606/VCF/.
Code availability
The MH screening tool and R scripts used in this study are available from authors upon request.
None.
References
Butler JM (2014) Advanced Topics in Forensic DNA Typing: Interpretationhttps://doi.org/10.1016/C2011-0-07649-4
Butler JM, Buel E, Crivellente F, McCord BR (2004) Forensic DNA typing by capillary electrophoresis using the ABI Prism 310 and 3100 genetic analyzers for STR analysis. Electrophoresis 25:1397–1412. https://doi.org/10.1002/elps.200305822
Oldoni F, Podini D (2019) Forensic molecular biomarkers for mixture analysis. Forensic Sci Int Genet 41:107–119. https://doi.org/10.1016/j.fsigen.2019.04.003
Kidd K, Pakstis AJ, Speed WC et al (2013) Microhaplotype loci are a powerful new type of forensic marker. Forensic Sci Int: Genet Supp Series 4:e123–e124. https://doi.org/10.1016/j.fsigss.2013.10.063
Oldoni F, Kidd KK, Podini D (2019) Microhaplotypes in forensic genetics. Forensic Sci Int Genet 38:54–69. https://doi.org/10.1016/j.fsigen.2018.09.009
Zhu J, Lv M, Zhou N et al (2019) Genotyping polymorphic microhaplotype markers through the Illumina(®) MiSeq platform for forensics. Forensic Sci Int Genet 39:1–7. https://doi.org/10.1016/j.fsigen.2018.11.005
Oldoni F, Hart R, Long K et al (2017) Microhaplotypes for ancestry prediction. Forensic Sci Int: Genet Supp Series 6:e513–e515. https://doi.org/10.1016/j.fsigss.2017.09.209
Ruiz Y, Phillips C, Gomez-Tato A et al (2013) Further development of forensic eye color predictive tests. Forensic Sci Int Genet 7:28–40. https://doi.org/10.1016/j.fsigen.2012.05.009
Crawford NG, Kelly DE, Hansen MEB et al (2017) Loci associated with skin pigmentation identified in African populations. Science (New York NY) 358:8433. https://doi.org/10.1126/science.aan8433
Turchi C, Melchionda F, Pesaresi M, Tagliabracci A (2019) Evaluation of a microhaplotypes panel for forensic genetics using massive parallel sequencing technology. Forensic Sci Int Genet 41:120–127. https://doi.org/10.1016/j.fsigen.2019.04.009
Gandotra N, Speed WC, Qin W et al (2020) Validation of novel forensic DNA markers using multiplex microhaplotype sequencing. Forensic Sci Int Genet 47:102275. https://doi.org/10.1016/j.fsigen.2020.102275
de la Puente M, Phillips C, Xavier C et al (2020) Building a custom large-scale panel of novel microhaplotypes for forensic identification using MiSeq and Ion S5 massively parallel sequencing systems. Forensic Science International: Genetics 45https://doi.org/10.1016/j.fsigen.2019.102213
Oldoni F, Bader D, Fantinato C et al (2020) A sequence-based 74plex microhaplotype assay for analysis of forensic DNA mixtures. Forensic Sci Int Genet 49:102367. https://doi.org/10.1016/j.fsigen.2020.102367
Pang JB, Rao M, Chen QF et al (2020) A 124-plex Microhaplotype Panel Based on Next-generation Sequencing Developed for Forensic Applications. 10: 1945 https://doi.org/10.1038/s41598-020-58980-x
Pakstis AJ, Gandotra N, Speed WC, Murtha M, Scharfe C, Kidd KK (2021) The population genetics characteristics of a 90 locus panel of microhaplotypes. Hum Genet 140:1753–1773. https://doi.org/10.1007/s00439-021-02382-0
Kidd KK, Speed WC (2015) Criteria for selecting microhaplotypes: mixture detection and deconvolution. Investig Genet 6:1. https://doi.org/10.1007/s00414-020-02288-y10.1186/s13323-014-0018-3
Rajeevan H, Soundararajan U, Kidd JR, Pakstis AJ, Kidd KK (2012) ALFRED: an allele frequency resource for research and teaching. Nucleic Acids Res 40:D1010–D1015. https://doi.org/10.1093/nar/gkr924
Standage DS, Mitchell RN (2020) MicroHapDB: A Portable and Extensible Database of All Published Microhaplotype Marker and Frequency Data. Front Genet 11:781. https://doi.org/10.3389/fgene.2020.00781
Auton A, Abecasis GR, Altshuler DM et al (2015) A global reference for human genetic variation. Nature 526:68–74. https://doi.org/10.1038/nature15393
Stephens M, Donnelly P (2003) A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet 73:1162–1169. https://doi.org/10.1086/379378
Chang W, Cheng J, Allaire J, Xie Y, McPherson J (2017) Shiny: web application framework for R. R package version 1:2017
Danecek P, Auton A, Abecasis G et al (2011) The variant call format and VCFtools. Bioinformatics 27:2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Team RC (2021) R: A Language and Environment for Statistical Computing.
Hao Z, Lv D, Ge Y et al RIdeogram: drawing SVG graphics to visualize and map genome-wide data on the idiograms.
Smigielski EM, Sirotkin K, Ward M, Sherry ST (2000) dbSNP: a database of single nucleotide polymorphisms. Nucleic Acids Res 28:352–355. https://doi.org/10.1093/nar/28.1.352
Hamosh A, Scott AF, Amberger JS, Bocchini CA, McKusick VA (2005) Online Mendelian Inheritance in Man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33:D514–D517. https://doi.org/10.1093/nar/gki033
Zhang R, Tan Y, Jian H et al (2020) A new approach to detect a set of SNP-SNP markers: Combining ARMS-PCR with SNaPshot technology. 41: 1189-97https://doi.org/10.1002/elps.202000009
Stephens M, Smith NJ, Donnelly P (2001) A new statistical method for haplotype reconstruction from population data. Am J Hum Genet 68:978–989. https://doi.org/10.1086/319501
Stephens M, Scheet P (2005) Accounting for decay of linkage disequilibrium in haplotype inference and missing-data imputation. Am J Hum Genet 76:449–462. https://doi.org/10.1086/428594
Kidd KK, Pakstis AJ, Speed WC et al (2014) Current sequencing technology makes microhaplotypes a powerful new type of genetic marker for forensics. Forensic Sci Int Genet 12:215–224. https://doi.org/10.1186/s13323-014-0018-310.1016/j.fsigen.2014.06.014
Debeljak M, Mocci E, Morrison MC et al (2017) Haplotype Counting for Sensitive Chimerism Testing: Potential for Early Leukemia Relapse Detection. J Mol Diagn 19:427–436. https://doi.org/10.1016/j.jmoldx.2017.01.005
Ou X, Qu N (2020) Noninvasive prenatal paternity testing by target sequencing microhaps. Forensic Sci Int Genet 48:102338. https://doi.org/10.1016/j.fsigen.2020.102338
Ou X, Bai Z (2021) A case of heteropaternal superfecundation identified by microhap sequencing of maternal plasma cell-free DNA: A case of HS identified by microhap sequencing. Forensic Sci Int Genet 51:102458. https://doi.org/10.1016/j.fsigen.2020.102458
Jian H, Wang L, Lv M et al (2021) A Novel SNP-STR System Based on a Capillary Electrophoresis Platform. Front Genet 12:636821. https://doi.org/10.3389/fgene.2021.636821
Qu S, Lv M, Xue J et al (2020) Multi-Indel: A Microhaplotype Marker Can Be Typed Using Capillary Electrophoresis Platforms. Front Genet 11:567082. https://doi.org/10.3389/fgene.2020.567082
Acknowledgements
We are grateful to the members of our laboratory for their helpful discussions.
Funding
This study was supported by grants from National Natural Science Foundation of China (00401054A1019).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by Ethics Committee at Sichuan university.
Consent to participate
None.
Conflicts of interest/Competing interests
There were no conflicts of interest in this study.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Xue, J., Qu, S., Tan, M. et al. An overview of SNP-SNP microhaplotypes in the 26 populations of the 1000 Genomes Project. Int J Legal Med 136, 1211–1226 (2022). https://doi.org/10.1007/s00414-022-02820-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00414-022-02820-2