Abstract
Severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) has spread all over the world and became a pandemic that named coronavirus disease-2019 (COVID-19). At present, several intramuscular vaccines have been successfully developed and mass vaccination has progressed in many countries. The aim of the study is to develop and examine an oral vaccine against COVID-19 with recombinant Lactococcus lactis IL1403, a strain of lactic acid bacteria, expressing SARS-CoV-2 spike (S) protein receptor-binding domain (RBD) S1 subunit as an immunizing antigen. PBS or cell extracts from recombinant L. lactis were orally administered into mice (control VS treatment), and formation of antigen-specific antibodies and changes in the gut microbiome were analyzed. Intracellular antigen was detected, but its secretion was not successful. After immunization, antigen-specific serum IgG and fecal IgA levels were 1.5-fold (P = 0.002) and 1.4-fold (P = 0.016) higher in the immunized mice (treatment) than control, respectively. Gut microbiome profiles were clearly separated between the two groups when analyzed for beta diversity with overall similarity. At the genus level, while Coprococcus (P = 0.036) and unclassified genus of Ruminococcaceae (P = 0.037) in treatment were more abundant than control, rc4-4 (P = 0.013) and Stenotrophomonas (P = 0.021) were less abundant. Our results indicate that cell extract containing SARS-CoV-2 antigen can induce mice to produce antigen-specific antibodies without overall changes in the gut microbiome. This strategy may be useful for the development of other oral viral vaccines.
Similar content being viewed by others
Introduction
The discovery of a novel coronavirus that the severe acute respiratory syndrome coronavirus 2 (SARS-CoV-2) in the human body in 2019 has gradually become a pandemic all over the world and named coronavirus disease-2019 (COVID-19) [1].
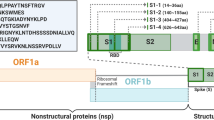
With the severity of the pandemic, many institutions began to develop several COVID-19 vaccines, such as recombinant protein [2, 3] and nucleic acid-based vaccine [4] for producing vaccine-induced neutralizing antibodies. SARS-CoV-2 mainly contains four protein structures including spike (S), membrane (M), envelop (E), and nucleocapsid (N) protein and these viral proteins may be potential to targets for vaccines to induce immune response [5]. Angiotensin-converting enzyme 2 (ACE2) is a cell entry receptor, while S protein of SARS-CoV-2 binding [6]. S protein is composed of S1 and S2 subunit, the S1 subunit recognizes the receptor site with receptor-binding domain (RBD) and the S2 subunit is responsible for membrane fusion. At present, many vaccines are developed with S1 subunit as the target. In other coronavirus research such as SARS and Middle East respiratory syndrome coronavirus (MERS-CoV), S protein S1 subunit binds to ACE2 receptor effectively [7, 8].
Several lactic acid bacteria (LAB) as probiotics be considered as carrier of oral vaccine candidates, these LAB strains originally live in the intestine of animals and can use plasmid vector system to produce heterologous protein. Recombinant LAB strains can elicit mucosal immune responses against selected antigens [9]. In anti-food-and-mouth disease virus (anti-FMDV) research, mice that immunized with that producing FMDV antigen recombinant Lactococcus lactis induced high levels of neutralizing antibodies [10]. In other research, oral immunization with purified Brachyspira membrane protein B from recombinant E. coli can produce antigen-specific serum IgG and fecal IgA in mice [11], and it can provide a certain amount of antigen cause immune response. Probiotics themselves are beneficial for the host, and purification process is not required for such immunization when recombinant LAB vaccines are used [12]. Limited studies have developed oral vaccines from the cell extracts of lactic acid bacteria.
With the advantages of next-generation sequencing (NGS) technology, it is easy to understand gut microbiome of animals. At present, many studies have focused on the changes of gut microbiome after the intake of live probiotics and not clear the changes of gut microbiome after intake of probiotics cell extracts and probiotic-based oral vaccine [13, 14]. Therefore, it is necessary to study whether probiotics cell extracts affect the gut microbiome, and what kind of changes do the gut microbiome have when vaccine of probiotics cell extracts enter the intestine.
In this study, we evaluated the impact of oral COVID-19 vaccine of recombinant L. lactis cell extracts on the immune response and profiling the gut microbiome of immunized mice.
Materials and Methods
Microorganism Strain and Growth Condition
L. lactis IL1403 was used as host strain and grown in M17 medium (MBcell, Korea) supplemented with 5 g/L of glucose (M17G) without antibiotics for wild type, and recombinant L. lactis IL1403 was grown in M17G media with erythromycin (5 µg/mL) and chloramphenicol (5 µg/mL) at 30 ℃.
Gene Synthesis and Plasmid Construction
Based on SARS coronavirus (GenBank: YP_009825051.1) [7] research, Fig. S2 shows using alignment method to obtain S1 subunit target of SARS-CoV-2 surface glycoprotein sequence (GenBank: YP_009724390.1) [15]. For detecting the expressed antigen, His-tag was added C-terminal. In order to make the recombinant L. lactis secrete the target protein, signal peptide of USP45 [16] was added N-terminal of the target protein (Fig. 1a). Codon optimization was conducted in DNAWorks v3.2.4 [17] based on L. lactis Il1403 codon usage table, and 33 primers were used to synthesize the insert sequence with overlap PCR method (Table S1). Plasmid DNA pILPtuf.Mb vector was used as a backbone [18], and AseI and XhoI restriction enzyme sites of target insert were ligated into NdeI and XhoI of vector backbone and transformed into L. lactis IL1403 competent cells.
Production and secretion of target antigen from recombinant L. lactis. a Schematic diagram for construction of pILPtuf.nCoV.h vector (modified from E.B. Kim et al., 2009). b Western blot for detecting SARS-CoV2 S protein RBD S1 subunit antigen from recombinant L. lactis. Target antigen was detected in intracellular recombinant L. lactis and not detected in wild-type L. lactis IL1403 (intracellular) and cultured broth of recombinant L. lactis. Lane1: L. lactis IL1403 WT; Lane2: L. lactis IL1403 (pILPtuf.nCoV.h); Lane3: L. lactis IL1403 (pILPtuf.nCoV.h) cultured broth (Cell free); Lane4–6: Commercial His-tagged Calmodulin (18 kDa) 1.5, 1, and 0.5μg, respectively
SDS-PAGE and Western Blot Assay
Wild type and recombinant L. lactis IL1403 were grown in M17G (50 mL) with or without antibiotics at 30 ℃ for 10 h. Total 50 mL of 1.5 × 1010 colony-forming unit (CFU) cell extracts were collected by centrifugation at 4000 rpm for 10 min and broken in bead beater with 0.5 g glass beads (0.5 mm) and 200 µL sterilized 1 × phosphate -buffered saline (PBS) and 3.5 µL extraction solution was mixed by 1 × PBS and 5 × loading dye to load. For preparation of extracellular protein, after above centrifugation, 40 mL of cell-cultured supernatant was filtered by 0.2 µm filter and precipitated by 16% trichloroacetic acid (TCA), and the precipitants were washed with ethanol and dissolved in 200 µL 1 × PBS and 24 µL extraction solution was mixed by 5 × loading dye to load. In order to quantify the protein production, commercial recombinant His-tagged human Calmodulin (MERCK, Darmstadt, Germany) of 18 kDa protein was used as the standard curve with 1.5, 1, and 0.5 µg.
The total cell extracts and cell-free supernatant extracts were separated by SDS-PAGE and transferred on to 0.2 µm nitrocellulose membrane. The membrane was blocked with 5% (w/v) skim milk with 1 × TBST (Tris-Buffered Saline, 0.1% Tween) at room temperature (RT) for 1 h. Blocked membrane was washed by 1 × TBST three times and incubated with anti-His6x monoclonal antibody (1:500, R&D Systems, USA) at 4 ℃ overnight with shaking. After three times washed with 1 × TBST, the membrane was visualized with ECL reagents (Bio-Rad, USA).
Immunization of Mice
Four-week-old female BALB/c mice were orally administered PBS (control) or immunized (treatment) with including recombinant SARS-CoV-2 spike protein RBD cell extracts of L. lactis, five mice in each group. Before oral administration with PBS (control) or cell extracts (treatment), 300 µL of neutralizing reagent (1.5% NaH2CO3) was administered orally. Recombinant L. lactis was cultured in M17G broth with erythromycin (5 µg/mL) and chloramphenicol (5 µg/mL) for 10 h. In treatment group, the total cell extracts (including 219 µg of target antigen) from 3.0 × 1010 CFU were dissolved in 200 µL-sterilized 1 × PBS and fed to each mouse, and control group fed only 200 µL-sterilized 1 × PBS. According to immunization of mice researches, the immunization dosage of target antigen and schedule were referred to that study using crude antigen to induce mucosal immune response with oral administration [11, 19]. Mice were immunized totally six times, priming, 1st boosting and 2nd boosting each two times. Fig 2a shows the immunization schedule. After 3 weeks, for analyzing anti-SARS-CoV-2 S protein RBD S1 subunit-specific immunoglobulins, serum samples were obtained from ventricle after centrifugation and fresh feces were sampled. The body weight of mice was measured before (At day 1) and after (At day 22) the experiment to compare the weight gain between the two groups.
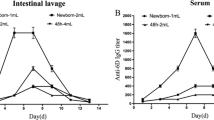
Humoral and mucosal immune response elicited by cell extracts of SARS-CoV-2 S protein RBD S1 subunit expressing recombinant L. lactis. a Schematic view of immunization test schedule (n = 5 in each group tested). b The levels of SARS-CoV-2 S protein RBD S1 subunit-specific serum IgG in the treatment group were 1.5-fold (P = 0.002) higher than control group at day 22. c The levels of SARS-CoV-2 S protein RBD S1 subunit-specific fecal IgA in the treatment group were 1.4-fold (P = 0.016) higher than control group at day 22. Control: Fed with PBS. Treatment: Fed with Cell extracts of SARS-CoV-2 S protein S1 subunit expressing recombinant L. lactis. For significance test, a Student’s t test was used and expressed as follows: *P < 0.05, **P < 0.01. The error bars on graphs represent the mean ± SD of data values
Enzyme-Linked Immunosorbent (Elisa) Assay
Recombinant human coronavirus SARS-CoV-2 spike glycoprotein S1 (GenBank: YP_009724390.1, Abcam, USA) was coated at 96-well plates (1 µg/well) for monitoring antigen-specific serum IgG and fecal IgA. Diluted serum (1:1000) or fecal (100 mg/mL PBS with protease inhibitor cocktail, 1:100) samples were loaded in each well and incubated at room temperature for 2 h. After three times washing with 1 × PBST (1 × PBS with 0.1% Tween) and diluted HRP-conjugated goat anti-mouse IgG or IgA was added to each well and incubated at room temperature for 1 h. TMB substrate buffer elicited HRP enzyme reaction and is stopped by H2SO4 stop solution. The ELISA results were expressed as optical density (OD) values using microplate reader at 450 nm.
DNA Extraction and Sequencing for Gut Microbiome
Genomic DNA was extracted from 20 mg each fecal sample with NucleoSpin Soil kit (Macherey–Nagel, Düren, Germany). After DNA extraction, the 16S ribosomal RNA (16S rRNA) V4 region was amplified by universal primer sets (Forward: 5′-GGACTACHVG GGTWTCTAAT-3′ and R: 5′-GTGCCAGCMGCCGCGGTA A-3′) [20] with Ex-taq polymerase (Takara, Shiga, Japan). After amplification of 16S rRNA V4 region, amplified DNA was normalized to 50 ng per sample using Spark 10 M multimode microplate reader (Tecan Group AG, Zurich, Switzerland). DNA library is constructed and sequenced by the Illumina MiSeq platform (eGenome, Inc., Korea) generating 2 × 250 bp paired-end reads.
Bioinformatic Analysis for Microbial Community
The quality trimming of raw reads was performed by in-house Perl script. After quality control process, microbiome analysis was using open-source bioinformatics pipeline that quantitative insights into microbial ecology (QIIME, http://qiime.org/index.html) version 1.9.1. Normalization was performed with assigned reads that 45,000 reads/per sample for comparison and clustered into operational taxonomic units (OTUs) and using the GreenGenes 13_8 database to pick OTUs with 97% similarity. According to picked OTUs, the relative abundances of phyla and genera were calculated. Alpha and beta diversity of gut microbiome were assessed by QIIME tool, respectively. Four of alpha diversity indices that observed OTUs, Chao1, phylogenetic diversity (PD) whole tree, and Shannon were assessed by rarefaction with ten iterations from 45,000 reads. The beta diversity was analyzed by UniFrac distances in QIIME.
Statistical Analysis
Statistical analysis was performed by R (version 4.1.0) language. For microbiome significance tests, a one-way analysis of variance (ANOVA) followed by Tukey’s post hoc test. A student’s t test was used for ELISA and gut microbiome significance tests.
Results
Production of a SARS-CoV-2 Antigen
In order to confirm the correct construction of the plasmid vector, we used Sanger sequencing (Macrogen, Inc., Korea) to detect the restriction mapping between the vector and the insert and insert sequencing. Sequencing result indicates the restriction mapping and insert sequencing were correct (Fig. S1). To examine production and secretion of SARS-CoV-2 antigen in recombinant L. lactis, we analyzed cell extracts and culture supernatants with western blot. Western blot result showed that there is no His-tagged protein in L. lactis IL1403 wild-type cells and there is His-tagged of 22.89 kDa protein in recombinant lactis (Fig. 1b). Although well-known signal peptide was located N-terminal of target protein and its secretion was not successful. To quantify the amount of expressed antigens, commercial His-tagged Calmodulin were used as standard curve. According to the standard curve, recombinant L. lactis produces his-tagged antigen of SARS-CoV-2 S protein RBD S1 subunit 2.19 µg/ml in M17G medium. These results demonstrated that the extracts of recombinant L. lactis contain the target antigens.
In vivo Evaluation of Recombinant SARS-CoV-2 Antigen
Next, the in vivo effect of the oral vaccine was examined. BALB/c mice were orally administered with the total cell extracts of recombinant L. lactis. The vaccination schedule is shown in Fig. 2a. On day 22 post-priming vaccination, the levels of antigen-specific serum IgG and fecal IgA were measured using ELISA to evaluate the systemic and mucosal immune responses. The levels of SARS-CoV-2 antigen-specific serum IgG in the treatment (OD450nm = 0.081 ± 0.009) group were 1.5-fold (P = 0.002) higher than those in the control (OD450nm = 0.054 ± 0.0008) group. Meanwhile, the levels of SARS-CoV-2 antigen-specific fecal IgA in the treatment (OD450nm = 0.375 ± 0.067) group were 1.4-fold (P = 0.016) higher than those in the control (OD450nm = 0.262 ± 0.033) group (Fig. 2b, c). The bodyweight gain during the experimental period (days 1–22) was monitored to evaluate the effect of the oral vaccine on the bodyweight of mice. The oral vaccine did not significantly affect the bodyweight of mice (Table S4). Thus, the oral vaccine elicited antigen-specific systemic and mucosal immune responses without significantly affecting the bodyweight.
Gut Microbial Diversity
To compare the bacterial diversity and communities, we investigated the alpha and beta diversity of two groups that control and treatment from normalized microbiome sequencing reads. In alpha diversity, we measured the four indices that observed OTUs, Chao1, PD whole tree and Shannon. All four indices showed no significant difference between the two groups (Fig. S3). In beta diversity, PCoA analysis of unweighted and weighted based on UniFrac distances. From the unweight result, the two groups are dispersed (Fig. S4a), and there was no difference between the two groups in the weighted result (Fig. S4b).
Gut Microbial Composition
In order to compare the difference of major gut microbial taxa between immunized and non-immunized groups, we examined the microbial composition in both archaea and bacteria with phylum and genus levels. Overall microbial composition in the gut was not so critically different between the two groups. However, certain microbial groups were significantly different. No significant differences in relative abundance at the phylum level were not found between control and treatment groups (Table S2). Relative abundance in genus of Archaea was not significantly different between the two groups (Table S3). In bacteria, compared with control group, Coprococcus and unclassified genus of Ruminococcaceae were significantly higher abundant in treatment group (P < 0.05) and rc4-4 and Stenotrophomonas were significantly less abundant in treatment group (P < 0.05) (Fig. 3). Moreover, the abundance of the genus Lactococcus, which was used as a host for the production of recombinant antigens, was not significantly different between the two groups (Table S3). These results demonstrated that oral vaccination with the cell extracts of lactic acid bacteria containing the recombinant antigen without overall alter the intestinal microbial community.
The relative abundance (%) of genus between control and treatment groups. a Coprococcus. b rc4_4. c Genus of Ruminococcaceae. d Stenotrophomonas. Control (n = 5): Fed with PBS. Treatment (n = 5): Fed with Cell extracts of SARS-CoV-2 S protein S1 subunit expressing recombinant L. lactis. For significance test, a Student’s t test was used. The error bars on graphs represent the mean ± SD of data values
Discussion
Because of the high infectivity of SARS-CoV-2 and vaccines may be the only public health measure against the pandemic. At present, there is no oral vaccine based on cell extracts of LAB. The purpose of this study is to verify whether the cell extracts containing antigen of SARS-CoV-2 S protein RBD S1 subunit from recombinant L. lactis can induce mucosal immunity to produce antigen-specific antibodies and profiling the changes of gut microbiome.
We used western blot assay to confirm whether the recombinant L. lactis produce antigen of SARS-CoV-2 S protein RBD S1 subunit. Although signal peptide of USP45 was located N-terminal of target protein to obtain secretory protein, His-tagged secretory protein was not detected in the cultured supernatant (Fig. 1b). Since the recombinant L. lactis do not secrete the target protein, we used cell extracts to immunize mice instead of using living modified organisms (LMO).
The S protein of the SARS-CoV-2 is composed of S1 and S2 that complete the initial binding with the virus and ACE2 that of cell surface receptor and then the virus enters the host cells [21]. Tai et al. research shows that the SARS-CoV RBD-specific antibodies could cross-react with SARS-CoV-2 RBD protein [22]. The alignment of two coronaviruses shows that S protein RBD of SARS-CoV and SARS-CoV-2 are similar (Fig. S2). In Wong et al. study shows the a 193 amino acid fragment of the SARS-CoV S protein (residues 381–510) bind with ACE2 more efficiently than full length of S1 domain [7]. In this study, we used 194 amino acid fragment of SARS-CoV-2 S protein RBD S1 subunit to produce antigen in recombinant L. lactis. In other research of producing neutralizing anti-SARS-CoV-2 RBD, SARS-CoV-2 S protein RBD (193 amino acid) proteins were expressed on surface of Saccharomyces cerevisiae, and antigen-specific antibodies were produced in mice after oral immunization with live-attenuated vaccine [23].
Some studies have shown that different gut microbiomes lead to different antibody production efficiencies of vaccines in human [24]. However, there are few studies on the changes of gut microbiome after vaccination. In this study, we used NGS to characterize how probiotic-based oral vaccine changed the gut microbiome. Cell extracts of probiotic-based oral vaccine did not change the diversity of gut microbiome. Unpublished studies from our laboratory shows cell extracts of wild-type L. lactis IL1403 did not affect genes expression in the intestine of mice. However, the expression of genes involved in the related specific protein pathway in feeding cell extracts of specific protein-expressing recombinant L. lactis group was higher than control group. This results show that the proteins fed with this form were not completely degraded when it passes through the esophagus and stomach of mice and has a certain effect on the intestine. Although there was no great change in gut microbiome after immunization, the abundances of four genera were significantly different compared with that of the control group, such as Coprococcus, rc4-4, genus of unclassified Ruminococcaceae, and Stenotrophomonas. Moreover, the abundance of the genus Lactococcus, which was used as a host for the production of recombinant antigens, was not significantly different between the two groups. This indicated that Lactococcus wild-type proteins did not elicit antibody responses. This may be because Lactococcus is a natural inhabitant of the mouse intestinal tract. Also, our results indicate that oral vaccine can induce antigen-specific immune response without critical changes in gut microbiome. It is not clear whether the changes of these four genera were caused by intestinal immune response. Ritzi et al. research showed that combination of probiotics and coccidiosis vaccines can effectively reduce the lesion score of small intestine [25]. Further studies are required to validate such beneficial effects of probiotic host cells for expression of viral antigens in oral vaccines. The intestinal expression of ACE2 affects the balance of gut microbiome [26]. The viral spike protein fragment may bind to ACE2 receptor in intestine when the mice were immunized orally. However, because the structure of mouse ACE2 is different from that of human, the viral spike protein could not bind to ACE2 receptor effectively [27]. This may be one of the reasons for the less changes in gut microbiome after oral immunization with corona spike protein-based antigen. The infection of corona virus is not only human, and it can infect the other animals, such as, dogs, cats, tigers, and lions [28]. Our results expected this strategy can be used in the prevention of other animal corona virus.
L. lactis is generally recognized as safe (GRAS) status by the Food and Drug Administration (FDA) [29]. L. lactis IL1403 is known as a strain widely used for recombinant protein production in laboratory [30], and there has been no evidence to date that metabolites of L. lactis IL1403 are toxic to experimental animals, which can directly use their cell extracts without purification process. Moreover, in this study, antigens produced from L. lactis induced immune response and did not cause critical changes in the gut microbiome in mice. These results demonstrate the advantages of probiotic-based vaccines.
In summary, the cell extracts of SARS-CoV-2 S protein RBD S1 subunit antigen expressing recombinant L. lactis induce mice to produce antigen-specific antibody, and there was no critical change in the gut microbiome after oral administration of the probiotic-based vaccine. This strategy may potentially be used in development of oral vaccines to induce humoral and mucosal immune responses.
Data Availability
Microbiome raw sequences were deposited in GenBank with the Accession Number PRJNA769238.
References
Helmy YA, Fawzy M, Elaswad A et al (2020) The COVID-19 pandemic: a comprehensive review of taxonomy, genetics, epidemiology, diagnosis, treatment, and control. J Clin Med 9:1225. https://doi.org/10.3390/jcm9041225
Pollet J, Chen W, Strych U (2020) Since January 2020 Elsevier has created a COVID-19 resource centre with free information in English and Mandarin on the novel coronavirus COVID-19. The COVID-19 resource centre is hosted on Elsevier Connect, the company ’ s public news and information
Wang M, Fu T, Hao J et al (2020) Since January 2020 Elsevier has created a COVID-19 resource centre with free information in English and Mandarin on the novel coronavirus COVID-19. The COVID-19 resource centre is hosted on Elsevier Connect, the company ’ s public news and information
Jackson LA, Anderson EJ, Rouphael NG et al (2020) An mRNA vaccine against SARS-CoV-2—preliminary report. N Engl J Med 383:1920–1931. https://doi.org/10.1056/nejmoa2022483
Dai L, Gao GF (2021) Viral targets for vaccines against COVID-19. Nat Rev Immunol 21:73–82. https://doi.org/10.1038/s41577-020-00480-0
Zhou P, Lou YX, Wang XG et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579:270–273. https://doi.org/10.1038/s41586-020-2012-7
Wong SK, Li W, Moore MJ et al (2004) A 193-amino acid fragment of the SARS coronavirus S protein efficiently binds angiotensin-converting enzyme 2. J Biol Chem 279:3197–3201. https://doi.org/10.1074/jbc.C300520200
Du L, Zhao G, Kou Z et al (2013) Identification of a receptor-binding domain in the S Protein of the novel human coronavirus middle east respiratory syndrome coronavirus as an essential target for vaccine development. J Virol 87:9939–9942. https://doi.org/10.1128/jvi.01048-13
Szatraj K, Szczepankowska AK, Chmielewska-Jeznach M (2017) Lactic acid bacteria—promising vaccine vectors: possibilities, limitations, doubts. J Appl Microbiol 123:325–339. https://doi.org/10.1111/jam.13446
Liu X, Qi L, Lv J et al (2020) The immune response to a recombinant Lactococcus lactis oral vaccine against foot-and-mouth disease virus in mice. Biotechnol Lett 42:1907–1917. https://doi.org/10.1007/s10529-020-02900-6
Kim JI, Park TE, Maharjan S et al (2015) Soluble RANKL expression in Lactococcus lactis and investigation of its potential as an oral vaccine adjuvant. BMC Immunol 16:1–11. https://doi.org/10.1186/s12865-015-0132-x
Taghinezhad-S S, Mohseni AH, Bermúdez-Humarán LG et al (2021) Probiotic-based vaccines may provide effective protection against covid-19 acute respiratory disease. Vaccines 9:1–21. https://doi.org/10.3390/vaccines9050466
Hemarajata P, Versalovic J (2013) Effects of probiotics on gut microbiota: mechanisms of intestinal immunomodulation and neuromodulation. Therap Adv Gastroenterol 6:39–51. https://doi.org/10.1177/1756283X12459294
Wieërs G, Belkhir L, Enaud R et al (2020) How Probiotics affect the microbiota. Front Cell Infect Microbiol. https://doi.org/10.3389/fcimb.2019.00454
Wu F, Zhao S, Yu B et al (2020) A new coronavirus associated with human respiratory disease in China. Nature 579:265–269. https://doi.org/10.1038/s41586-020-2008-3
van Asseldonk M, de Vos WM, Simons G (1993) Functional analysis of the Lactococcus lactis usp45 secretion signal in the secretion of a homologous proteinase and a heterologous α-amylase. MGG Mol Gen Genet 240:428–434. https://doi.org/10.1007/BF00280397
Hoover DM, Lubkowski J (2002) DNAWorks: an automated method for designing oligonucleotides for PCR-based gene synthesis. Nucleic Acids Res 30:1–7. https://doi.org/10.1093/nar/30.10.e43
Kim EB, Piao DC, Son JS, Choi YJ (2009) Cloning and characterization of a novel tuf promoter from lactococcus lactis subsp. Lactis IL1403. Curr Microbiol 59:425–431. https://doi.org/10.1007/s00284-009-9455-2
Li HS, Piao DC, Jiang T et al (2015) Recombinant interleukin 6 with M cell-targeting moiety produced in Lactococcus lactis IL1403 as a potent mucosal adjuvant for peroral immunization. Vaccine 33:1959–1967. https://doi.org/10.1016/j.vaccine.2015.02.061
Han GG, Lee JY, Jin GD et al (2018) Tracing of the fecal microbiota of commercial pigs at five growth stages from birth to shipment. Sci Rep 8:1–9. https://doi.org/10.1038/s41598-018-24508-7
Jackson CB, Farzan M, Chen B, Choe H (2022) Mechanisms of SARS-CoV-2 entry into cells. Nat Rev Mol Cell Biol 23:3–20. https://doi.org/10.1038/s41580-021-00418-x
Tai W, He L, Zhang X et al (2020) Characterization of the receptor-binding domain (RBD) of 2019 novel coronavirus: implication for development of RBD protein as a viral attachment inhibitor and vaccine. Cell Mol Immunol 17:613–620. https://doi.org/10.1038/s41423-020-0400-4
Gao T, Ren Y, Li S et al (2021) Immune response induced by oral administration with a Saccharomyces cerevisiae-based SARS-CoV-2 vaccine in mice. Microb Cell Fact 20:1–10. https://doi.org/10.1186/s12934-021-01584-5
Ciabattini A, Olivieri R, Lazzeri E, Medaglini D (2019) Role of the microbiota in the modulation of vaccine immune responses. Front Microbiol. https://doi.org/10.3389/fmicb.2019.01305
Ritzi MM, Abdelrahman W, Van-Heerden K et al (2016) Combination of probiotics and coccidiosis vaccine enhances protection against an Eimeria challenge. Vet Res 47:1–8. https://doi.org/10.1186/s13567-016-0397-y
Hashimoto T, Perlot T, Rehman A et al (2012) ACE2 links amino acid malnutrition to microbial ecology and intestinal inflammation. Nature 487:477–481. https://doi.org/10.1038/nature11228
Wan Y, Shang J, Graham R et al (2020) Receptor recognition by the novel coronavirus from wuhan: an analysis based on decade-long structural studies of SARS coronavirus. J Virol 94:1–9. https://doi.org/10.1128/jvi.00127-20
Sharun K, Dhama K, Pawde AM et al (2021) SARS-CoV-2 in animals: potential for unknown reservoir hosts and public health implications. Vet Q 41:181–201. https://doi.org/10.1080/01652176.2021.1921311
Song AAL, In LLA, Lim SHE, Rahim RA (2017) A review on Lactococcus lactis: from food to factory. Microb Cell Fact 16:1–15. https://doi.org/10.1186/s12934-017-0669-x
Tavares LM, de Jesus LCL, da Silva TF et al (2020) Novel strategies for efficient production and delivery of live biotherapeutics and biotechnological uses of lactococcus lactis: the lactic acid bacterium model. Front Bioeng Biotechnol 8:1–19. https://doi.org/10.3389/fbioe.2020.517166
Funding
This work was supported by the National Research Foundation of Korea (NRF, 2019R1A2C1009406). Biao Xuan was supported by the BK21 Plus Program from Ministry of Education.
Author information
Authors and Affiliations
Contributions
EBK conceived the original idea, funding acquisition, project administration, and supervision. EBK, BX, JP, and JHY were involved in DNA working and animal experiment. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical Approval
An animal experiment was approved by the Institutional Animal Care and Use Committee (IACUC Accept Number: KW-190605-1) in Kangwon National University.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
284_2022_2866_MOESM1_ESM.tif
Supplementary file1 (TIF 330 kb) Fig. S1 Sequence alignment between reference (synthetic gene) and pILPtuf.nCoV.h. pILPtuf.nCoV.h:Vector backbone (1-10bp), NdeI/AseI compatible site (11-16bp), start codon (17-19bp), usp45 (20-103bp), NdeI (104-106bp), SARS-CoV-2 S protein RBD S1 subunit (107-691bp), his-tag (692-709bp),stop codon (710-712bp), XhoI (713-718bp), and vector backbone (719-728bp).
284_2022_2866_MOESM2_ESM.tif
Supplementary file2 (TIF 97 kb) Fig. S2 Sequence alignment between SARS-CoV S protein (318-510bp) and SARS-CoV-2 S protein (331-524bp).
284_2022_2866_MOESM3_ESM.tif
Supplementary file3 (TIF 30 kb) Fig. S3 Alpha diversity of gut microbiomes between control and treatmentgroups. (a) Rarefaction analysis observed features (Number of operationaltaxonomic units), (b) Chao1, (c) phylogenetic diversity (PD whole tree), and(d) Shannon index. Control (n = 5): Fed with PBS. Treatment (n = 5): Fedwith Cell extracts of SARS-CoV-2 S protein S1 subunit expressingrecombinant L. lactis. For significance test, a Student’s t test was used. The error bars on graphs represent the mean ± SD of data values.
284_2022_2866_MOESM7_ESM.tif
Supplementary file7 (TIF 128 kb) Fig. S4 Principal coordinate analysis of the microbiota between control and treatment groups. (a) Unweighted and (b) weighted based on UniFrac distances. Subject color: blue, control(n = 5); red, treatment (n = 5).
Rights and permissions
About this article
Cite this article
Xuan, B., Park, J., Yoo, J.H. et al. Oral Immunization of Mice with Cell Extracts from Recombinant Lactococcus lactis Expressing SARS-CoV-2 Spike Protein. Curr Microbiol 79, 167 (2022). https://doi.org/10.1007/s00284-022-02866-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-022-02866-w