Abstract
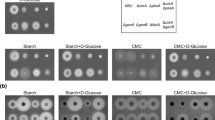
CpcA is a conserved transcriptional activator for the cross-pathway control of amino acid biosynthetic genes in filamentous fungi. Previous studies of this regulator mainly revealed its function under amino acid starvation condition, where amino acid biosynthetic inhibitors were added in the culture. In this study, the biological function of CpcA in Penicillium oxalicum was investigated under different cultivation conditions. Disruption of cpcA led to decreased cell growth either in the presence or absence of histidine biosynthetic inhibitor, and the phenotype could be rescued by the addition of exogenous amino acid sources. In addition, CpcA was required for the rapid production of cellulase when cells were cultured on cellulose. Transcript abundance measurement showed that a set of amino acid biosynthetic genes as well as two major cellulase genes were significantly down-regulated in cpcA deletion mutant relative to wild type. Taken together, the results revealed the biological role of CpcA in supporting normal growth and extracellular enzyme production of P. oxalicum under amino acid non-starvation condition.



Similar content being viewed by others
References
Hinnebusch AG (2005) Translational regulation of GCN4 and the general amino acid control of yeast. Annu Rev Microbiol 59:407–450
Paluh JL, Orbach MJ, Legerton TL, Yanofsky C (1988) The cross-pathway control gene of Neurospora crassa, cpc-1, encodes a protein similar to GCN4 of yeast and the DNA-binding domain of the oncogene v-jun-encoded protein. Proc Natl Acad Sci USA 85(11):3728–3732
Wanke C, Eckert S, Albrecht G, van Hartingsveldt W, Punt PJ, van den Hondel CA, Braus GH (1997) The Aspergillus niger GCN4 homologue, cpcA, is transcriptionally regulated and encodes an unusual leucine zipper. Mol Microbiol 23(1):23–33
Hoffmann B, Valerius O, Andermann M, Braus GH (2001) Transcriptional autoregulation and inhibition of mRNA translation of amino acid regulator gene cpcA of filamentous fungus Aspergillus nidulans. Mol Bio Cell 12(9):2846–2857
Hinnebusch AG (1993) Gene-specific translational control of the yeast GCN4 gene by phosphorylation of eukaryotic initiation factor 2. Mol Microbiol 10(2):215–223
Kornitzer D, Raboy B, Kulka RG, Fink GR (1994) Regulated degradation of the transcription factor Gcn4. EMBO J 13(24):6021–6030
Ebbole DJ, Paluh JL, Plamann M, Sachs MS, Yanofsky C (1991) cpc-1, the general regulatory gene for genes of amino acid biosynthesis in Neurospora crassa, is differentially expressed during the asexual life cycle. Mol Cell Biol 11(2):928–934
Tian C, Kasuga T, Sachs MS, Glass NL (2007) Transcriptional profiling of cross pathway control in Neurospora crassa and comparative analysis of the Gcn4 and CPC1 regulons. Eukaryot Cell 6(6):1018–1029
Albrecht G, Mosch HU, Hoffmann B, Reusser U, Braus GH (1998) Monitoring the Gcn4 protein-mediated response in the yeast Saccharomyces cerevisiae. J Biol Chem 273(21):12696–12702
Yang R, Wek SA, Wek RC (2000) Glucose limitation induces GCN4 translation by activation of Gcn2 protein kinase. Mol Cell Biol 20(8):2706–2717
Liu G, Qin Y, Li Z, Qu Y (2013) Improving lignocellulolytic enzyme production with Penicillium: from strain screening to systems biology. Biofuels 4(5):523–534
Liu G, Zhang L, Wei X, Zou G, Qin Y, Ma L, Li J, Zheng H, Wang S, Wang C, Xun L, Zhao GP, Zhou Z, Qu Y (2013) Genomic and secretomic analyses reveal unique features of the lignocellulolytic enzyme system of Penicillium decumbens. PLoS ONE 8(2):e55185
Cullen D, Leong SA, Wilson LJ, Henner DJ (1987) Transformation of Aspergillus nidulans with the hygromycin-resistance gene, hph. Gene 57(1):21–26
Yu JH, Hamari Z, Han KH, Seo JA, Reyes-Dominguez Y, Scazzocchio C (2004) Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet Biol 41(11):973–981
Kubodera T, Yamashita N, Nishimura A (2002) Transformation of Aspergillus sp. and Trichoderma reesei using the pyrithiamine resistance gene (ptrA) of Aspergillus oryzae. Biosci Biotechnol Biochem 66(2), 404–406.
Gao L, Li Z, Xia C, Qu Y, Liu M, Yang P, Yu L, Song X (2017) Combining manipulation of transcription factors and overexpression of the target genes to enhance lignocellulolytic enzyme production in Penicillium oxalicum. Biotechnol Biofuels 10:100
Li ZH, Du CM, Zhong YH, Wang TH (2010) Development of a highly efficient gene targeting system allowing rapid genetic manipulations in Penicillium decumbens. Appl Microbiol Biotechnol 87(3):1065–1076
Vogel HJ (1956) A convenient growth medium for Neurospora (Medium N). Microb Genet Bull 13:42–43
Chevallet M, Luche S, Rabilloud T (2006) Silver staining of proteins in polyacrylamide gels. Nat Protoc 1(4):1852–1858
Ivanov IP, Wei J, Caster SZ, Smith KM, Michel AM, Zhang Y, Firth AE, Freitag M, Dunlap JC, Bell-Pedersen D, Atkins JF, Sachs MS (2017) Translation initiation from conserved non-AUG codons provides additional layers of regulation and coding capacity. MBio 8(3):e00844
Ellenberger TE, Brandl CJ, Struhl K, Harrison SC (1992) The GCN4 basic region leucine zipper binds DNA as a dimer of uninterrupted alpha helices: crystal structure of the protein-DNA complex. Cell 71(7):1223–1237
Fordyce PM, Gerber D, Tran D, Zheng J, Li H, DeRisi JL, Quake SR (2010) De novo identification and biophysical characterization of transcription-factor binding sites with microfluidic affinity analysis. Nat Biotechnol 28(9):970–975
Sun X, Liu Z, Qu Y, Li X (2008) The effects of wheat bran composition on the production of biomass-hydrolyzing enzymes by Penicillium decumbens. Appl Biochem Biotechnol 146(1–3):119–128
Li C, Yang Z, Zhang RH, Zhang D, Chen S, Ma L (2013) Effect of pH on cellulase production and morphology of Trichoderma reesei and the application in cellulosic material hydrolysis. J Biotechnol 168(4):470–477
He R, Ma L, Li C, Jia W, Li D, Zhang D, Chen S (2014) Trpac1, a pH response transcription regulator, is involved in cellulase gene expression in Trichoderma reesei. Enzyme Microb Technol 67:17–26
Christianson DD, Cavins JF, Wall JS (1965) Identification and determination of nonprotein nitrogenous substances in corn steep liquor. J Agric Food Chem 13(3):277–280
Liu G, Zhang L, Qin Y et al (2013) Long-term strain improvements accumulate mutations in regulatory elements responsible for hyper-production of cellulolytic enzymes. Sci Rep 3:1569
Carle-Urioste JC, Escobar-Vera J, El-Gogary S, Henrique-Silva F, Torigoi E, Crivellaro O, Herrera-Estrella A, El-Dorry H (1997) Cellulase induction in Trichoderma reesei by cellulose requires its own basal expression. J Biol Chem 272(15):10169–10174
Acknowledgements
This work was supported by the National Key R&D Program of China (grant number 2018YFA0900503), the National Natural Science Foundation of China (31700062), the Project funded by China Postdoctoral Science Foundation (2017M612260), and the Young Scholars Program of Shandong University (YSPSDU).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Pan, Y., Gao, L., Zhang, X. et al. The Role of Cross-Pathway Control Regulator CpcA in the Growth and Extracellular Enzyme Production of Penicillium oxalicum. Curr Microbiol 77, 49–54 (2020). https://doi.org/10.1007/s00284-019-01803-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-019-01803-8