Abstract
Key message
TFL1-like genes of the basal eudicot Platanus acerifolia have conserved roles in maintaining vegetative growth and inhibiting flowering, but may act through distinct regulatory mechanism.
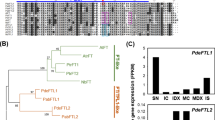
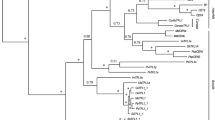
Three TERMINAL FLOWER 1 (TFL1)-like genes were isolated and characterized from London plane tree (Platanus acerifolia). All genes have conserved genomic organization and characteristic of the phosphatidylethanolamine-binding protein (PEBP) family. Sequence alignment and phylogenetic analysis indicated that two genes belong to the TFL1 clade, designated as PlacTFL1a and PlacTFL1b, while another one was grouped in the BFT clade, named as PlacBFT. qRT-PCR analysis showed that all three genes primarily expressed in vegetative phase, but the expression of PlacTFL1a was much higher and wider than that of PlacTFL1b, with the latter only detected at relatively low expression levels in apical and lateral buds in April. PlacBFT was mainly expressed in young stems of adult trees followed by juvenile tissues. Ectopic expression of any TFL1-like gene in Arabidopsis showed phenotypes of delayed or repressed flowering. Furthermore, overexpression of PlacTFL1a gene in petunia also resulted in extremely delayed flowering. In non-flowering 35:PlacTFL1a transgenic petunia plants, the FT-like gene (PhFT) gene was significantly upregulated and AP1 homologues PFG, FBP26 and FBP29 were significantly down-regulated in leaves. Yeast two-hybrid analysis indicated that only weak interactions were detected between PlacTFL1a and PlacFDL, and PlacTFL1a showed no interaction with PhFDL1/2. These results indicated that the TFL1-like genes of Platanus have conserved roles in repressing flowering, but probably via a distinct regulatory mechanism.









Similar content being viewed by others
Availability of data and materials
All data used in this research are included in this published article and its supplementary information files.
Code availability
Not applicable.
References
Ahn JH, Miller D, Winter VJ, Banfield MJ, Lee JH, Yoo SY, Henz SR, Brady RL, Weigel D (2006) A divergent external loop confers antagonistic activity on floral regulators FT and TFL1. EMBO J 25:605–614
Bai S, Tuan PA, Saito T, Ito A, Ubi BE, Ban Y, Moriguchi T, Wilson Z (2017) Repression of TERMINAL FLOWER1 primarily mediates floral induction in pear (Pyrus pyrifolia Nakai) concomitant with change in gene expression of plant hormone-related genes and transcription factors. J Exp Bot 68:4899–4914
Bai M, Liu J, Fan C, Chen Y, Chen H, Lu J, Sun J, Ning G, Wang C (2021) KSN heterozygosity is associated with continuous flowering of Rosa rugosa Purple branch. Hortic Res 8:26
Bi Z, Tahir AT (2019) Cloning and functional analysis of five TERMINAL FLOWER 1/CENTRORADIALIS-like genes from Hevea brasiliensis. Physiol Plantarum 166:612–627
Bi X, van Esse W, Mulki MA (2019) CENTRORADIALIS interacts with FLOWERING LOCUS T-like genes to control floret development and grain number. Plant Physiol 180:1013–1030
Blackman BK, Strasburg JL, Raduski AR, Michaels SD, Rieseberg LH (2010) The role of recently derived FT paralogs in sunflower domestication. Curr Biol 20:629–635
Bradley D, Carpenter R, Copsey L, Vincent C, Rothstein S, Coen E (1996) Control of inflorescence architecture in Antirrhinum. Nature 379:791–797
Bradley D, Ratcliffe O, Vincent C, Carpenter R, Coen E (1997) Inflorescence commitment and architecture in Arabidopsis. Science 275:80–83
Byng JW, Chase MW, Christenhusz MJM, Fay MF, Judd WS, Mabberley DJ, Sennikov AN, Soltis DE, Soltis PS, Stevens PF, Briggs B, Brockington S, Chautems A, Clark JC, Conran J, Haston E, Moller M, Moore M, Olmstead R, Perret M, Skog L, Smith J, Tank D, Vorontsova M, Weber A, Grp AP (2016) An update of the Angiosperm Phylogeny Group classification for the orders and families of flowering plants: APG IV. Bot J Linn Soc 181:1–20
Cai F, Shao C, Zhang Y, Bao Z, Li Z, Shi G, Bao M, Zhang J (2019) Identification and characterisation of a novel FT orthologous gene in London plane with a distinct expression response to environmental stimuli compared to PaFT. Plant Biol 21:1039–1051
Cao K, Cui L, Zhou X, Ye L, Zou Z, Deng S (2015) Four tomato FLOWERING LOCUS T-like proteins act antagonistically to regulate floral initiation. Front Plant Sci 6:1213
Cheng X, Li G, Tang Y, Wen J (2018) Dissection of genetic regulation of compound inflorescence development in Medicago truncatula. Development 145.3:dev158766-dev158766.
Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16:735–743
Corbesier L, Vincent C, Jang S, Fornara F, Fan Q, Searle I, Giakountis A, Farrona S, Gissot L, Turnbull C, Coupland G (2007) FT protein movement contributes to long-distance signaling in floral induction of Arabidopsis. Science 316:1030–1033
Fernandez L, Torregrosa L, Segura V, Bouquet A, Martinez-Zapater JM (2010) Transposon-induced gene activation as a mechanism generating cluster shape somatic variation in grapevine. Plant J 61:545–557
Foucher F, Morin J, Courtiade J, Cadioux S, Ellis N, Banfield MJ, Rameau C (2003) DETERMINATE and LATE FLOWERING are two TERMINAL FLOWER1/CENTRORADIALIS homologs that control two distinct phases of flowering initiation and development in pea. Plant Cell 15:2742–2754
Goretti D, Silvestre M (2020) TERMINAL FLOWER1 Functions as a mobile transcriptional cofactor in the shoot apical meristem. Plant Physiol 182:2081–2095
Goslin K, Zheng B, Serrano-Mislata A (2017) Transcription factor interplay between LEAFY and APETALA1/CAULIFLOWER during floral initiation. Plant Physiol 174:1097–1109
Guo JL, Yu CL, Fan CY, Lu QN, Yin JM, Zhang YF, Yang Q (2010) Cloning and characterization of a potato TFL1 gene involved in tuberization regulation. Plant Cell Tiss Org 103:103–109
Hanano S, Goto K (2011) Arabidopsis TERMINAL FLOWER1 is involved in the regulation of flowering time and inflorescence development through transcriptional repression. Plant Cell 23:3172–3184
Hanzawa Y, Money T, Bradley D (2005) A single amino acid converts a repressor to an activator of flowering. Proc Natl Acad Sci USA 102:7748–7753
Harig L, Beinecke FA, Oltmanns J, Muth J, Müller O, Rüping B, Twyman RM, Fischer R, Prüfer D, Noll GA (2012) Proteins from the FLOWERING LOCUS T-like subclade of the PEBP family act antagonistically to regulate floral initiation in tobacco. Plant J 72:908–921
Hedman H, Källman T, Lagercrantz U (2009) Early evolution of the MFT-like gene family in plants. Plant Mol Biol 70:359–369
Ho WW, Weigel D (2014) Structural features determining flower-promoting activity of Arabidopsis FLOWERING LOCUS T. Plant Cell 26:552–564
Huang NC, Jane WN, Chen J, Yu TS (2012) Arabidopsis thaliana CENTRORADIALIS homologue (ATC) acts systemically to inhibit floral initiation in Arabidopsis. Plant J 72:175–184
Immink RG, Hannapel DJ, Ferrario S, Busscher M, Franken J, Lookeren Campagne MM, Angenent GC (1999) A petunia MADS box gene involved in the transition from vegetative to reproductive development. Development 126:5117–5126
Jin S, Nasim Z, Susila H, Ahn JH (2021) Evolution and functional diversification of FLOWERING LOCUS T/TERMINAL FLOWER 1 family genes in plants. Semin Cell Dev Biol 109:20–30
Kaneko-Suzuki M, Kurihara-Ishikawa R, Okushita-Terakawa C, Kojima C, Nagano-Fujiwara M, Ohki I, Tsuji H, Shimamoto K, Taoka KI (2018) TFL1-like proteins in rice antagonize rice ft-like protein in inflorescence development by competition for complex formation with 14-3-3 and FD. Plant Cell Physiol 59:458–468
Karlgren A, Gyllenstrand N, Källman T, Sundström JF, Moore D, Lascoux M, Lagercrantz U (2011) Evolution of the PEBP gene family in plants: functional diversification in seed plant evolution. Plant Physiol 156:1967–1977
Kobayashi Y, Kaya H, Goto K, Iwabuchi M, Araki T (1999) A pair of related genes with antagonistic roles in mediating flowering signals. Science 286:1960–1962
Kumar S, Tamura K, Nei M (2004) MEGA3: Integrated software for molecular evolutionary genetics analysis and sequence alignment. Brief Bioinform 5:150–163
Lee C, Kim SJ, Jin S, Susila H, Youn G, Nasim Z, Alavilli H, Chung KS, Yoo SJ, Ahn JH (2019) Genetic interactions reveal the antagonistic roles of FT/TSF and TFL1 in the determination of inflorescence meristem identity in Arabidopsis. Plant J 99:452–464
Li Y, Zhang B (2021) DOTFL1 affects the floral transition in orchid Dendrobium Chao Praya Smile. Plant Physiol 186:2021–2036
Li Z, Liu G, Zhang J, Zhang J, Bao M (2008) Extraction of high-quality tissue-specific RNA from London plane trees (Platanus acerifolia), permitting the construction of a female inflorescence cDNA library. Funct Plant Biol 35:159–165
Li C, Luo L, Fu Q, Niu L, Xu ZF (2015) Identification and characterization of the FT/TFL1 gene family in the biofuel plant Jatropha curcas. Springer, New York, p 2
Li C, Fu Q, Niu L, Luo L, Chen J, Xu ZF (2017) Three TFL1 homologues regulate floral initiation in the biofuel plant Jatropha curcas. Sci Rep 7:43090
Liu G, Li Z, Bao M (2007) Colchicine-induced chromosome doubling in Platanus acerifolia and its effect on plant morphology. Euphytica 157:145–154
Liu Y, Zhang D, Ping J, Li S, Chen Z, Ma J (2016a) Innovation of a regulatory mechanism modulating semi-determinate stem growth through artificial selection in soybean. PLoS Genet 12:e1005818
Liu YY, Yang KZ, Wei XX, Wang XQ (2016b) Revisiting the phosphatidylethanolamine-binding protein (PEBP) gene family reveals cryptic FLOWERING LOCUS T gene homologs in gymnosperms and sheds new light on functional evolution. New Phytol 212:730–744
Lu S, Li Z, Zhang J, Yi S, Liu L, Bao M, Liu G (2012) Isolation and expression analysis of a LEAFY/FLORICAULA homolog and its promoter from London plane (Platanus acerifolia Willd.). Plant Cell Rep 31:1851–1865
Mallona I, Lischewski S, Weiss J, Hause B, Egea-Cortines M (2010) Validation of reference genes for quantitative real-time PCR during leaf and flower development in Petunia hybrida. BMC Plant Biol 10:4
McGarry RC, Prewitt SF, Culpepper S, Eshed Y, Lifschitz E, Ayre BG (2016) Monopodial and sympodial branching architecture in cotton is differentially regulated by the Gossypium hirsutum SINGLE FLOWER TRUSS and SELF-PRUNING orthologs. New Phytol 212:244–258
Mimida N, Goto K, Kobayashi Y, Araki T, Ahn JH, Weigel D, Murata M, Motoyoshi F, Sakamoto W (2001) Functional divergence of the TFL1-like gene family in Arabidopsis revealed by characterization of a novel homologue. Genes Cells 6:327–336
Mimida N, Kotoda N, Ueda T, Igarashi M, Hatsuyama Y, Iwanami H, Moriya S, Abe K (2009) Four TFL1/CEN-like genes on distinct linkage groups show different expression patterns to regulate vegetative and reproductive development in apple (Malus x domestica Borkh.). Plant Cell Physiol 50:394–412
Ming R, VanBuren R, Liu Y, Yang M, Han Y, Li LT, Zhang Q, Kim MJ, Schatz MC, Campbell M, Li J, Bowers JE, Tang H, Lyons E, Ferguson AA, Narzisi G, Nelson DR, Blaby-Haas CE, Gschwend AR, Jiao Y, Der JP, Zeng F, Han J, Min XJ, Hudson KA, Singh R, Grennan AK, Karpowicz SJ, Watling JR, Ito K, Robinson SA, Hudson ME, Yu Q, Mockler TC, Carroll A, Zheng Y, Sunkar R, Jia R, Chen N, Arro J, Wai CM, Wafula E, Spence A, Han Y, Xu L, Zhang J, Peery R, Haus MJ, Xiong W, Walsh JA, Wu J, Wang ML, Zhu YJ, Paull RE, Britt AB, Du C, Downie SR, Schuler MA, Michael TP, Long SP, Ort DR, Schopf JW, Gang DR, Jiang N, Yandell M, dePamphilis CW, Merchant SS, Paterson AH, Buchanan BB, Li S, Shen-Miller J (2013) Genome of the long-living sacred lotus (Nelumbo nucifera Gaertn.). Genome Biol 14:R41
Morel P, Chambrier P (2019) Divergent functional diversification patterns in the SEP/AGL6/AP1 MADS-box transcription factor superclade. Plant Cell 31:3033–3056
Nakagawa M, Shimamoto K, Kyozuka J (2002) Overexpression of RCN1 and RCN2, rice TERMINAL FLOWER 1/CENTRORADIALIS homologs, confers delay of phase transition and altered panicle morphology in rice. Plant J 29:743–750
Parcy F, Bomblies K, Weigel D (2002) Interaction of LEAFY, AGAMOUS and TERMINAL FLOWER1 in maintaining floral meristem identity in Arabidopsis. Development 129:2519–2527
Périlleux C, Bouché F, Randoux M, Orman-Ligeza B (2019) Turning meristems into fortresses. Trends Plant Sci 24:431–442
Pin PA, Benlloch R, Bonnet D, Wremerth-Weich E, Kraft T, Gielen JJ, Nilsson O (2010) An antagonistic pair of FT homologs mediates the control of flowering time in sugar beet. Science 330:1397–1400
Pnueli L, Carmel-Goren L, Hareven D, Gutfinger T, Alvarez J, Ganal M, Zamir D, Lifschitz E (1998) The SELF-PRUNING gene of tomato regulates vegetative to reproductive switching of sympodial meristems and is the ortholog of CEN and TFL1. Development 125:1979–1989
Randoux M, Davière JM, Jeauffre J, Thouroude T, Pierre S, Toualbia Y, Perrotte J, Reynoird JP, Jammes MJ, Hibrand-Saint Oyant L, Foucher F (2014) RoKSN, a floral repressor, forms protein complexes with RoFD and RoFT to regulate vegetative and reproductive development in rose. New Phytol 202:161–173
Ratcliffe OJ, Amaya I, Vincent CA, Rothstein S, Carpenter R, Coen ES, Bradley DJ (1998) A common mechanism controls the life cycle and architecture of plants. Development 125:1609–1615
Ratcliffe OJ, Bradley DJ, Coen ES (1999) Separation of shoot and floral identity in Arabidopsis. Development 126:1109–1120
Repinski SL, Kwak M, Gepts P (2012) The common bean growth habit gene PvTFL1y is a functional homolog of Arabidopsis TFL1. Theor Appl Genet 124:1539–1547
Serrano-Mislata A, Fernández-Nohales P, Doménech MJ, Hanzawa Y, Bradley D, Madueño F (2016) Separate elements of the TERMINAL FLOWER 1 cis-regulatory region integrate pathways to control flowering time and shoot meristem identity. Development 143:3315–3327
Serrano-Mislata A, Goslin K, Zheng B, Rae L, Wellmer F, Graciet E, Madueño F (2017) Regulatory interplay between LEAFY, APETALA1/CAULIFLOWER and TERMINAL FLOWER1: New insights into an old relationship. Plant Signal Behav 12:e1370164
Si Z, Liu H, Zhu J, Chen J, Wang Q, Fang L, Gao F, Tian Y, Chen Y, Chang L, Liu B, Han Z, Zhou B, Hu Y, Huang X, Zhang T (2018) Mutation of SELF-PRUNING homologs in cotton promotes short-branching plant architecture. J Exp Bot 69:2543–2553
Tian Z, Wang X, Lee R, Li Y, Specht JE, Nelson RL, McClean PE, Qiu L, Ma J (2010) Artificial selection for determinate growth habit in soybean. Proc Natl Acad Sci USA 107:8563–8568
Varkonyi-Gasic E, Moss SMA, Voogd C, Wang T, Putterill J, Hellens RP (2013) Homologs of FT, CEN and FD respond to developmental and environmental signals affecting growth and flowering in the perennial vine kiwifruit. New Phytol 198:732–746
Wang R, Albani MC, Vincent C, Bergonzi S, Luan M, Bai Y, Kiefer C, Castillo R, Coupland G (2011a) Aa TFL1 confers an age-dependent response to vernalization in perennial Arabis alpina. Plant Cell 23:1307–1321
Wang Z, Ye S, Li J, Zheng B, Bao M, Ning G (2011b) Fusion primer and nested integrated PCR (FPNI-PCR): a new high-efficiency strategy for rapid chromosome walking or flanking sequence cloning. BMC Biotechnol 11:109
Wang Y, Fan G, Liu Y, Sun F, Shi C, Liu X, Peng J, Chen W, Huang X, Cheng S, Liu Y, Liang X, Zhu H, Bian C, Zhong L, Lv T, Dong H, Liu W, Zhong X, Chen J, Quan Z, Wang Z, Tan B, Lin C, Mu F, Xu X, Ding Y, Guo AY, Wang J, Ke W (2013) The sacred lotus genome provides insights into the evolution of flowering plants. Plant J 76:557–567
Wang Z, Zhou Z, Liu Y, Liu T, Li Q, Ji Y, Li C, Fang C, Wang M, Wu M, Shen Y, Tang T, Ma J, Tian Z (2015) Functional evolution of phosphatidylethanolamine binding proteins in soybean and Arabidopsis. Plant Cell 27:323–336
Wickland DP, Hanzawa Y (2015) The FLOWERING LOCUS T/TERMINAL FLOWER 1 gene family: functional evolution and molecular mechanisms. Mol Plant 8:983–997
Wu L, Li F, Deng Q, Zhang S, Zhou Q, Chen F, Liu B, Bao M, Liu G (2019) Identification and characterization of the FLOWERING LOCUS T/TERMINAL FLOWER 1 gene family in Petunia. DNA Cell Biol 38:982–995
Xi W, Liu C, Hou X, Yu H (2010) MOTHER OF FT AND TFL1 regulates seed germination through a negative feedback loop modulating ABA signaling in Arabidopsis. Plant Cell 22:1733–1748
Yamaguchi A, Kobayashi Y, Goto K, Abe M, Araki T (2005) TWIN SISTER OF FT (TSF) acts as a floral pathway integrator redundantly with FT. Plant Cell Physiol 46:1175–1189
Yoo SY, Kardailsky I, Lee JS, Weigel D, Ahn JH (2004) Acceleration of flowering by overexpression of MFT (MOTHER OF FT AND TFL1). Mol Cells 17:95–101
Yoo SJ, Chung KS, Jung SH, Yoo SY, Lee JS, Ahn JH (2010) BROTHER OF FT AND TFL1 (BFT) has TFL1-like activity and functions redundantly with TFL1 in inflorescence meristem development in Arabidopsis. Plant J63:241–253
Zhang J, Liu G, Guo C, He Y, Li Z, Ning G, Shi X, Bao M (2011) The FLOWERING LOCUS T orthologous gene of Platanus acerifolia is expressed as alternatively spliced forms with distinct spatial and temporal patterns. Plant Biol 13:809–820
Zhang S, Lu S, Yi S, Han H, Zhou Q, Cai F, Bao M, Liu G (2019) Identification and characterization of FRUITFULL-like genes from Platanus acerifolia, a basal eudicot tree. Plant Sci 280:206–218
Zhang F, Wang Y, Irish VF (2021) CENTRORADIALIS maintains shoot meristem indeterminacy by antagonizing THORN IDENTITY1 in Citrus. Curr Biol 31:2237-2242.e2234
Zhu Y, Klasfeld S, Jeong CW, Jin R, Goto K, Yamaguchi N (2020) TERMINAL FLOWER 1-FD complex target genes and competition with FLOWERING LOCUS T. Nat Commun 11:5118
Funding
This work was financially supported by the National Natural Science Foundation of China (31272206, 32102425, 31772345) and the Bureau of Forestry and Landscaping of Wuhan Municipality (Grant No. WHGF2022A09, WHGF2019A05). We thank all colleagues in our laboratory for discussions and technical assistance.
Author information
Authors and Affiliations
Contributions
LGF and BMZ designed the experiments. ZSS, ZQ, YXY, LYJ and SMM performed the experiments. ZSS, WJQ, JJ and NCR analyzed the data. ZSS and LGF wrote and revised the manuscript. All authors participated in the research and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Peishan Yi.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, S., Zhou, Q., Yang, X. et al. Functional characterization of three TERMINAL FLOWER 1-like genes from Platanus acerifolia. Plant Cell Rep 42, 1071–1088 (2023). https://doi.org/10.1007/s00299-023-03014-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-023-03014-9