Abstract
Background
Healthy eating index (HEI), a measure of diet quality, associates with metabolic health outcomes; however, the molecular basis is unclear. We conducted a multi-omic study to examine whether HEI associates with the circulatory and gut metabolome and investigated the gut microbiome–HEI interaction on circulating and gut metabolites.
Methods
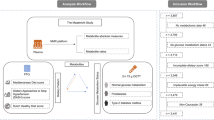
Through a cross-sectional study, we evaluated diet quality in healthy individuals [the ABO Glycoproteomics in Platelets and Endothelial Cells (ABO) Study, n = 73], metabolites (measured at Metabolon Inc.) in plasma (n = 800) and gut (n = 767) and the gut microbiome at enterotype and microbial taxa (n = 296) levels. Pathway analysis was conducted using Metaboanalyst 4.0. We performed multi-variable linear regression to explore both the HEI-metabolites and HEI-microbiome associations and how metabolites were affected by the HEI–microbiome interaction. In the Fish oils and Adipose Inflammation Reduction (FAIR) Study (n = 25), analyses on HEI and plasma metabolites were replicated. Estimates of findings from both studies were pooled in random-effects meta-analysis.
Results
The HEI-2015 was associated with 74 plasma and 73 gut metabolites (mostly lipids) and with 47 metabolites in the meta-analysis of the ABO and FAIR Studies. Compared to Enterotype-1 participants, those with Enterotype-2 had higher diet quality (p = 0.01). We also identified 9 microbial genera associated with HEI, and 35 plasma and 40 gut metabolites linked to the HEI–gut microbiome interaction. Pathways involved in the metabolism of polar lipids, amino acids and caffeine strongly associated with diet quality. However, the HEI–microbiome interaction not only influenced the pathways involved in the metabolism of branch-chain amino acids, it also affected upstream pathways including nucleotide metabolism and amino acids biosynthesis.
Conclusions
Our multi-omic analysis demonstrated that changes in metabolism, measured by either circulatory/gut metabolites or metabolic pathways, are influenced by not only diet quality but also gut microbiome alterations shaped by the quality of diet consumed. Future work is needed to explore the causality in the interplay between HEI and gut-microbiome composition in metabolism.



Similar content being viewed by others
Abbreviations
- AA:
-
Amino acid
- AHEI:
-
Alternate healthy eating index
- AMP:
-
Adenosine 5′-monophosphate
- DAG:
-
Diacylglycerol
- DASH:
-
Dietary approaches to stop hypertension
- DHQ:
-
Diet History Questionnaire
- FAIR:
-
Fish oils and Adipose Inflammation Reduction
- FFQ:
-
Food Frequency Questionnaires
- HEI:
-
Healthy Eating Index
- IQR:
-
Interquartile range
- MUFA:
-
Monounsaturated
- PAM:
-
Partitioning Around Medoids
- PUFA:
-
Polyunsaturated fatty acid
- SAH:
-
Adenosylhomocysteine
- UPenn NGSC:
-
University of Pennsylvania Next-Generation Sequencing Center
- VANTAGE:
-
Vanderbilt University Technologies for Advanced Genomics
References
Gil Á, de Victoria EM, Olza J (2015) Indicators for the evaluation of diet quality. Nutr Hosp 31(Suppl 3):128–144. https://doi.org/10.3305/nh.2015.31.sup3.8761
Wirt A, Collins CE (2009) Diet quality—what is it and does it matter? Public Health Nutr 12:2473–2492. https://doi.org/10.1017/S136898000900531X
Imamura F, Micha R, Khatibzadeh S et al (2015) Dietary quality among men and women in 187 countries in 1990 and 2010: a systematic assessment. Lancet Glob Health 3:e132–e142. https://doi.org/10.1016/S2214-109X(14)70381-X
George SM, Ballard-Barbash R, Manson JE et al (2014) Comparing indices of diet quality with chronic disease mortality risk in postmenopausal women in the Women’s Health Initiative Observational Study: evidence to inform national dietary guidance. Am J Epidemiol 180:616–625. https://doi.org/10.1093/aje/kwu173
Playdon MC, Moore SC, Derkach A et al (2017) Identifying biomarkers of dietary patterns by using metabolomics. Am J Clin Nutr 105:450–465. https://doi.org/10.3945/ajcn.116.144501
Guasch-Ferré M, Bhupathiraju SN, Hu FB (2018) Use of metabolomics in improving assessment of dietary intake. Clin Chem 64:82–98. https://doi.org/10.1373/clinchem.2017.272344
Bagheri M, Willett W, Townsend MK et al (2020) A lipid-related metabolomic pattern of diet quality. Am J Clin Nutr. https://doi.org/10.1093/ajcn/nqaa242
Tang Z-Z, Chen G, Hong Q et al (2019) Multi-omic analysis of the microbiome and metabolome in healthy subjects reveals microbiome-dependent relationships between diet and metabolites. Front Genet 10:454. https://doi.org/10.3389/fgene.2019.00454
Visconti A, Le Roy CI, Rosa F et al (2019) Interplay between the human gut microbiome and host metabolism. Nat Commun 10:4505. https://doi.org/10.1038/s41467-019-12476-z
Subramanian I, Verma S, Kumar S et al (2020) Multi-omics data integration, interpretation, and its application. Bioinform Biol Insights 14:1177932219899051–1177932219899051. https://doi.org/10.1177/1177932219899051
Hasin Y, Seldin M, Lusis A (2017) Multi-omics approaches to disease. Genome Biol 18:83. https://doi.org/10.1186/s13059-017-1215-1
Shah RD, Tang Z-Z, Chen G et al (2020) Soy food intake associates with changes in the metabolome and reduced blood pressure in a gut microbiota dependent manner. Nutr Metab Cardiovasc Dis 30:1500–1511. https://doi.org/10.1016/j.numecd.2020.05.001
Perng W, Aslibekyan S (2020) Find the needle in the haystack, then find it again: replication and validation in the ‘omics era. Metabolites 10:286. https://doi.org/10.3390/metabo10070286
Subar AF, Crafts J, Zimmerman TP et al (2010) Assessment of the accuracy of portion size reports using computer-based food photographs aids in the development of an automated self-administered 24-hour recall. J Am Diet Assoc 110:55–64. https://doi.org/10.1016/j.jada.2009.10.007
Ford L, Kennedy AD, Goodman KD et al (2020) Precision of a clinical metabolomics profiling platform for use in the identification of inborn errors of metabolism. J Appl Lab Med 5:342–356. https://doi.org/10.1093/jalm/jfz026
Wilmanski T, Rappaport N, Earls JC et al (2019) Blood metabolome predicts gut microbiome α-diversity in humans. Nat Biotechnol 37:1217–1228. https://doi.org/10.1038/s41587-019-0233-9
Borenstein M, Hedges LV, Higgins JPT, Rothstein HR (2010) A basic introduction to fixed-effect and random-effects models for meta-analysis. Res Synth Methods 1:97–111. https://doi.org/10.1002/jrsm.12
R Core Team (2017) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Hartman PE (1990) Ergothioneine as antioxidant. Methods in enzymology. Academic Press, pp 310–318
McCullough ML, Maliniak ML, Stevens VL et al (2019) Metabolomic markers of healthy dietary patterns in US postmenopausal women. Am J Clin Nutr 109:1439–1451. https://doi.org/10.1093/ajcn/nqy385
Sorrentino V, Menzies KJ, Auwerx J (2018) Repairing mitochondrial dysfunction in disease. Annu Rev Pharmacol Toxicol 58:353–389. https://doi.org/10.1146/annurev-pharmtox-010716-104908
Murphy MP (2013) Mitochondrial dysfunction indirectly elevates ROS production by the endoplasmic reticulum. Cell Metab 18:145–146. https://doi.org/10.1016/j.cmet.2013.07.006
Menni C, Fauman E, Erte I et al (2013) Biomarkers for type 2 diabetes and impaired fasting glucose using a nontargeted metabolomics approach. Diabetes 62:4270–4276. https://doi.org/10.2337/db13-0570
Wu GD, Chen J, Hoffmann C et al (2011) Linking long-term dietary patterns with gut microbial enterotypes. Science 334:105–108. https://doi.org/10.1126/science.1208344
Sandberg J, Kovatcheva-Datchary P, Björck I et al (2019) Abundance of gut Prevotella at baseline and metabolic response to barley prebiotics. Eur J Nutr 58:2365–2376. https://doi.org/10.1007/s00394-018-1788-9
Kovatcheva-Datchary P, Nilsson A, Akrami R et al (2015) Dietary fiber-induced improvement in glucose metabolism is associated with increased abundance of Prevotella. Cell Metab 22:971–982. https://doi.org/10.1016/j.cmet.2015.10.001
Johnson AJ, Zheng JJ, Kang JW et al (2020) A guide to diet-microbiome study design. Front Nutr 7:79–79. https://doi.org/10.3389/fnut.2020.00079
Maskarinec G, Hullar MAJ (2020) Understanding the interaction of diet quality with the gut microbiome and their effect on disease. J Nutr 150:654–655. https://doi.org/10.1093/jn/nxaa015
Rodríguez-Carrio J, Salazar N, Margolles A et al (2017) Free fatty acids profiles are related to gut microbiota signatures and short-chain fatty acids. Front Immunol 8:823–823. https://doi.org/10.3389/fimmu.2017.00823
Saini RK, Keum Y-S (2018) Omega-3 and omega-6 polyunsaturated fatty acids: dietary sources, metabolism, and significance—a review. Life Sci 203:255–267. https://doi.org/10.1016/j.lfs.2018.04.049
Ríos-Covián D, Ruas-Madiedo P, Margolles A et al (2016) Intestinal short chain fatty acids and their link with diet and human health. Front Microbiol 7:185–185. https://doi.org/10.3389/fmicb.2016.00185
Xu F, Tavintharan S, Sum CF et al (2013) Metabolic signature shift in type 2 diabetes mellitus revealed by mass spectrometry-based metabolomics. J Clin Endocrinol Metab 98:E1060-1065. https://doi.org/10.1210/jc.2012-4132
Krebs M, Krssak M, Bernroider E et al (2002) Mechanism of amino acid-induced skeletal muscle insulin resistance in humans. Diabetes 51:599–605. https://doi.org/10.2337/diabetes.51.3.599
Holmsen H, Hindenes JO, Fukami M (1992) Glycerophospholipid metabolism: back to the future. Thromb Res 67:313–323. https://doi.org/10.1016/0049-3848(92)90006-v
Nehlig A (2018) Interindividual differences in caffeine metabolism and factors driving caffeine consumption. Pharmacol Rev 70:384. https://doi.org/10.1124/pr.117.014407
Yang A, Palmer AA, de Wit H (2010) Genetics of caffeine consumption and responses to caffeine. Psychopharmacology 211:245–257. https://doi.org/10.1007/s00213-010-1900-1
Siddiqui A, Ceppi P (2020) A non-proliferative role of pyrimidine metabolism in cancer. Mol Metab 35:100962–100962. https://doi.org/10.1016/j.molmet.2020.02.005
Willett W (2012) Nutritional epidemiology. Oxford University Press
Althouse AD (2016) Adjust for multiple comparisons? It’s not that simple. Ann Thorac Surg 101:1644–1645. https://doi.org/10.1016/j.athoracsur.2015.11.024
Funding
This study was supported by the NIH, (R01 HL142856), by an AHA Scientist Development Grant (15SDG24890015), and a P&F Award from the Vanderbilt University Medical Center’s Digestive Disease Research Center supported by NIH grant P30DK058404.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have no conflicts of interest to disclose.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bagheri, M., Shah, R.D., Mosley, J.D. et al. A metabolome and microbiome wide association study of healthy eating index points to the mechanisms linking dietary pattern and metabolic status. Eur J Nutr 60, 4413–4427 (2021). https://doi.org/10.1007/s00394-021-02599-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00394-021-02599-9