Abstract
The disease-associated nuclease–helicase DNA2 has been implicated in DNA end-resection during DNA double-strand break repair, Okazaki fragment processing, and the recovery of stalled DNA replication forks (RFs). Its role in Okazaki fragment processing has been proposed to explain why DNA2 is indispensable for cell survival across organisms. Unexpectedly, we found that DNA2 has an essential role in suppressing homologous recombination (HR)-dependent replication restart at stalled RFs. In the absence of DNA2-mediated RF recovery, excessive HR-restart of stalled RFs results in toxic levels of abortive recombination intermediates that lead to DNA damage-checkpoint activation and terminal cell-cycle arrest. While HR proteins protect and restart stalled RFs to promote faithful genome replication, these findings show how HR-dependent replication restart is actively constrained by DNA2 to ensure cell survival. These new insights disambiguate the effects of DNA2 dysfunction on cell survival, and provide a framework to rationalize the association of DNA2 with cancer and the primordial dwarfism disorder Seckel syndrome based on its role in RF recovery.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
In every cell cycle, cells face the enormous task of generating a faithful copy of their genome. Replisomes containing the Cdc45/Mcm2-7/GINS (CMG) replicative helicase unwind the parental chromosomes, forming DNA replication forks (RFs). At RFs, DNA polymerases ε and δ catalyse highly accurate leading- and lagging-strand DNA synthesis. The progression of RFs is routinely stalled by a range of obstacles including polymerase-blocking DNA lesions, DNA secondary structures, DNA-binding proteins, and DNA–RNA hybrids (Zeman and Cimprich 2014). Alterations in replication-origin firing caused by oncogene activation, depletion of replication factors, and low deoxyribonucleotide-triphosphate levels are further sources of replication stress that negatively impact RF progression. If stalled RFs are not properly dealt with, chromosomes remain partially unreplicated, jeopardizing chromosome disjunction at mitosis and the faithful transmission of the genome to daughter cells (Falquet and Rass 2019). Importantly, elevated replication stress has been linked to common fragile site expression, chromosome instability, and cancer (Gaillard et al. 2015; Glover et al. 2017).
In eukaryotes, DNA replication initiates at a multitude of replication origins, each giving rise to two RFs travelling in opposite directions along the parental chromosome. An inter-origin stretch of DNA is, therefore, replicated by a pair of converging RFs from adjacent origins (with the exception of DNA at the very tips of chromosomes), which provides protection against underreplication when individual RFs become blocked. In addition, dormant origins can activate in regions where DNA replication does not progress normally, providing another mechanism to make up for replication shortfalls upon RF-stalling or arrest. However, active and dormant origins are finite and cannot be set up de novo, while replication is ongoing (Siddiqui et al. 2013). Consequently, double-stalling events affecting a pair of converging RFs or single-stalling events in origin-poor regions of the genome cannot always be passively rescued by fork convergence to complete genome replication (Al Mamun et al. 2016). Mitigating the risk of local chromosome underreplication, cells have evolved mechanisms to protect, recover, and restart perturbed RFs, and homologous recombination (HR) proteins are intimately linked to these processes (Ait Saada et al. 2018).
Homologous recombination-dependent replication restart
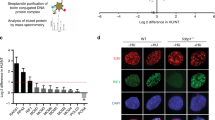
If stalled RFs remain replication-competent by retaining the replisome and their structural integrity, they may resume DNA synthesis once the replication impediment is resolved. However, stalled RFs quickly attract HR proteins (Nguyen et al. 2015), and more persistent RF-stalling renders replication completion more and more reliant on HR factors such as RAD51 (Petermann et al. 2010). This is in large part due to protective functions of various HR factors at stalled RFs that are independent of signature recombination reactions such as homology search and DNA strand invasion mediated by the Rad51-single-stranded DNA (ssDNA) filament (Mason et al. 2019; Zellweger et al. 2015). Accordingly, human RAD51 has been shown to promote a process known as RF reversal, which entails the dissociation of the nascent leading and lagging strands from the parental template and their annealing with one another, essentially turning three-way RFs into four-way DNA junctions known as reversed RFs, chicken-foot structures, or Holliday junctions (Fig. 1a). Reversed RFs have emerged as key intermediates for the protection of stalled RFs and the resumption of DNA synthesis by a variety of mechanisms (Neelsen and Lopes 2015). RAD51 and its primary loader in mammalian cells, BRCA2, protect the DNA end exposed at reversed RFs from degradation, maintaining them in a configuration that allows subsequent restart and/or fusion with an approaching active RF (Berti et al. 2020; Schlacher et al. 2011). Similarly, fork protection by Rad51 and its loader Rad52 is indispensable for replication completion upon RF-blockage in yeast (Ait Saada et al. 2017; Pardo et al. 2020).
Processing and restart of stalled RFs. a When a stalled RF cannot easily resume DNA synthesis, it may backtrack into a reversed position, forming a four-way DNA junction. Reversed RFs have emerged as key intermediates of RF recovery, facilitating passive rescue by fork convergence or further processing to restore RF activity. b Breakage of stalled and reversed RFs can occur through attack by structure-specific nucleases (black arrowheads). The resulting single-ended DNA double-strand break undergoes DNA end-resection and Rad51-mediated strand invasion to initiate HR-dependent replication restart by BIR. c HR-dependent replication restart can also occur at unbroken reversed RFs. End-resection at the regressed branch produces the 3′-DNA overhang for Rad51-mediated strand-invasion, restarting replication along the RDR pathway
Long-term exposure to replication stress is associated with the breakage of stalled RFs (Petermann et al. 2010), at least in part due to the action of structure-specific nucleases (also known as Holliday-junction resolvases) (Rass 2013). Nucleolytic cleavage results in severance of one sister chromatid from the fork and the formation of a single-ended DNA double-strand break. DNA end-resection at the broken sister chromatid results in the formation of a 3′-overhang, which can serve as a substrate to prime renewed DNA synthesis by DNA polymerase δ in the context of a displacement loop (D-loop) upon Rad51-mediated strand invasion of the unbroken parental chromosome. This pathway, which has been studied extensively in the budding yeast Saccharomyces cerevisiae (Lydeard et al. 2007, 2010), is known as break-induced replication (BIR) (Kramara et al. 2018) (Fig. 1b). During BIR, D-loop migration and extensive DNA synthesis are uniquely dependent upon the conserved helicase PIF1 (Saini et al. 2013; Wilson et al. 2013). Although the fate of the CMG replicative helicase during HR-dependent replication restart is not entirely clear, substituting CMG with Pif1 during D-loop DNA synthesis may solve the problem that the replicative helicase, once lost, cannot reload onto DNA in the S phase of the cell cycle.
A mechanism similar to BIR operates to restart stalled but unbroken RFs. Using the site-specific RF barrier provided by Schizosaccharomyces pombe RTS1 (replication termination sequence 1), it has been demonstrated that persistently stalled RFs restart through recombination-dependent replication (RDR) (Ahn et al. 2005; Lambert et al. 2005). Stalled RFs are engaged by the double-stranded DNA (dsDNA) end-binding protein Ku, which suggests that RF reversal precedes RDR (Teixeira-Silva et al. 2017). Ku stabilizes the reversed-fork structure, but can subsequently be removed through nucleolytic cleavage mediated by the Mre11–Rad50–Nbs1 (MRN) complex and CtIP. MRN-CtIP action results in limited DNA end-resection, followed by more extensive resection by Exo1, which generates a 3′-overhang for intramolecular, Rad51-dependent strand invasion ahead of the site of fork reversal to restart replication (Fig. 1c). DNA synthesis within the resulting D-loop is dependent—as in the case of BIR—on Pif1 (Jalan et al. 2019).
Homologous recombination-dependent replication: a salvage pathway with a cost to genetic stability
RF protection and restart by HR factors is vital for the completion of DNA replication, particularly under replication stress conditions. However, BIR and RDR are associated with a cost to genetic stability: BIR is characterized by high mutation frequencies (Deem et al. 2011) and the D-loop structures involved are intrinsically unstable. Thus, nascent DNA strands may undergo multiple strand eviction/invasion episodes, increasing the risk of ectopic recombination and formation of chromosome rearrangements (Costantino et al. 2014; Lambert et al. 2010; Mayle et al. 2015; Mizuno et al. 2013; Smith et al. 2007). While BIR/RDR can proceed for thousands of nucleotides, fusion with oncoming conventional RFs usually limits the extent of error-prone replication. In addition, the aforementioned mechanisms attenuating DNA end-resection at stalled and remodelled RFs antagonize the commitment to HR-dependent replication restart. For example, the initial protection by Ku from end-resection would favour a merger of a stalled RF with an approaching fork over RDR (Teixeira-Silva et al. 2017). Similarly, the resection nuclease Exo1 is constrained by the S-phase checkpoint (Tsang et al. 2014), while various regulators of RF remodelling and degradation have been identified in mammalian cells (Rickman and Smogorzewska 2019). Thus, DNA end-resection is a critical and highly controlled juncture for RF protection versus HR-dependent restart of stalled RFs. In budding yeast, protein sumoylation mediates the relocation of perturbed RFs to the nuclear pore, promoting Rad51-dependent replication restart and suggesting a spatial control mechanism governing commitment to HR-dependent replication restart (Nagai et al. 2008; Whalen et al. 2020). Quite clearly, cells carefully balance the need to restart persistently stalled RFs to avoid chromosome underreplication against the potential cost to genome stability that is associated with HR-dependent replication mechanisms.
Anything in excess is a poison: Dna2 limits homologous recombination-dependent restart at stalled replication forks
Our recent work suggests that the Dna2 nuclease–helicase plays a critical role in limiting RDR at stalled RFs (Falquet et al. 2020). DNA2 is essential across organisms and implicated in a variety of DNA metabolic processes including DNA double-strand break repair, checkpoint activation, Okazaki fragment processing, telomere homeostasis, centromeric DNA replication, and RF recovery (Zheng et al. 2020). In budding yeast, it has long been known that the lethality associated with loss of DNA2 can be suppressed by deletion (or nuclear exclusion) of PIF1 and/or DNA damage-checkpoint mediator RAD9. In the absence of Dna2, Pif1 and Rad9, therefore, mediate a toxic process. To explain this, it has been proposed that Pif1 stimulates strand-displacement DNA synthesis by DNA polymerase δ on the lagging strand, resulting in Okazaki fragments with extended 5′-flaps; if not cleaved by Dna2, these flaps are thought to associate with ssDNA-binding protein RPA, eliciting a Rad9-dependent DNA damage-checkpoint response, cell-cycle arrest, and ultimately cell death. These considerations have led to the concept that DNA2 fulfils its essential function by policing the generation of long 5′-flaps during Okazaki fragment maturation (Budd et al. 2011). However, overt lagging-strand DNA replication defects associated with DNA2 dysfunction have failed to transpire in yeast or human (Duxin et al. 2012; Kahli et al. 2019).
We found that—in the absence of DNA2 and PIF1, or DNA2 and RAD9—budding yeast cells fail to complete chromosome replication (Falquet et al. 2020), which is in keeping with Dna2′s documented role in stalled-RF recovery (Hu et al. 2012; Ölmezer et al. 2016; Thangavel et al. 2015). Under these conditions, survival became dependent upon Yen1, a Holliday-junction resolvase that is activated in anaphase (Blanco et al. 2014; Garcia-Luis et al. 2014; Ip et al. 2008), and which resolves persistent replication intermediates to facilitate viable chromosome segregation in DNA2-defective and dna2Δ cells (Falquet et al. 2020; Falquet and Rass 2017; Ölmezer et al. 2016). This showed that dna2Δ cells suffer from severe replication defects, and require failsafe resolution of underreplicated chromosomes by Yen1, and that these defects are fully independent of any action of Pif1. Accordingly, dna2Δ pif1Δ cells exhibit hypersensitivity upon exposure to exogenous replication stress. Next, we addressed whether the toxicity that is generated if Pif1 and Rad9 are present might be related to stalled RFs that are not properly processed when Dna2 is dysfunctional. Using hypomorphic dna2 cells, which—in contrast to dna2Δ cells—can be grown in the presence PIF1, we determined that Pif1 exacerbates dna2-linked DNA replication problems. Thus, Pif1 proved responsible for an unscheduled, post-replicative checkpoint response in dna2 cells that we have previously described (Ölmezer et al. 2016). This checkpoint response occurs immediately following passage through S phase under replication stress conditions, is distinct from Mec1–Ddc2/Mrc1/Rad53-mediated replication checkpoint signalling, and fully dependent upon DNA damage-checkpoint mediator Rad9. Consequently, Dna2 dysfunction in the presence of stalled RFs renders cells susceptible to cell-cycle arrest at the G2/M transition, which is enforced by the DNA damage checkpoint and elicited by the actions of Pif1. We surmised that RFs, which are not properly processed in the absence of DNA2, become susceptible to Pif1-mediated fork transitions, ultimately resulting in DNA damage-checkpoint activation and cell death. This provided an alternative to the Okazaki fragment processing model by explaining the essential nature of DNA2 with its role in stalled-RF recovery (Falquet et al. 2020).
What might toxic fork transitions mediated by Pif1 consist of? While Pif1 is a versatile helicase with multiple functions in DNA replication and repair (Chistol and Walter 2014), we found that unrelated point mutations within a stress-responsive phosphorylation motif and Pif1’s PCNA-interacting PIP box suppressed unscheduled DNA damage-checkpoint activation in response to RF-stalling and cell death in hypomorphic dna2 cells and dna2Δ cells, respectively (Falquet et al. 2020). Both of these Pif1 mutations converge on a specific Pif1 activity, blocking its ability to promote HR-coupled DNA synthesis (Buzovetsky et al. 2017; Vasianovich et al. 2014). From these observations, we concluded that Pif1/Rad9 toxicity is a direct consequence of inappropriate HR-dependent replication restart at RFs that are not properly processed by Dna2.
Given that HR-dependent replication restart provides a means to reboot DNA synthesis at perturbed RFs and complete DNA replication under stress conditions, why should this pathway be harmful in DNA2-defective cells? An important clue to answer this question came from a recent study investigating the types of aberrant DNA structures that accumulate upon depletion of Dna2 using electron microscopy (Rossi et al. 2018). In line with the previous findings in yeast and human cells (Hu et al. 2012; Thangavel et al. 2015), the authors found elevated levels of DNA four-way junctions, suggesting stalled RFs that escape processing by Dna2 accumulate in a reversed-fork configuration. Another intermediate, observed at even higher levels, consisted of dsDNA with one long branch of ssDNA that sometimes exceeded 10,000 nucleotides, representing a significant trigger for checkpoint activation upon coating with RPA. Since the Okazaki fragment processing machinery is biased against the formation of long DNA flaps, and extensive strand-displacement synthesis by DNA polymerases δ does not occur on the chromatinised lagging strand in vivo (Devbhandari et al. 2017; Garg et al. 2004; Kahli et al. 2019; Smith and Whitehouse 2012; Stodola and Burgers 2016), we suggest that these dsDNA/ssDNA intermediates are generated through RDR at stalled RFs by way of D-loop collapse. We propose a model where Dna2 counteracts RF reversal by degrading the nascent DNA strands at stalled RFs (Hu et al. 2012; Thangavel et al. 2015). Upon Dna2 dysfunction, reversed RFs accumulate and become available for HR-dependent replication restart. Consequently, there is excessive RDR (which can be suppressed by disrupting PIF1) and extensive D-loop DNA synthesis of thousands of nucleotides worth of DNA. Given the intrinsic instability of D-loops, nascent strands may disengage and remain permanently exposed as ssDNA. Accumulating levels of RPA-bound ssDNA will then activate the checkpoint (which can be suppressed by disrupting RAD9), causing a futile cell-cycle arrest in Dna2-deficient cells.
Future questions and concluding remarks
As shown in the model presented in Fig. 2, we envisage a dual role that makes Dna2 essential: first, Dna2 promotes RF recovery, thereby facilitating complete genome replication; second, Dna2 acts as gatekeeper to RDR, guarding cells against the lethal consequences of an excessive use of HR-dependent replication restart at stalled RFs. It remains unclear how the actions of Dna2 might integrate with other replication and repair factors at stalled RFs, limiting HR-dependent replication restart while allowing for it when required. Regulatory mechanisms could involve Dna2 phosphorylation (Chen et al. 2011; Hu et al. 2012), sumoylation (Ranjha et al. 2019), or protein inhibitors (Higgs et al. 2015). The exact processing reaction catalysed by Dna2 at stalled and/or reversed RFs also remains to be determined. Dna2 shows preferential binding of complex DNA structures including partially reversed RFs with a dissociated leading or lagging strand exposed as ssDNA (Park et al. 2020). Since Dna2 is capable of degrading ssDNA from the 5′- and 3′-ends (Cejka et al. 2010), its actions could, in principle, remove the dissociating nascent leading and lagging stands at stalled RFs as proposed previously (Hu et al. 2012), effectively pre-empting their annealing and formation of reversed RF structures. This is very different from the well-established role of Dna2 in DNA end-resection during DNA double-strand break repair by HR, where, alongside Exo1, Dna2 cooperates with the Sgs1 helicase (BLM in human) to unwind the DNA duplex and, stimulated by RPA, specifically degrades the 5′-terminated DNA strand (Cejka et al. 2010). Interestingly, in fission yeast, Exo1, but not Dna2-Rqh1 (the Sgs1 homologue), has been implicated in end-resection at reversed RFs to generate the 3′-overhand needed for RDR (Teixeira-Silva et al. 2017). Moreover, Pxd1 has been shown to associate with Dna2, suppressing the stimulatory effect that RPA has on Dna2 activity (Zhang et al. 2014). Dna2 activity thus appears malleable in a context-specific manner, but more work is required to determine whether Dna2 indiscriminately degrades nascent DNA strands at stalled RFs. While protection from replication stress depends chiefly on Dna2’s nuclease activity (Falquet et al. 2020), its helicase activity contributes to RF recovery (Ölmezer et al. 2016). This suggests that at least a subset of replication intermediates requires unwinding as well as DNA degradation by Dna2; this activity may target fully reversed RFs, for which Dna2 also shows preferential binding (Park et al. 2020).
Dna2 is an essential gatekeeper to HR-dependent restart of stalled replication forks. By degrading dissociated nascent DNA at stalled RFs, Dna2 counteracts RF reversal, promoting the resumption of DNA synthesis and/or RF convergence to mediate complete genome replication. By processing stalled RFs, Dna2 also limits opportunities for RF restart by RDR, and this makes Dna2 indispensable for cell survival. While RDR provides an important salvage pathway for stalled RFs, DNA synthesis in the context of a displacement loop (D-loop) is unstable. In the absence of Dna2, excessive RDR results in an accumulation of unsustainably high levels of recombination by-products by way of D-loop collapse. Exposure of long ssDNA tracts then causes Rad9-dependent checkpoint activation and terminal cell-cycle arrest. For further details, see main text
In conclusion, cells are set up to protect stalled RFs, resume DNA synthesis where possible, and ultimately rescue more persistently stalled forks using HR-dependent replication restart. While RDR/BIR are associated with a cost to genetic stability, their use reflects the overriding necessity to ensure that chromosomes are fully replicated before cell division. Our work indicates that RDR by-products can lead to cell death by eliciting a checkpoint-mediated cell-cycle arrest, uncovering an additional danger linked to the promiscuous use of HR-dependent replication restart. DNA2 averts this danger through the processing of stalled RFs, thereby limiting opportunities for RDR (Falquet et al. 2020). These findings provide a new rationale for the essential requirement of DNA2 for cell viability, which consists of establishing the correct balance between different RF recovery and restart pathways. Based on our work, an unbalanced response to RF-stalling resulting in diminished cell proliferation during development (Klingseisen and Jackson 2011) provides a plausible explanation for causative DNA2 mutations in patients with the primordial dwarfism disorder Seckel syndrome (Shaheen et al. 2014; Tarnauskaitė et al. 2019). In cancer, DNA2 has been found overexpressed (Peng et al. 2012; Strauss et al. 2014), potentially providing a survival advantage under elevated replication stress, which is prevalent in cancer cells. In future, it will be important to further address the RDR gatekeeper role of DNA2 in human cells.
References
Ahn JS, Osman F, Whitby MC (2005) Replication fork blockage by RTS1 at an ectopic site promotes recombination in fission yeast. EMBO J 24:2011–2023. https://doi.org/10.1038/sj.emboj.7600670
Ait Saada A et al (2017) Unprotected replication forks are converted into mitotic sister chromatid bridges. Mol Cell 66:398–410. https://doi.org/10.1016/j.molcel.2017.04.002
Ait Saada A, Lambert SAE, Carr AM (2018) Preserving replication fork integrity and competence via the homologous recombination pathway. DNA Repair (Amst) 71:135–147. https://doi.org/10.1016/j.dnarep.2018.08.017
Al Mamun M, Albergante L, Moreno A, Carrington JT, Blow JJ, Newman TJ (2016) Inevitability and containment of replication errors for eukaryotic genome lengths spanning megabase to gigabase. Proc Natl Acad Sci USA 113:E5765–E5774. https://doi.org/10.1073/pnas.1603241113
Berti M, Cortez D, Lopes M (2020) The plasticity of DNA replication forks in response to clinically relevant genotoxic stress. Nat Rev Mol Cell Biol. https://doi.org/10.1038/s41580-020-0257-5
Blanco MG, Matos J, West SC (2014) Dual control of Yen1 nuclease activity and cellular localization by Cdk and Cdc14 prevents genome instability. Mol Cell 54:94–106. https://doi.org/10.1016/j.molcel.2014.02.011
Budd ME, Antoshechkin IA, Reis C, Wold BJ, Campbell JL (2011) Inviability of a DNA2 deletion mutant is due to the DNA damage checkpoint. Cell Cycle 10:1690–1698. https://doi.org/10.4161/cc.10.10.15643
Buzovetsky O, Kwon Y, Pham NT, Kim C, Ira G, Sung P, Xiong Y (2017) Role of the Pif1–PCNA complex in Pol delta-dependent strand displacement DNA synthesis and break-induced replication. Cell Rep 21:1707–1714. https://doi.org/10.1016/j.celrep.2017.10.079
Cejka P, Cannavo E, Polaczek P, Masuda-Sasa T, Pokharel S, Campbell JL, Kowalczykowski SC (2010) DNA end resection by Dna2–Sgs1–RPA and its stimulation by Top3–Rmi1 and Mre11–Rad50–Xrs2. Nature 467:112–116. https://doi.org/10.1038/nature09355
Chen X et al (2011) Cell cycle regulation of DNA double-strand break end resection by Cdk1-dependent Dna2 phosphorylation. Nat Struct Mol Biol 18:1015–1019. https://doi.org/10.1038/nsmb.2105
Chistol G, Walter J (2014) Molecular watchdogs on genome patrol. Elife 3:e02854. https://doi.org/10.7554/eLife.02854
Costantino L et al (2014) Break-induced replication repair of damaged forks induces genomic duplications in human cells. Science 343:88–91. https://doi.org/10.1126/science.1243211
Deem A et al (2011) Break-induced replication is highly inaccurate. PLoS Biol 9:e1000594. https://doi.org/10.1371/journal.pbio.1000594
Devbhandari S, Jiang J, Kumar C, Whitehouse I, Remus D (2017) Chromatin constrains the initiation and elongation of DNA replication. Mol Cell 65:131–141. https://doi.org/10.1016/j.molcel.2016.10.035
Duxin JP et al (2012) Okazaki fragment processing-independent role for human Dna2 enzyme during DNA replication. J Biol Chem 287:21980–21991. https://doi.org/10.1074/jbc.M112.359018
Falquet B, Rass U (2017) A new role for Holliday junction resolvase Yen1 in processing DNA replication intermediates exposes Dna2 as an accessory replicative helicase. Microb Cell 4:32–34. https://doi.org/10.15698/mic2017.01.554
Falquet B, Rass U (2019) Structure-specific endonucleases and the resolution of chromosome underreplication. Genes (Basel) 10:232. https://doi.org/10.3390/genes10030232
Falquet B et al (2020) Disease-associated DNA2 nuclease–helicase protects cells from lethal chromosome under-replication. Nucleic Acids Res 48:7265–7278. https://doi.org/10.1093/nar/gkaa524
Gaillard H, Garcia-Muse T, Aguilera A (2015) Replication stress and cancer. Nat Rev Cancer 15:276–289. https://doi.org/10.1038/nrc3916
Garcia-Luis J, Clemente-Blanco A, Aragon L, Machin F (2014) Cdc14 targets the Holliday junction resolvase Yen1 to the nucleus in early anaphase. Cell Cycle 13:1392–1399. https://doi.org/10.4161/cc.28370
Garg P, Stith CM, Sabouri N, Johansson E, Burgers PM (2004) Idling by DNA polymerase delta maintains a ligatable nick during lagging-strand DNA replication. Genes Dev 18:2764–2773. https://doi.org/10.1101/gad.1252304
Glover TW, Wilson TE, Arlt MF (2017) Fragile sites in cancer: more than meets the eye. Nat Rev Cancer 17:489–501. https://doi.org/10.1038/nrc.2017.52
Higgs MR et al (2015) BOD1L is required to suppress deleterious resection of stressed replication forks. Mol Cell 59:462–477. https://doi.org/10.1016/j.molcel.2015.06.007
Hu J et al (2012) The intra-S phase checkpoint targets Dna2 to prevent stalled replication forks from reversing. Cell 149:1221–1232. https://doi.org/10.1016/j.cell.2012.04.030
Ip SC, Rass U, Blanco MG, Flynn HR, Skehel JM, West SC (2008) Identification of Holliday junction resolvases from humans and yeast. Nature 456:357–361. https://doi.org/10.1038/nature07470
Jalan M, Oehler J, Morrow CA, Osman F, Whitby MC (2019) Factors affecting template switch recombination associated with restarted DNA replication. Elife 8:e41697. https://doi.org/10.7554/eLife.41697
Kahli M, Osmundson JS, Yeung R, Smith DJ (2019) Processing of eukaryotic Okazaki fragments by redundant nucleases can be uncoupled from ongoing DNA replication in vivo. Nucleic Acids Res 47:1814–1822. https://doi.org/10.1093/nar/gky1242
Klingseisen A, Jackson AP (2011) Mechanisms and pathways of growth failure in primordial dwarfism. Genes Dev 25:2011–2024. https://doi.org/10.1101/gad.169037
Kramara J, Osia B, Malkova A (2018) Break-induced replication: the where, the why, and the how. Trends Genet 34:518–531. https://doi.org/10.1016/j.tig.2018.04.002
Lambert S, Watson A, Sheedy DM, Martin B, Carr AM (2005) Gross chromosomal rearrangements and elevated recombination at an inducible site-specific replication fork barrier. Cell 121:689–702. https://doi.org/10.1016/j.cell.2005.03.022
Lambert S et al (2010) Homologous recombination restarts blocked replication forks at the expense of genome rearrangements by template exchange. Mol Cell 39:346–359. https://doi.org/10.1016/j.molcel.2010.07.015
Lydeard JR, Jain S, Yamaguchi M, Haber JE (2007) Break-induced replication and telomerase-independent telomere maintenance require Pol32. Nature 448:820–823. https://doi.org/10.1038/nature06047
Lydeard JR, Lipkin-Moore Z, Sheu YJ, Stillman B, Burgers PM, Haber JE (2010) Break-induced replication requires all essential DNA replication factors except those specific for pre-RC assembly. Genes Dev 24:1133–1144. https://doi.org/10.1101/gad.1922610
Mason JM, Chan YL, Weichselbaum RW, Bishop DK (2019) Non-enzymatic roles of human RAD51 at stalled replication forks. Nat Commun 10:4410. https://doi.org/10.1038/s41467-019-12297-0
Mayle R et al (2015) DNA REPAIR. Mus81 and converging forks limit the mutagenicity of replication fork breakage. Science 349:742–747. https://doi.org/10.1126/science.aaa8391
Mizuno K, Miyabe I, Schalbetter SA, Carr AM, Murray JM (2013) Recombination-restarted replication makes inverted chromosome fusions at inverted repeats. Nature 493:246–249. https://doi.org/10.1038/nature11676
Nagai S et al (2008) Functional targeting of DNA damage to a nuclear pore-associated SUMO-dependent ubiquitin ligase. Science 322:597–602. https://doi.org/10.1126/science.1162790
Neelsen KJ, Lopes M (2015) Replication fork reversal in eukaryotes: from dead end to dynamic response. Nat Rev Mol Cell Biol 16:207–220. https://doi.org/10.1038/nrm3935
Nguyen MO, Jalan M, Morrow CA, Osman F, Whitby MC (2015) Recombination occurs within minutes of replication blockage by RTS1 producing restarted forks that are prone to collapse. Elife 4:e04539. https://doi.org/10.7554/eLife.04539
Ölmezer G, Levikova M, Klein D, Falquet B, Fontana GA, Cejka P, Rass U (2016) Replication intermediates that escape Dna2 activity are processed by Holliday junction resolvase Yen1. Nat Commun 7:13157. https://doi.org/10.1038/ncomms13157
Pardo B, Moriel-Carretero M, Vicat T, Aguilera A, Pasero P (2020) Homologous recombination and Mus81 promote replication completion in response to replication fork blockage. EMBO Rep 21:e49367. https://doi.org/10.15252/embr.201949367
Park S, Karatayeva N, Demin AA, Munashingha PR, Seo YS (2020) The secondary-structured DNA-binding activity of Dna2 endonuclease/helicase is critical to cell growth under replication stress. FEBS J. https://doi.org/10.1111/febs.15475(published online ahead of print, 2020 Jul 7)
Peng G et al (2012) Human nuclease/helicase DNA2 alleviates replication stress by promoting DNA end resection. Cancer Res 72:2802–2813. https://doi.org/10.1158/0008-5472.CAN-11-3152
Petermann E, Orta ML, Issaeva N, Schultz N, Helleday T (2010) Hydroxyurea-stalled replication forks become progressively inactivated and require two different RAD51-mediated pathways for restart and repair. Mol Cell 37:492–502. https://doi.org/10.1016/j.molcel.2010.01.021
Ranjha L, Levikova M, Altmannova V, Krejci L, Cejka P (2019) Sumoylation regulates the stability and nuclease activity of Saccharomyces cerevisiae Dna2. Commun Biol 2:174. https://doi.org/10.1038/s42003-019-0428-0
Rass U (2013) Resolving branched DNA intermediates with structure-specific nucleases during replication in eukaryotes. Chromosoma 122:499–515. https://doi.org/10.1007/s00412-013-0431-z
Rickman K, Smogorzewska A (2019) Advances in understanding DNA processing and protection at stalled replication forks. J Cell Biol 218:1096–1107. https://doi.org/10.1083/jcb.201809012
Rossi SE, Foiani M, Giannattasio M (2018) Dna2 processes behind the fork long ssDNA flaps generated by Pif1 and replication-dependent strand displacement. Nat Commun 9:4830. https://doi.org/10.1038/s41467-018-07378-5
Saini N et al (2013) Migrating bubble during break-induced replication drives conservative DNA synthesis. Nature 502:389–392. https://doi.org/10.1038/nature12584
Schlacher K, Christ N, Siaud N, Egashira A, Wu H, Jasin M (2011) Double-strand break repair-independent role for BRCA2 in blocking stalled replication fork degradation by MRE11. Cell 145:529–542. https://doi.org/10.1016/j.cell.2011.03.041
Shaheen R et al (2014) Genomic analysis of primordial dwarfism reveals novel disease genes. Genome Res 24:291–299. https://doi.org/10.1101/gr.160572.113
Siddiqui K, On KF, Diffley JF (2013) Regulating DNA replication in eukarya. Cold Spring Harb Perspect Biol. https://doi.org/10.1101/cshperspect.a012930
Smith DJ, Whitehouse I (2012) Intrinsic coupling of lagging-strand synthesis to chromatin assembly. Nature 483:434–438. https://doi.org/10.1038/nature10895
Smith CE, Llorente B, Symington LS (2007) Template switching during break-induced replication. Nature 447:102–105. https://doi.org/10.1038/nature05723
Stodola JL, Burgers PM (2016) Resolving individual steps of Okazaki-fragment maturation at a millisecond timescale. Nat Struct Mol Biol 23:402–408. https://doi.org/10.1038/nsmb.3207
Strauss C et al (2014) The DNA2 nuclease/helicase is an estrogen-dependent gene mutated in breast and ovarian cancers. Oncotarget 5:9396–9409. https://doi.org/10.18632/oncotarget.2414
Tarnauskaitė Ž et al (2019) Biallelic variants in DNA2 cause microcephalic primordial dwarfism. Hum Mutat 40:1063–1070. https://doi.org/10.1002/humu.23776
Teixeira-Silva A, Ait Saada A, Hardy J, Iraqui I, Nocente MC, Freon K, Lambert S (2017) The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks. Nat Commun 8:1982. https://doi.org/10.1038/s41467-017-02144-5
Thangavel S et al (2015) DNA2 drives processing and restart of reversed replication forks in human cells. J Cell Biol 208:545–562. https://doi.org/10.1083/jcb.201406100
Tsang E, Miyabe I, Iraqui I, Zheng J, Lambert S, Carr AM (2014) The extent of error-prone replication restart by homologous recombination is controlled by Exo1 and checkpoint proteins. J Cell Sci 127:2983–2994. https://doi.org/10.1242/jcs.152678
Vasianovich Y, Harrington LA, Makovets S (2014) Break-induced replication requires DNA damage-induced phosphorylation of Pif1 and leads to telomere lengthening. PLoS Genet 10:e1004679. https://doi.org/10.1371/journal.pgen.1004679
Whalen JM, Dhingra N, Wei L, Zhao X, Freudenreich CH (2020) Relocation of collapsed forks to the nuclear pore complex depends on sumoylation of DNA repair proteins and permits Rad51 association. Cell Rep 31:107635. https://doi.org/10.1016/j.celrep.2020.107635
Wilson MA et al (2013) Pif1 helicase and Polδ promote recombination-coupled DNA synthesis via bubble migration. Nature 502:393–396. https://doi.org/10.1038/nature12585
Zellweger R et al (2015) Rad51-mediated replication fork reversal is a global response to genotoxic treatments in human cells. J Cell Biol 208:563–579. https://doi.org/10.1083/jcb.201406099
Zeman MK, Cimprich KA (2014) Causes and consequences of replication stress. Nat Cell Biol 16:2–9. https://doi.org/10.1038/ncb2897
Zhang JM et al (2014) Fission yeast Pxd1 promotes proper DNA repair by activating Rad16 XPF and inhibiting Dna2. PLoS Biol 12:e1001946. https://doi.org/10.1371/journal.pbio.1001946
Zheng L, Meng Y, Campbell JL, Shen B (2020) Multiple roles of DNA2 nuclease/helicase in DNA metabolism, genome stability and human diseases. Nucleic Acids Res 48:16–35. https://doi.org/10.1093/nar/gkz1101
Acknowledgements
Work in the Rass laboratory is supported by the UK Government’s Department of Business, Energy and Industrial Strategy through the Academy of Medical Sciences Professorship Scheme.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by M. Kupiec.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Appanah, R., Jones, D., Falquet, B. et al. Limiting homologous recombination at stalled replication forks is essential for cell viability: DNA2 to the rescue. Curr Genet 66, 1085–1092 (2020). https://doi.org/10.1007/s00294-020-01106-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00294-020-01106-7