Abstract
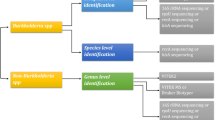
Burkholderia cepacia complex (Bcc) comprises 24 related species genetically distinct, associated with high mortality in cystic fibrosis (CF) patients. Due to a high level of similarity among Bcc species, accurate identification has been problematic, and most conventional and automated phenotypic tests have shown low accuracy. We evaluated accuracy of MALDI-ToF MS decreasing the cut-off score value to distinguish Bcc species compared to recA gene sequencing. A total of 145 Bcc isolates were analyzed. B. vietnamiensis (41.37%), B. cenocepacia IIIA (23.44%), B. multivorans (20%), B. cenocepacia IIIB (11.03%), and B. contaminans (2.75%) among other species were identified by recA sequencing. MALDI-ToF MS identified 100% of Bcc isolates at the genus level and 53.1% at the species level. By decreasing cut-off values for ≥1.70, the correct identification at the species level increased to 74.5%. MALDI-ToF MS proved to be useful at the genus level identification, but it still requires improvements that allow more precise identification, requiring continuous updates and addition of new spectra to its database. A review of interpretative criteria is a field to be explored with a large collection of Bcc species.

Similar content being viewed by others
References
Ragupathi NKD, Veeraraghavan B (2019) Accurate identification and epidemiological characterization of Burkholderia cepacia complex: an update. Ann Clin Microbiol Antimicrob 18(1):7. https://doi.org/10.1186/s12941-019-0306-0
Alexander BD, Petzold EW, Reller LB et al (2008) Survival after lung transplantation of cystic fibrosis patients infected with Burkholderia cepacia complex. Am J Transplant 8(5):1025–1030. https://doi.org/10.1111/j.1600-6143.2008.02186.x
Kenna DTD, Lilley D, Coward A, Martin K, Perry C, Pike R et al (2017) Prevalence of Burkholderia species, including members of Burkholderia cepacia complex, among UK cystic and non-cystic fibrosis patients. J Med Microbiol 66:490–501. https://doi.org/10.1099/jmm.0.000458
Furlan JPR, Pitondo-Silva A, Braz VS, Gallo IFL, Stehling EG (2019) Evaluation of different molecular and phenotypic methods for identification of environmental Burkholderia cepacia complex. J Microbiol Biotechnol 35(3):39. https://doi.org/10.1007/s11274-019-2614-0
Folescu TW, da Costa CH, Cohen RW et al (2015) Burkholderia cepacia complex: clinical course in cystic fibrosis patients. BMC Pulm Med 15:158. https://doi.org/10.1186/s12890-015-0148-2
Coenye T, Vandamme P, Govan JR, LiPuma JJ (2001) Taxonomy and identification of the Burkholderia cepacia complex. J Clin Microbiol 39:3427–3436. https://doi.org/10.1128/JCM.39.10.3427-3436.2001
Martina P, Leguizamon M, Prieto CI, Sousa SA, Montanaro P, Draghi WO et al (2018) Burkholderia puraquae sp. nov., a novel species of the Burkholderia cepacia complex isolated from hospital settings and agricultural soils. Int J Syst Evolut Microbiol 68(1):14–20. https://doi.org/10.1099/ijsem.0.002293
Jin Y, Zhou J, Zhou J et al (2020) Genome-based classification of Burkholderia cepacia complex provides new insight into its taxonomic status. Biol Direct 15(1):6. https://doi.org/10.1186/s13062-020-0258-5
Wolk DM, Clark AE (2018) Matrix-assisted laser desorption time of flight mass spectrometry. Clin Lab Med 38(3):471–486. https://doi.org/10.1016/j.cll.2018.05.008
Fehlberg LC, Andrade LH, Assis DM, Pereira RH, Gales AC, Marques EA (2013) Performance of MALDI-ToF MS for species identification of Burkholderia cepacia complex clinical isolates. Diagn Microbiol Infect Dis 77:126–128
Sambrook J et al (1982) Molecular cloning: a laboratory manual, vol 545. Cold Spring harbor laboratory, New York
Mahenthiralingam E et al (2000) DNA-based diagnostic approaches for identification of Burkholderia cepacia Complex, Burkholderia vietnamiensis, Burkholderia multivorans, Burkholderia stabilis, and Burkholderia cepacia Genomovars I and III. J Clin Microbiol 38:3165–3173
Wong KSK, Dhaliwal S, Bilawka J et al (2020) Matrix-assisted laser desorption/ionization time-of-flight MS for the accurate identification of Burkholderia cepacia complex and Burkholderia gladioli in the clinical microbiology laboratory. J Med Microbiol 69(8):1105–1113. https://doi.org/10.1099/jmm.0.001223
Sfeir MM (2018) Burkholderia cepacia complex infections: more complex than the bacterium name suggest. J Infect 77:166–170
De Dios J, Martínez CL, Tato M, Morosini MI, Cobo M et al (2016) Comparison between MALDI-TOF and recA gene sequencing for the identification of Burkholderia cepacia complex species isolated in a cystic fibrosis unit. J Cyst Fibros 15:S75
Vicenz FJ, Pillonetto M, Souza HA, Palmeiro JK, Riedi CA, RosarioFilho NA, Dalla-Costa LM (2016) Polyphasic characterisation of Burkholderia cepacia complex species isolated from children with cystic fibrosis. Mem Inst Oswaldo Cruz 111:37–42. https://doi.org/10.1590/0074-02760150314
Desai AP, Stanley T, Atuan M, McKey J, Lipuma JJ, Rogers B, Jerris R (2012) Use of matrix assisted laser desorption ionisation time of flight mass spectrometry in a paediatric clinical laboratory for identification of bacteria commonly isolated from cystic fibrosis patients. J Clin Pathol 65:835–838
Poonawala H, Marrs Conner T, Peaper DR (2018) The brief case: misidentification of Brucella melitensis as Ochrobactrum anthropi by matrixassisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS). J Clin Microbiol 56:e00914-e917. https://doi.org/10.1128/JCM.00914-17
Wong KSK, Dhaliwal S, Bilawka J et al (2020) Matrix-assisted laser desorption/ionization time-of-flight MS for the accurate identification of Burkholderia cepacia complex and Burkholderia gladioli in the clinical microbiology laboratory. J Med Microbiol. https://doi.org/10.1099/jmm.0.001223 (published online ahead of print, 2020 Jun 29)
Clinical CLSI, Institute LS (2017) Methods for the identification of cultured microorganisms using matrix-assisted laser desorption/ionization time-of-flight mass spectrometry, 1st edn. CLSI, Wayne
Barberis C, Almuzara M, Join-Lambert O, Ramírez MS, Famiglietti A, Vay C (2014) Comparison of the Bruker MALDI-TOF mass spectrometry system and conventional phenotypic methods for identification of gram-positive rods. PLoS ONE 9(9):e106303. https://doi.org/10.1371/journal.pone.0106303
Alatoom AA, Cazanave CJ, Cunningham SA, Ihde SM, Patel R (2012) Identification of non-diphtheriae corynebacterium by use of matrix-assisted laser desorption ionization-time of flight mass spectrometry. J Clin Microbiol 50(1):160–163. https://doi.org/10.1128/JCM.05889-11
Funding
This study was financed by the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq). Grant No. E-26/010.001040/2015.
Author information
Authors and Affiliations
Contributions
EFV, EAM, RSL, RMA Proposed the research. EFV, NIMM, LSSP, FADF, Prepared samples and performed experiments. EFV, EAM, CCFL, designed the experiments. EFV, FADF, RMA performed bio informatics analysis. EFV, Wrote manuscript and prepared figures. EAM, supervised the project. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical Approval
This study was approved by the Research Ethics Committee of our centre (University Hospital Pedro Ernesto, State University of Rio de Janeiro, Brazil), CAAE 16427019.8.0000.5259 (ref. 3.460.239).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Vianna, E.F., Pentagna, L.S.S., Menezes, N.I.M. et al. Decreasing the Cut-off Score Value of MALDI-ToF MS Increase the Identities of Burkholderia cepacia Complex Species. Curr Microbiol 78, 2259–2263 (2021). https://doi.org/10.1007/s00284-021-02493-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02493-x