Abstract
Objectives
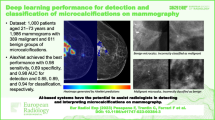
To analyze the performance of deep learning in isodense/obscure masses in dense breasts. To build and validate a deep learning (DL) model using core radiology principles and analyze its performance in isodense/obscure masses. To show performance on screening mammography as well as diagnostic mammography distribution.
Methods
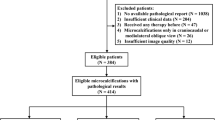
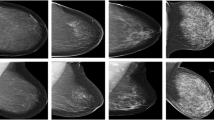
This was a retrospective, single-institution, multi-centre study with external validation. For model building, we took a 3-pronged approach. First, we explicitly taught the network to learn features other than density differences: such as spiculations and architectural distortion. Second, we used the opposite breast to enable the detection of asymmetries. Third, we systematically enhanced each image by piece-wise-linear transformation. We tested the network on a diagnostic mammography dataset (2569 images with 243 cancers, January to June 2018) and a screening mammography dataset (2146 images with 59 cancers, patient recruitment from January to April 2021) from a different centre (external validation).
Results
When trained with our proposed technique (and compared with baseline network), sensitivity for malignancy increased from 82.7 to 84.7% at 0.2 False positives per image (FPI) in the diagnostic mammography dataset, 67.9 to 73.8% in the subset of patients with dense breasts, 74.6 to 85.3 in the subset of patients with isodense/obscure cancers and 84.9 to 88.7 in an external validation test set with a screening mammography distribution. We showed that our sensitivity exceeded currently reported values (0.90 at 0.2 FPI) on a public benchmark dataset (INBreast).
Conclusion
Modelling traditional mammographic teaching into a DL framework can help improve cancer detection accuracy in dense breasts.
Clinical relevance statement
Incorporating medical knowledge into neural network design can help us overcome some limitations associated with specific modalities. In this paper, we show how one such deep neural network can help improve performance on mammographically dense breasts.
Key Points
• Although state-of-the-art deep learning networks achieve good results in cancer detection in mammography in general, isodense, obscure masses and mammographically dense breasts posed a challenge to deep learning networks.
• Collaborative network design and incorporation of traditional radiology teaching into the deep learning approach helped mitigate the problem.
• The accuracy of deep learning networks may be translatable to different patient distributions. We showed the results of our network on screening as well as diagnostic mammography datasets.






Similar content being viewed by others
Change history
22 June 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00330-023-09851-2
Abbreviations
- CABD:
-
Contrast adjusted for breast density
- CBIS-DDSM:
-
Curated Breast Imaging Subset of DDSM, a publicly available dataset
- CI:
-
Confidence interval
- DICOM:
-
Digital imaging and communications in medicine
- DL:
-
Deep learning
- DM:
-
Diagnostic mammography dataset
- DNN:
-
Deep neural network
- EfficientDet:
-
A state-of-art object detection network
- FFDM:
-
Full-field digital mammography
- FPI:
-
False positives per image
- FRCNN:
-
Faster RCNN, a state-of-art object detection network
- PACS:
-
Picture archiving communication system
- PNG:
-
Portable network graphics file format
- RIS:
-
Radiology information system
- RPN:
-
Region proposal network
- SM:
-
Screening mammography dataset
- SOTA:
-
State-of-the art
- TI:
-
Thresholded image
- YOLO v3:
-
You Only Look Once version 3, a State-of-art Object Detection Network
References
Seely JM, Alhassan T (2018) Screening for breast cancer in 2018-what should we be doing today? Curr Oncol Tor Ont. https://doi.org/10.3747/co.25.3770
Mandelson MT, Oestreicher N, Porter PL et al (2000) Breast density as a predictor of mammographic detection: comparison of interval- and screen-detected cancers. J Natl Cancer Inst. https://doi.org/10.1093/jnci/92.13.1081
Majid AS, de Paredes ES, Doherty RD, Sharma NR, Salvador X (2003) Missed breast carcinoma: pitfalls and pearls. Radiographics. https://doi.org/10.1148/rg.234025083
Freer PE (2015) Mammographic breast density: impact on breast cancer risk and implications for screening. Radiographics. https://doi.org/10.1148/rg.352140106
Patel MR, Whitman GJ (1998) Negative mammograms in symptomatic patients with breast cancer. Acad Radiol. https://doi.org/10.1016/s1076-6332(98)80008-1
Kopans DB (2007) Breast Imaging. Lippincott Williams & Wilkins, 1135 p
Abdelhafiz D, Yang C, Ammar R, Nabavi S (2019) Deep convolutional neural networks for mammography: advances, challenges and applications. BMC Bioinform. https://doi.org/10.1186/s12859-019-2823-4
Freeman K, Geppert J, Stinton C et al (2021) Use of artificial intelligence for image analysis in breast cancer screening programmes: systematic review of test accuracy. BMJ. https://doi.org/10.1136/bmj.n1872
Lee R, Gimenez F, Hoogi A et al (2017) A curated mammography data set for use in computer-aided detection and diagnosis research. Sci Data. https://doi.org/10.1038/sdata.2017.177
Moreira IC, Amaral I, Domingues I et al (2012) INbreast: toward a full-field digital mammographic database. Acad Radiol. https://doi.org/10.1016/j.acra.2011.09.014
Ren S, He K, Girshick R, Sun J (2015) Faster R-CNN: Towards real-time object detection with region proposal networks. In: Advances in Neural Information Processing Systems (NIPS)
Chakraborty DP (2013) A brief history of FROC paradigm data analysis. Acad Radiol. https://doi.org/10.1016/j.acra.2013.03.001
Kozegar E, Soryani M, Minaei B, Domingues I (2013) Assessment of a novel mass detection algorithm in mammograms. J Can Res Ther. https://doi.org/10.4103/0973-1482.126453
Akselrod-Ballin A, Karlinsky L, Hazan A et al (2017) Deep learning for automatic detection of abnormal findings in breast mammography. In: Lecture Notes in Computer Science 321–329. https://doi.org/10.1007/978-3-319-67558-9_37
Dhungel N, Carneiro G, Bradley AP (2015) Automated mass detection in mammograms using cascaded deep learning and random forests. In: 2015 International Conference on Digital Image Computing: Techniques and Applications (DICTA). https://doi.org/10.1109/DICTA.2015.7371234
Ribli D, Horváth A, Unger Z, Pollner P, Csabai I (2018) Detecting and classifying lesions in mammograms with deep learning. Sci Rep. https://doi.org/10.1038/s41598-018-22437-z
Agarwal R, Diaz O, Lladó X, Yap MH, Martí R (2019) Automatic mass detection in mammograms using deep convolutional neural networks. J Med Imaging. https://doi.org/10.1117/1.JMI.6.3.031409
Yala A, Schuster T, Miles R, Barzilay R, Lehman C (2019) A deep learning model to triage screening mammograms: a simulation study. Radiology. https://doi.org/10.1148/radiol.2019182908
Dembrower K, Wåhlin E, Liu Y et al (2020) Effect of artificial intelligence-based triaging of breast cancer screening mammograms on cancer detection and radiologist workload: a retrospective simulation study. Lancet Digit Health. https://doi.org/10.1016/S2589-7500(20)30185-0
Hepsağ PU, Özel SA, Yazıcı A (2017) Using deep learning for mammography classification. International Conference on Computer Science and Engineering (UBMK). https://doi.org/10.1109/UBMK.2017.8093429
Zhu W, Lou Q, Vang YS, Xie X (2017) Deep multi-instance networks with sparse label assignment for whole mammogram classification. bioRxiv. https://doi.org/10.1101/095794
Jung H, Kim B, Lee I et al (2018) Detection of masses in mammograms using a one-stage object detector based on a deep convolutional neural network. PLoS One. https://doi.org/10.1371/journal.pone.0203355
Shen L, Margolies LR, Rothstein JH, Fluder E, McBride R, Sieh W (2019) Deep learning to improve breast cancer detection on screening mammography. Sci Rep. https://doi.org/10.1038/s41598-019-48995-4
Zebari DA, Ibrahim DA, Zeebaree DQ et al (2021) Systematic review of computing approaches for breast cancer detection based computer aided diagnosis using mammogram images. Appl Artif Intell. https://doi.org/10.1080/08839514.2021.2001177
Maghsoudi OH, Gastounioti A, Scott C (2021) Deep-LIBRA: an artificial-intelligence method for robust quantification of breast density with independent validation in breast cancer risk assessment. Med Image Anal. https://doi.org/10.1016/j.media.2021.102138
Gong X, Yang Z, Wang D, Qi Y, Guo Y, Ma Y (2019) Breast density analysis based on glandular tissue segmentation and mixed feature extraction. Multimed Tools Appl. https://doi.org/10.1007/s11042-019-07917-2
Guan Y, Wang X, Li H et al (2020) Detecting asymmetric patterns and localizing cancers on mammograms. Patterns. https://doi.org/10.1016/j.patter.2020.100106
Hagos YB, Merida AG, Teuwen J (2018) Improving breast cancer detection using symmetry information with deep learning. ArXiv180808273. https://doi.org/10.1007/978-3-030-00946-5_10
Kooi T, Karssemeijer N (2017) Classifying symmetrical differences and temporal change for the detection of malignant masses in mammography using deep neural networks. J Med Imaging. https://doi.org/10.1117/1.JMI.4.4.044501
Acknowledgements
We acknowledge the help provided by Dr M Kalaivani, Professor, Department of Biostatistics, AIIMS for her valuable guidance in the project. We also acknowledge the role of our data entry operator Hema Malhotra in meticulously compiling the data required for this project. In addition, we also acknowledge Dr Pankaj Hari, Professor of Pediatrics, AIIMS Delhi for his help in the biostatistical analysis.
Funding
This work was supported in part by the Department of Biotechnology, Government of India, under Grant BT/PR33193/AI/133/5/2019.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Dr Krithika Rangarajan, Assistant Professor, Radiology, AIIMS, New Delhi.
Conflict of interest
The authors of this manuscript declare no relationships with any companies whose products or services may be related to the subject matter of the article.
Statistics and biometry
Dr M. Kalaivani, Professor, Department of Biostatistics, AIIMS kindly provided statistical advice for this manuscript.
Informed consent
Informed consent was waived by the institutional review board.
Ethical approval
Institutional Review Board approval was obtained. (IEC-247-4.05.2018). The title of the project for which ethical clearance was obtained is “Deep learning for detection and classification of abnormalities on full field digital mammography.” The current manuscript is one of the works done under this project.
Study subjects or cohorts overlap
Mammograms from study subjects are being used for the development of other AI algorithms for cancer detection on mammograms as well. However, none of the other studies deals with the detection of cancers in dense breasts and uses completely different neural networks. The current neural network is not reported on the data presented in the manuscript in any other work.
Methodology
This was a retrospective diagnostic accuracy study performed at 2 centres of one institution. These centres were the Department of Radiology, All India Institute of Medical Sciences New Delhi (centre 1) and the Department of Oncoradiology, BR Ambedkar Institute Rotary Cancer Hospital, AIIMS (centre 2), New Delhi. The DM dataset and Institutional training dataset described in the manuscript were obtained from centre 1, and the SM dataset was obtained from centre 2. The original ethical clearance obtained from IRB was for mammography and pathology information from centre 1 (IEC-247–4.05.2018). Centre 2 mammography and pathology information was allowed by IRB as an addendum to the same ethical clearance (IEC-247–4.05.2018 OP03/04.06.2021). The other 2 datasets described are publicly available datasets which are downloadable. These can be accessed from:
• The CBIS-DDSM dataset is available from https://wiki.cancerimagingarchive.net/display/Public/CBIS-DDSM
• The INBreast dataset is available upon request from http://medicalresearch.inescporto.pt/breastresearch/GetINb
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: In this article the author name Pranjal Aggarwal was incorrectly written as Pranjal Agarwal.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Rangarajan, K., Aggarwal, P., Gupta, D.K. et al. Deep learning for detection of iso-dense, obscure masses in mammographically dense breasts. Eur Radiol 33, 8112–8121 (2023). https://doi.org/10.1007/s00330-023-09717-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-023-09717-7