Abstract
Key message
The identification of N -glycosylated proteins with information about changes in the level of N -glycosylation during de-etiolation provides a database that will aid further research on plant N -glycosylation and de-etiolation.
Abstract
N-glycosylation is one of the most prominent and abundant protein post-translational modifications in all eukaryotes and in plants it plays important roles in development, stress tolerance and immune responses. Because light-induced de-etiolation is one of the most dramatic developmental processes known in plants, seedlings undergoing de-etiolation are an excellent model for investigating dynamic proteomic profiles. Here, we present a comprehensive, quantitative N-glycoproteomic profile of maize seedlings undergoing 12 h of de-etiolation obtained using Concanavalin A (Con A) lectin affinity chromatography enrichment coupled with a nano-LC–MS/MS-based iTRAQ approach. In total, 1084 unique N-glycopeptides carrying 909 N-glycosylation sites and corresponding to 609 proteins were identified and quantified, including 186 N-glycosylation sites from 162 proteins that were significantly regulated over the course of the 12 h de-etiolation period. Based on hierarchical clustering analysis, the significantly regulated N-glycopeptides were divided into seven clusters that showed different N-glycosylation patterns during de-etiolation. We found no obvious difference in the enriched MapMan bincode categories for each cluster, and these clustered significantly regulated N-glycoproteins (SRNPs) are enriched in miscellaneous, protein, cell wall and signaling, indicating that although the N-glycosylation regulation patterns of these SRNPs might differ, they are involved in similar biological processes. Overall, this study represents the first large-scale quantitative N-glycoproteome of the model C4 plant, maize, which is one of the most important cereal and biofuel crops. Our results greatly expand the maize N-glycoproteomic database and also shed light on the potential roles of N-glycosylation modification during the greening of maize leaves.






Similar content being viewed by others
Abbreviations
- Con A:
-
Concanavalin A
- ER:
-
Endoplasmic reticulum
- GO:
-
Gene ontology
- iTRAQ:
-
Isobaric tags for relative and absolute quantification
- LAC:
-
Lectin affinity chromatography
- LC–MS/MS:
-
Liquid chromatography and tandem mass spectrometry
- RLK:
-
Receptor-like kinase
- SRNP:
-
Significantly regulated N-glycoprotein
References
Albert M, Jehle AK, Mueller K, Eisele C, Lipschis M, Felix G (2010) Arabidopsis thaliana pattern recognition receptors for bacterial elongation factor Tu and flagellin can be combined to form functional chimeric receptors. J Biol Chem 285:19035–19042. doi:10.1074/jbc.M110.124800
An HJ, Froehlich JW, Lebrilla CB (2009) Determination of glycosylation sites and site-specific heterogeneity in glycoproteins. Curr Opin Chem Biol 13:421–426. doi:10.1016/j.cbpa.2009.07.022
Arsovski AA, Galstyan A, Guseman JM, Nemhauser JL (2012) Photomorphogenesis. Arabidopsis Book 10:e0147. doi:10.1199/tab.0147
Barba-Espin G, Dedvisitsakul P, Hagglund P, Svensson B, Finnie C (2014) Gibberellic acid-induced aleurone layers responding to heat shock or tunicamycin provide insight into the N-glycoproteome, protein secretion, and endoplasmic reticulum stress. Plant Physiol 164:951–965. doi:10.1104/pp.113.233163
Borisov OV, Field M, Ling VT, Harris RJ (2009) Characterization of oligosaccharides in recombinant tissue plasminogen activator produced in Chinese hamster ovary cells: two decades of analytical technology development. Anal Chem 81:9744–9754. doi:10.1021/ac901498k
Bouton S et al (2002) QUASIMODO1 encodes a putative membrane-bound glycosyltransferase required for normal pectin synthesis and cell adhesion in Arabidopsis. Plant Cell 14:2577–2590
Breton C, Mucha J, Jeanneau C (2001) Structural and functional features of glycosyltransferases. Biochimie 83:713–718. doi:10.1016/S0300-9084(01)01298-6
Buren S et al (2011) Importance of post-translational modifications for functionality of a chloroplast-localized carbonic anhydrase (CAH1) in Arabidopsis thaliana. PLoS One 6:e21021. doi:10.1371/journal.pone.0021021
Cano-Delgado A et al (2004) BRL1 and BRL3 are novel brassinosteroid receptors that function in vascular differentiation in Arabidopsis. Development 131:5341–5351. doi:10.1242/Dev.01403
Capone S et al (2015) Combining Protein and Strain Engineering for the Production of Glyco-Engineered Horseradish Peroxidase C1A in Pichia pastoris. Int J Mol Sci 16:23127–23142. doi:10.3390/ijms161023127
Casal JJ, Whitelam GC, Smith H (1990) Phytochrome control of extracellular peroxidase-activity in mustard internodes—correlation with growth, and comparison with the effect of wounding. Photochem Photobiol 52:165–172. doi:10.1111/j.1751-1097.1990.tb01770.x
Catala C, Howe KJ, Hucko S, Rose JKC, Thannhauser TW (2011) Towards characterization of the glycoproteome of tomato (Solanum lycopersicum) fruit using Concanavalin A lectin affinity chromatography and LC-MALDI-MS/MS analysis. Proteomics 11:1530–1544. doi:10.1002/pmic.201000424
Chen M, Chory J, Fankhauser C (2004) Light signal transduction in higher plants. Annu Rev Genet 38:87–117. doi:10.1146/annurev.genet.38.072902.092259
Chi YH, Koo YD, Dai SY, Ahn JE, Yun DJ, Lee SY, Zhu-Salzman K (2010) N-glycosylation at non-canonical Asn-X-Cys sequence of an insect recombinant cathepsin B-like counter-defense protein. Comp Biochem Phys B 156:40–47. doi:10.1016/j.cbpb.2010.01.017
Chou MF, Schwartz D (2011) Biological sequence motif discovery using motif-x. Current protocols in bioinformatics, Chap 13, Unit 13, pp 15–24. doi:10.1002/0471250953.bi1315s35
Craig R, Beavis RC (2004) TANDEM: matching proteins with tandem mass spectra. Bioinformatics 20:1466–1467. doi:10.1093/bioinformatics/bth092
Craig R, Cortens JP, Beavis RC (2004) Open source system for analyzing, validating, and storing protein identification data. J Proteome Res 3:1234–1242. doi:10.1021/pr049882h
De Smet I, Voss U, Jurgens G, Beeckman T (2009) Receptor-like kinases shape the plant. Nat Cell Biol 11:1166–1173. doi:10.1038/ncb1009-1166
DeYoung BJ, Clark SE (2008) BAM receptors regulate stem cell specification and organ development through complex interactions with CLAVATA signaling. Genetics 180:895–904
Doblin MS, Mohnen D, Bacic A (2016) Plant glycobiology: a current snap shot! Glycobiology 26:911–912
Dodds PN, Rathjen JP (2010) Plant immunity: towards an integrated view of plant-pathogen interactions. Nat Rev Genet 11:539–548. doi:10.1038/nrg2812
Fernández-Pérez F, Pomar F, Pedreño MA, Novo-Uzal E (2015) The suppression of AtPrx52 affects fibers but not xylem lignification in Arabidopsis by altering the proportion of syringyl units. Physiol Plant 154:395–406
Fitchette A-C, Dinh OT, Faye L, Bardor M (2007) Plant Proteomics and Glycosylation. In: Thiellement H, Zivy M, Damerval C, Méchin V (eds) Plant proteomics: methods and protocols. Humana Press, Totowa, pp 317–342. doi:10.1385/1-59745-227-0:317
Gilgunn S, Conroy PJ, Saldova R, Rudd PM, O’Kennedy RJ (2013) Aberrant PSA glycosylation-a sweet predictor of prostate cancer. Nat Rev Urol 10:99–107. doi:10.1038/nruro1.2012.258
Gupta N, Bandeira N, Keich U, Pevzner PA (2011) Target-decoy approach and false discovery rate: when things may go wrong. J Am Soc Mass Spectrom 22:1111–1120
Haltiwanger RS et al (1992) Glycosylation of nuclear and cytoplasmic proteins is ubiquitous and dynamic. Biochem Soc T 20:264–269
Hamamoto K et al (2012) Proteomic characterization of the greening process in rice seedlings using the MS spectral intensity-based label free method. J Proteome Res 11:331–347. doi:10.1021/pr200852q
Haweker H et al (2010) Pattern recognition receptors require N-glycosylation to mediate plant immunity. J Biolo Chem 285:4629–4636. doi:10.1074/jbc.M109.063073
Hayashi S et al (2008) The glycerophosphoryl diester phosphodiesterase-like proteins SHV3 and its homologs play important roles in cell wall organization. Plant Cell Physiol 49:1522–1535. doi:10.1093/pcp/pcn120
Helenius A, Aebi M (2004) Roles of N-linked glycans in the endoplasmic reticulum. Annu Rev Biochem 73:1019–1049. doi:10.1146/annurev.biochem.73.011303.073752
Jiao YL, Ma LG, Strickland E, Deng XW (2005) Conservation and divergence of light-regulated genome expression patterns during seedling development in rice and Arabidopsis The Plant cell 17:3239–3256. doi:10.1105/tpc.105.035840
Jin LJ, Wessely O, Marcusson EG, Ivan C, Calin GA, Alahari SK (2013) Prooncogenic factors miR-23b and miR-27b are regulated by Her2/Neu, EGF, and TNF-alpha in breast cancer. Cancer Res 73:2884–2896. doi:10.1158/0008-5472.CAN-12-2162
Jones J, Krag SS, Betenbaugh MJ (2005) Controlling N-linked glycan site occupancy. Biochimica et Biophysica Acta (BBA)-General Subj 1726:121–137
Kall L, Storey JD, MacCoss MJ, Noble WS (2008) Assigning significance to peptides identified by tandem mass spectrometry using decoy databases. J Proteome Res 7:29–34. doi:10.1021/pr700600n
Karam AK, Karlan BY (2010) Ovarian cancer: the duplicity of CA125 measurement. Nat Rev Clin Oncol 7:335–339. doi:10.1038/nrclinonc.2010.44
Kasturi L, Eshleman JR, Wunner WH, Shakineshleman SH (1995) The hydroxy amino-acid in an Asn-X-Ser/Thr sequon can influence n-linked core glycosylation efficiency and the level of expression of a cell-surface glycoprotein. J Biol Chem 270:14756–14761
Kendrick RE, Kronenberg GH (2012) Photomorphogenesis in plants. Springer Science & Business Media, New York
Klahre U et al (1998) The Arabidopsis DIMINUTO/DWARF1 gene encodes a protein involved in steroid synthesis. Plant Cell 10:1677–1690. doi:10.1105/tpc.10.10.1677
Kumar S et al (2013) Glycoproteome of Elongating Cotton Fiber Cells. Mol Cell Proteomics 12:3677–3689. doi:10.1074/mcp.M113.030726
Lannoo N, Van Damme EJ (2015) Review/N-glycans: the making of a varied toolbox. Plant Sci 239:67–83
Lehtimaki N, Koskela MM, Mulo P (2015) Posttranslational modifications of chloroplast proteins: an emerging field. Plant Physiol 168:768–775. doi:10.1104/pp.15.00117
Lehti-Shiu MD, Shiu SH (2012) Diversity, classification and function of the plant protein kinase superfamily. Philos T R Soc B 367:2619–2639. doi:10.1098/rstb.2012.0003
Lopez-Juez E et al (2008) Distinct light-initiated gene expression and cell cycle programs in the shoot apex and cotyledons of Arabidopsis. Plant Cell 20:947–968. doi:10.1105/tpc.107.057075
Ma LG, Li JM, Qu LJ, Hager J, Chen ZL, Zhao HY, Deng XW (2001) Light control of Arabidopsis development entails coordinated regulation of genome expression and cellular pathways. Plant cell 13:2589–2607. doi:10.1105/tpc.13.12.2589
Ma J, Wang D, She J, Li J, Zhu JK, She YM (2016) Endoplasmic reticulum-associated N-glycan degradation of cold-upregulated glycoproteins in response to chilling stress in Arabidopsis. New Phytol 212:282–296
Malaby HL, Kobertz WR (2014) The middle X residue influences cotranslational N-glycosylation consensus site skipping. Biochemistry-Us 53:4884–4893. doi:10.1021/bi500681p
Matsui T et al (2011) N-glycosylation at noncanonical Asn-X-Cys sequences in plant cells. Glycobiology 21:994–999. doi:10.1093/glycob/cwq198
McCarty DR, Chory J (2000) Conservation and innovation in plant signaling pathways. Cell 103:201–209. doi:10.1016/S0092-8674(00)00113-6
Minic Z, Jamet E, Négroni L, Der Garabedian PA, Zivy M, Jouanin L (2007) A sub-proteome of Arabidopsis thaliana mature stems trapped on Concanavalin A is enriched in cell wall glycoside hydrolases. J Exp Bot 58:2503–2512
Mohnen D, Tierney ML (2011) Plants Get Hyp to O-glycosylation. Science 332:1393–1394. doi:10.1126/science.1208641
Mullet JE (1988) Chloroplast development and gene-expression. Annu Rev Plant Phys 39:475–502. doi:10.1146/annurev.arplant.39.1.475
Mustafa G, Komatsu S (2014) Quantitative proteomics reveals the effect of protein glycosylation in soybean root under flooding stress. Front Plant Sci 5:Artn 627. doi:10.3389/Fpls.2014.00627
Nagai K, Ihara Y, Wada Y, Taniguchi N (1997) N-glycosylation is requisite for the enzyme activity and Golgi retention of N-acetylglucosaminyltransferase III. Glycobiology 7:769–776. doi:10.1093/glycob/7.6.769
Nagata N, Min YK, Nakano T, Asami T, Yoshida S (2000) Treatment of dark-grown Arabidopsis thaliana with a brassinosteroid-biosynthesis inhibitor, brassinazole, induces some characteristics of light-grown plants. Planta 211:781–790. doi:10.1007/s004250000351
Nanjo Y et al (2006) Rice plastidial N-glycosylated nucleotide pyrophosphatase/phosphodiesterase is transported from the ER-Golgi to the chloroplast through the secretory pathway. Plant Cell 18:2582–2592. doi:10.1105/tpc.105.039891
Nedelkov D, Nelson RW, Beavis RC (2006) Using the global proteome machine for protein identification. Methods Mol Biol 328:217–228. doi:10.1385/1-59745-026-X:217
Ning DL et al (2016) Large-scale comparative phosphoprotein analysis of maize seedling leaves during greening. Planta 243:501–517. doi:10.1007/s00425-015-2420-3
Olczak M, Olczak T (2002) Diphosphonucleotide phosphatase/phosphodiesterase from yellow lupin (Lupinus luteus L.) belongs to a novel group of specific metallophosphatases. FEBS Lett 519:159–163. doi:10.1016/S0014-5793(02)02740-0
Pantin F, Simonneau T, Rolland G, Dauzat M, Muller B (2011) Control of leaf expansion: a developmental switch from metabolics to hydraulics. Plant Physiol 156:803–815. doi:10.1104/pp.111.176289
Passardi F, Penel C, Dunand C (2004) Performing the paradoxical: how plant peroxidases modify the cell wall. Trends Plant Sci 9:534–540. doi:10.1016/j.tplants.2004.09.002
Pedersen CT et al (2017) N-glycan maturation mutants in Lotus japonicus for basic and applied glycoprotein research. Plant J 91:394–407. doi:10.1111/tpj.13570
Peschke M, Moog D, Klingl A, Maier UG, Hempel F (2013) Evidence for glycoprotein transport into complex plastids. Proc Natl Acad Sci USA 110:10860–10865. doi:10.1073/pnas.1301945110
Petersen TN, Brunak S, von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786. doi:10.1038/nmeth.1701
Pless DD, Lennarz WJ (1977) Enzymatic Conversion of Proteins to Glycoproteins. Proc Natl Acad Sci USA 74:134–138. doi:10.1073/pnas.74.1.134
Price NJ, Pinheiro C, Soares CM, Ashford DA, Ricardo CP, Jackson PA (2003) A biochemical and molecular characterization of LEP1, an extensin peroxidase from lupin. J Biol Chem 278:41389–41399. doi:10.1074/jbc.M304519200
Rappsilber J, Mann M, Ishihama Y (2007) Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nat Protoc 2:1896–1906. doi:10.1038/nprot.2007.261
Rips S, Bentley N, Jeong IS, Welch JL, von Schaewen A, Koiwa H (2014) Multiple N-glycans cooperate in the subcellular targeting and functioning of Arabidopsis KORRIGAN1. Plant Cell 26:3792–3808. doi:10.1105/tpc.114.129718
Rose JKC, Lee SJ (2010) Straying off the highway: trafficking of secreted plant proteins and complexity in the plant cell wall proteome. Plant Physiol 153:433–436. doi:10.1104/pp.110.154872
Ruiz-May E, Kim SJ, Brandizzi F, Rose JK (2012) The secreted plant N-glycoproteome and associated secretory pathways. Front Plant Sci 3:117. doi:10.3389/fpls.2012.00117
Ruiz-May E, Hucko S, Howe KJ, Zhang S, Sherwood RW, Thannhauser TW, Rose JKC (2014) A comparative study of lectin affinity based plant N-glycoproteome profiling using tomato fruit as a model. Mol Cell Proteom 13:566–579. doi:10.1074/mcp.M113.028969
Saito K, Fujimura-Kamada K, Furuta N, Kato U, Umeda M, Tanaka K (2004) Cdc50p, a protein required for polarized growth, associates with the Drs2p P-type ATPase implicated in phospholipid translocation in Saccharomyces cerevisiae. Mol Biol Cell 15:3418–3432. doi:10.1091/mbc.E03-11-0829
Schwartz D, Gygi SP (2005) An iterative statistical approach to the identification of protein phosphorylation motifs from large-scale data sets. Nat Biotechnol 23:1391–1398. doi:10.1038/nbt1146
Sedbrook JC, Carroll KL, Hung KF, Masson PH, Somerville CR (2002) The Arabidopsis SKU5 gene encodes an extracellular glycosyl phosphatidylinositol-anchored glycoprotein involved in directional root growth. Plant Cell 14:1635–1648. doi:10.1105/tpc.002360
Shen Z et al (2009) Label-free quantitative proteomics analysis of etiolated maize seedling leaves during greening. Mol Cell Proteom 8:2443–2460. doi:10.1074/mcp.M900187-MCP200
Shen JB, Ding Y, Gao CJ, Rojo E, Jiang LW (2014) N-linked glycosylation of AtVSR1 is important for vacuolar protein sorting in Arabidopsis. Plant J 80:977–992. doi:10.1111/tpj.12696
Song W et al (2011) N-glycoproteomics in plants: perspectives and challenges. J Proteom 74:1463–1474
Song W et al (2013) N-glycan occupancy of Arabidopsis N-glycoproteins. Journal of proteomics 93:343–355
Spiro RG (2002) Protein glycosylation: nature, distribution, enzymatic formation, and disease implications of glycopeptide bonds. Glycobiology 12:43R–56R
Strasser R (2014) Biological significance of complex N-glycans in plants and their impact on plant physiology. Front Plant Sci 5:Artn 363. doi:10.3389/Fpls.2014.00363
Strasser R, Schoberer J, Jin CS, Glossl J, Mach L, Steinkellner H (2006) Molecular cloning and characterization of Arabidopsis thaliana Golgi alpha-mannosidase II, a key enzyme in the formation of complex N-glycans in plants. Plant J 45:789–803. doi:10.1111/j.1365-313X.2005.02648.x
Sutimantanapi D, Pater D, Smith LG (2014) Divergent roles for maize PAN1 and PAN2 receptor-like proteins in cytokinesis and cell morphogenesis1[W][OPEN]. Plant Physiol 164:1905–1917. doi:10.1104/pp.113.232660
Takasaki T, Hatakeyama K, Suzuki G, Watanabe M, Isogai A, Hinata K (2000) The S receptor kinase determines self-incompatibility in Brassica stigma. Nature 403:913–916. doi:10.1038/35002628
Usadel B et al (2005) Extension of the visualization tool MapMan to allow statistical analysis of arrays, display of corresponding genes, and comparison with known responses. Plant Physiol 138:1195–1204. doi:10.1104/pp.105.060459
Van den Steen P, Rudd PM, Dwek RA, Opdenakker G (1998) Concepts and principles of O-linked glycosylation. Crit Rev Biochem Mol Biol 33:151–208
van der Hoorn RA, Wulff BB, Rivas S, Durrant MC, van der Ploeg A, de Wit PJ, Jones JD (2005) Structure-function analysis of cf-9, a receptor-like protein with extracytoplasmic leucine-rich repeats. Plant Cell 17:1000–1015
Wang BC, Pan YH, Meng DZ, Zhu YX (2006) Identification and quantitative analysis of significantly accumulated proteins during the Arabidopsis seedling de-etiolation process. J Integr Plant Biol 48:104–113. doi:10.1111/j.1744-7909.2006.00215.x
Wang Z-Y, Gehring C, Zhu J, Li F-M, Zhu J-K, Xiong L (2015) The Arabidopsis vacuolar sorting receptor1 is required for osmotic stress-induced abscisic acid biosynthesis. Plant Physiol 167:137–152
Wu XN, Rodriguez CS, Pertl-Obermeyer H, Obermeyer G, Schulze WX (2013) Sucrose-induced receptor kinase SIRK1 regulates a plasma membrane aquaporin in Arabidopsis. Mol Cell Proteomics 12:2856–2873. doi:10.1074/mcp.M113.029579
Xu S-L, Medzihradszky KF, Wang Z-Y, Burlingame AL, Chalkley RJ (2016) N-glycopeptide profiling in arabidopsis inflorescence. Mol Cell Proteom 15:2048–2054
Yamamoto M, Tantikanjana T, Nishio T, Nasrallah ME, Nasrallah JB (2014) Site-specific N-glycosylation of the S-locus receptor kinase and its role in the self-incompatibility response of the Brassicaceae. Plant cell 26:4749–4762
Yan AX, Lennarz WJ (2005) Unraveling the mechanism of protein N-glycosylation. J Biol Chem 280:3121–3124. doi:10.1074/jbc.R400036200
Yang PF, Chen H, Liang Y, Shen SH (2007) Proteomic analysis of de-etiolated rice seedlings upon exposure to light. Proteomics 7:2459–2468. doi:10.1002/pmic.200600215
Yang TB, Chaudhuri S, Yang LH, Du LQ, Poovaiah BW (2010) A calcium/calmodulin-regulated member of the receptor-like kinase family confers cold tolerance in plants. J Biol Chem 285:7119–7126. doi:10.1074/jbc.M109.035659
Zhang S et al (2012) Comparative characterization of the glycosylation profiles of an influenza hemagglutinin produced in plant and insect hosts. Proteomics 12:1269–1288. doi:10.1002/pmic.201100474
Zhang M, Chen GX, Lv DW, Li XH, Yan YM (2015) N-Linked glycoproteome profiling of seedling leaf in Brachypodium distachyon L. J Proteome Res 14:1727–1738. doi:10.1021/pr501080r
Zielinska DF, Gnad F, Schropp K, Wisniewski JR, Mann M (2012) Mapping N-glycosylation sites across seven evolutionarily distant species reveals a divergent substrate proteome despite a common core machinery. Mol Cell 46:542–548. doi:10.1016/j.molcel.2012.04.031
Zuo WL et al (2015) A maize wall-associated kinase confers quantitative resistance to head smut. Nat Genet 47:151–157. doi:10.1038/ng.3170
Acknowledgements
This study was funded by the “Strategic Priority Research Program” of the Chinese Academy of Sciences (Grant No. XDA08010206.5).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Mark C. Jordan.
Electronic supplementary material
Below is the link to the electronic supplementary material.
299_2017_2209_MOESM1_ESM.pdf
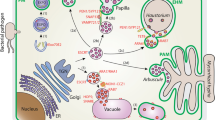
Fig. S1. Schematic of protein N-glycosylation in the secretory pathway. Enzymes identified as being N-glycosylated in this study are highlighted in yellow. (PDF 3978 kb)
Rights and permissions
About this article
Cite this article
Bu, Tt., Shen, J., Chao, Q. et al. Dynamic N-glycoproteome analysis of maize seedling leaves during de-etiolation using Concanavalin A lectin affinity chromatography and a nano-LC–MS/MS-based iTRAQ approach. Plant Cell Rep 36, 1943–1958 (2017). https://doi.org/10.1007/s00299-017-2209-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00299-017-2209-x