Abstract
Objective
To automate the segmentation of whole liver parenchyma on multi-echo chemical shift encoded (MECSE) MR examinations using convolutional neural networks (CNNs) to seamlessly quantify precise organ-related imaging biomarkers such as the fat fraction and iron load.
Methods
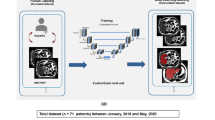
A retrospective multicenter collection of 183 MECSE liver MR examinations was conducted. An encoder-decoder CNN was trained (107 studies) following a 5-fold cross-validation strategy to improve the model performance and ensure lack of overfitting. Proton density fat fraction (PDFF) and R2* were quantified on both manual and CNN segmentation masks. Different metrics were used to evaluate the CNN performance over both unseen internal (46 studies) and external (29 studies) validation datasets to analyze reproducibility.
Results
The internal test showed excellent results for the automatic segmentation with a dice coefficient (DC) of 0.93 ± 0.03 and high correlation between the quantification done with the predicted mask and the manual segmentation (rPDFF = 1 and rR2* = 1; p values < 0.001). The external validation was also excellent with a different vendor but the same magnetic field strength, proving the generalization of the model to other manufacturers with DC of 0.94 ± 0.02. Results were lower for the 1.5-T MR same vendor scanner with DC of 0.87 ± 0.06. Both external validations showed high correlation in the quantification (rPDFF = 1 and rR2* = 1; p values < 0.001). In both internal and external validation datasets, the relative error for the PDFF and R2* quantification was below 4% and 1% respectively.
Conclusion
Liver parenchyma can be accurately segmented with CNN in a vendor-neutral virtual approach, allowing to obtain reproducible automatic whole organ virtual biopsies.
Key points
• Whole liver parenchyma can be automatically segmented using convolutional neural networks.
• Deep learning allows the creation of automatic pipelines for the precise quantification of liver-related imaging biomarkers such as PDFF and R2*.
• MR “virtual biopsy” can become a fast and automatic procedure for the assessment of chronic diffuse liver diseases in clinical practice.






Similar content being viewed by others
Abbreviations
- AI:
-
Artificial intelligence
- ASSD:
-
Average symmetric surface distance
- CNN:
-
Convolutional neural network
- DC:
-
Dice coefficient
- FDR:
-
False discovery rate
- MECSE:
-
Multi-echo chemical shift encoded
- MSD:
-
Maximum surface distance
- NAFLD:
-
Non-alcoholic fatty liver disease
- NASH:
-
Non-alcoholic steatohepatitis
- PDFF:
-
Proton density fat fraction
- RVD:
-
Relative volume difference
- VOE:
-
Volumetric overlap error
References
Donato H, França M, Candelária I, Caseiro-Alves F (2017) Liver MRI: from basic protocol to advanced techniques. Eur J Radiol 93:30–39
Farrell GC, Larter CZ (2006) Nonalcoholic fatty liver disease: from steatosis to cirrhosis. Hepatology. 43(2 Suppl 2):S99–S112
Kinner S, Reeder SB, Yokoo T (2016) Quantitative imaging biomarkers of NAFLD. Dig Dis Sci 61(5):1337–1347
França M, Alberich-Bayarri Á, Martí-Bonmatí L et al (2017) Accurate simultaneous quantification of liver steatosis and iron overload in diffuse liver diseases with MRI. Abdom Radiol (NY) 42(5):1434–1443
Kühn JP, Hernando D, Muñoz del Rio A et al (2012) Effect of multipeak spectral modeling of fat for liver iron and fat quantification: correlation of biopsy with MR imaging results. Radiology. 265(1):133–142
Rostoker G, Laroudie M, Blanc R et al (2017) Signal-intensity-ratio MRI accurately estimates hepatic iron load in hemodialysis patients. Heliyon. 3(1):e00226
Ratziu V, Charlotte F, Heurtier A et al (2005) Sampling variability of liver biopsy in nonalcoholic fatty liver disease. Gastroenterology. 128(7):1898–1906
Butensky E, Fischer R, Hudes M et al (2005) Variability in hepatic iron concentration in percutaneous needle biopsy specimens from patients with transfusional hemosiderosis. Am J Clin Pathol 123(1):146–152
Deugnier Y, Turlin B (2007) Pathology of hepatic iron overload. World J Gastroenterol 13(35):4755–4760
Martí-Bonmatí L, Alberich-Bayarri A, Sánchez-González J (2012) Overload hepatitides: quanti-qualitative analysis. Abdom Imaging 37(2):180–187
McCarville MB, Hillenbrand CM, Loeffler RB et al (2010) Comparison of whole liver and small region-of-interest measurements of MRI liver R2* in children with iron overload. Pediatr Radiol 40(8):1360–1367
Campo CA, Hernando D, Schubert T, Bookwalter CA, Pay AJV, Reeder SB (2017) Standardized approach for ROI-based measurements of proton density fat fraction and R2* in the liver. AJR Am J Roentgenol 209(3):592–603
Esfandiarkhani M, Foruzan AH (2017) A generalized active shape model for segmentation of liver in low-contrast CT volumes. Comput Biol Med 82:59–70
Yan Z, Zhang S, Tan C et al (2015) Atlas-based liver segmentation and hepatic fat-fraction assessment for clinical trials. Comput Med Imaging Graph 41:80–92
Dura E, Domingo J, Göçeri E, Martí-Bonmatí L (2017) A method for liver segmentation in perfusion MR images using probabilistic atlases and viscous reconstruction. Pattern Anal Appl 21(4):1083–1095
Xu Y, Xu C, Kuang X et al (2016) 3D-SIFT-Flow for atlas-based CT liver image segmentation. Med Phys 43(5):2229
Yuan Z, Wang Y, Yang J, Liu Y (2010). A novel automatic liver segmentation technique for MR images. 2010 3rd International Congress on Image and Signal Processing, Yantai. 1282–1286
Göçeri E (2016) Fully automated liver segmentation using Sobolev gradient-based level set evolution. Int J Numer Method Biomed Eng 32(11)
Yang D, Xu D, Zhou SK et al (2017). Automatic liver segmentation using an adversarial image-to-image network. Medical Image Computing and Computer-Assisted. MICCAI 2017. Lecture Notes in Computer Science, vol 10435. Springer, Cham
Han X (2017). Automatic liver lesion segmentation using a deep convolutional neural network method. arXiv: 1704.07239
Qin W, Wu J, Han F et al (2018) Superpixel-based and boundary-sensitive convolutional neural network for automated liver segmentation. Phys Med Biol 63(9):095017
Christ PF, Ettlinger F, Grün F et al (2017). Automatic liver and tumor segmentation of CT and MRI volumes using cascaded fully convolutional neural networks. arXiv: 1702.05970
Wang K, Mamidipalli A, Retson T et al (2019) Automated CT and MRI liver segmentation and biometry using a generalized convolutional neural network. Radiol Artif Intell 1(2):180022
Lavdas I, Glocker B, Kamnitsas K et al (2017) Fully automatic, multiorgan segmentation in normal whole body magnetic resonance imaging (MRI), using classification forests (CFs), convolutional neural networks (CNNs), and a multi-atlas (MA) approach. Med Phys 44(10):5210–5220
Jansen MJA, Kuijf HJ, Niekel M et al (2019) Liver segmentation and metastases detection in MR images using convolutional neural networks. J Med Imaging (Bellingham) 6(4):044003
Montagnon E, Cerny M, Cadrin-Chênevert A et al (2020) Deep learning workflow in radiology: a primer. Insights Imaging 11(1)
Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted. MICCAI 2015. Lecture Notes in Computer Science, vol 9351. Springer, Cham
Wang L, Yang Y, Min R, Chakradhar S (2017) Accelerating deep neural network training with inconsistent stochastic gradient descent. Neural Netw 93:219–229
Lee C, Xie S, Gallagher P, Zhang Z, Tu Z (2015). Deeply-supervised nets. Artif Intell Stat 562–570
Dou Q, Chen H, Jin Y, Yu L, Qin J, Heng P (2016) 3D deeply supervised network for automatic liver segmentation from CT volumes. Medical Image Computing and Computer-Assisted. MICCAI 2016. Springer, Cham
Sudre CH, Li W, Vercauteren T, Ourselin S, Cardoso MJ (2017). Generalised Dice overlap as a deep learning loss function for highly unbalanced segmentations. Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. DLMIA 2017, ML-CDS 2017. Lecture Notes in Computer Science, vol 10553. Springer, Cham
Kingma DP, Ba J (2015) Adam: a method for stochastic optimization. international conference on learning representations. 1–13. arXiv: 1412.6980
Stocker D, Bashir MR, Kannengiesser SAR, Reiner CS (2018) Accuracy of automated liver contouring, fat fraction, and R2* measurement on gradient multiecho magnetic resonance images. J Comput Assist Tomogr 42(5):697–706
Fezza SA, Bakhti Y, Hamidouche W, Deforges O (2019) Perceptual evaluation of adversarial attacks for CNN-based image classification. 2019 Eleventh International Conference on Quality of Multimedia Experience (QoMEX). 1–6
Chen L, Bentley P, Mori K, Misawa K, Fujiwara M, Rueckert D (2019) Intelligent image synthesis to attack a segmentation CNN using adversarial learning. Simulation and Synthesis in Medical Imaging. SASHIMI 2019. Lecture Notes in Computer Science, vol 11827. Springer, Cham
Yan W, Huang L, Xia L et al (2020) MRI manufacturer shift and adaptation: increasing the generalizability of deep learning segmentation for MR images acquired with different scanners. Radiol Artif Intell 2(4):e190195
Acknowledgements
DMA is recipient of a Río Hortega award (CM19/00212), Instituto de Salud Carlos III.
Funding
This study was partially funded by the Spanish Ministry of Science and innovation, Instituto de Salud Carlos III (PI19/0380) and GILEAD Sciences (Grant Number: GLD19/00050). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Guarantor
The scientific guarantor of this publication is Luis Marti-Bonmati.
Conflict of interest
The authors of this manuscript declare relationships with the following companies: QUIBIM SL.
Statistics and biometry
No complex statistical methods were necessary for this paper.
Informed consent
Written informed consent was waived by the Institutional Review Board.
Ethical approval
Institutional Review Board approval was obtained.
Methodology
• Retrospective
• Observational
• Multicenter study
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Jimenez-Pastor, A., Alberich-Bayarri, A., Lopez-Gonzalez, R. et al. Precise whole liver automatic segmentation and quantification of PDFF and R2* on MR images. Eur Radiol 31, 7876–7887 (2021). https://doi.org/10.1007/s00330-021-07838-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00330-021-07838-5