Abstract
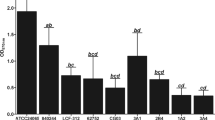
Bacterial biofilms can enhance survival in adverse environments and promote infection. However, little is known about biofilm formation by Clostridium perfringens. To better characterize this process, we used SEM to observe the surfaces of C. perfringens biofilms after 12, 24, 48, and 72 h of incubation. Biofilm cells appeared to be encased in a dense matrix material, and the total biomass of the biofilm increased with incubation time. To gain insight into the differentially expressed genes (DEGs) between biofilm and planktonic cells, we carried out comparative transcriptomic analysis using RNA sequencing. In total, 91 genes were significantly differentially expressed, with 40 being up-regulated and 51 down-regulated. In particular, genes encoding sortase, ribosomal proteins, and ATP synthase were up-regulated in biofilms, while genes coding for clostripain and phospholipase C were down-regulated. To validate the RNA sequencing results, qRT-PCR analysis was performed using five randomly selected DEGs. Results showed that all five genes were up-regulated, which was in accordance with the RNA sequencing results. To examine the functional differences, the DEGs were characterized by GO and KEGG pathway enrichment analyses. Results showed that the up-regulated genes were divided into 32 significantly enriched GO terms, with “macromolecular complex” being the most common. Oxidative phosphorylation was the only significantly enriched pathway, suggesting that ATP is required for biofilm stability. This study provides valuable insights into the morphology and transcriptional regulation of C. perfringens during biofilm formation, and will be useful for understanding and developing biofilm-based processes.






Similar content being viewed by others
References
Charlebois A, Jacques M, Boulianne M, Archambault M (2017) Tolerance of Clostridium perfringens biofilms to disinfectants commonly used in the food industry. Food Microbiol 62:32–38
Petit L, Gibert M, Popoff MR (1999) Clostridium perfringens: toxinotype and genotype. Trends Microbiol 7(3):104–110
Thomas MK, Murray R, Flockhart L, Pintar K, Pollari F, Fazil A, Nesbitt A, Marshall B (2013) Estimates of the burden of foodborne illness in Canada for 30 specified pathogens and unspecified agents, circa 2006. Foodborne Pathog Dis 10(7):639–648
Bryant AE (2003) Biology and pathogenesis of thrombosis and procoagulant activity in invasive infections caused by group A streptococci and Clostridium perfringens. Clin Microbiol Rev 16(3):451–462
Varga JJ, Therit B, Melville SB (2008) Type IV pili and the CcpA protein are needed for maximal biofilm formation by the gram-positive anaerobic pathogen Clostridium perfringens. Infect Immun 76(11):4944–4951
Davey ME, O’Toole GA (2000) Microbial biofilms: from ecology to molecular genetics. Microbiol Mol Biol Rev 64(4):847–867
Costerton JW, Stewart PS, Greenberg EP (1999) Bacterial biofilms: a common cause of persistent infections. Science 284(5418):1318–1322
Jorge EV, Joshua RS, Canizalez-Roman A (2015) The CpAL quorum sensing system regulates production of hemolysins CPA and PFO to build Clostridium perfringens biofilms. Infect Immun 83(6):2430–2442
Obana N, Nakamura K, Nomura N (2014) A Sporulation factor is involved in the morphological change of Clostridium perfringens biofilms in response to temperature. J Bacteriol 196(8):1540–1550
Høiby N, Bjarnsholt T, Givskov M, Molin S, Ciofu O (2010) Antibiotic resistance of bacterial biofilms. Int J Antimicrobial Agents 35(4):322–332
Costerton JW (2001) Cystic fibrosis pathogenesis and the role of biofilms in persistent infection. Trends Microbiol 9(2):50–52
Ha KY, Chung YG, Ryoo SJ (2005) Adherence and biofilm formation of Staphylococcus epidermidis and Mycobacterium tuberculosis on various spinal implants. Spine 30(1):38–43
Modi N, Wilcox MH (2001) Evidence for antibiotic induced Clostridium perfringens diarrhoea. J Clin Pathol 54(10):748–751
Charlebois A, Jacques M, Archambault M (2014) Biofilm formation of Clostridium perfringens and its exposure to low-dose antimicrobials. Front Microbiol 5:183
Charlebois A, Jacques M, Archambault M (2016) Comparative transcriptomic analysis of Clostridium perfringens biofilms and planktonic cells. Avian Pathol 45(5):593–601
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat methods 9(4):357–359
Wang L, Feng Z, Wang X, Wang X, Zhang X (2010) DEGseq: an R package for identifying differentially expressed genes from RNA-seq data. Bioinformatics 26(1):136–138
Young MD, Wakefield MJ, Smyth GK, Oshlack A (2010) Gene ontology analysis for RNA-seq: accounting for selection bias. Genome Biol 11(2):R14
Suzuki S, Tanigawa O, Akanuma G, Nanamiya H, Kawamura F, Tagami K, Nomura N, Kawabata T, Sekine Y (2014) Enhanced expression of Bacillus subtilis yaaA can restore both the growth and the sporulation defects caused by mutation of rplB, encoding ribosomal protein L2. Microbiology 160(Pt 6):1040–1053
Maracci C, Wohlgemuth I, Rodnina MV (2015) Activities of the peptidyl transferase center of ribosomes lacking protein L27. RNA 21(12):2047–2052
Schmohl L, Bierlmeier J, von Kügelgen N, Kurz L, Reis P, Barthels F, Mach P, Schutkowski M, Freund C, Schwarzer D (2017) Identification of sortase substrates by specificity profiling. Bioorg Med Chem 25(18):5002–5007
Jonsson IM, Mazmanian SK, Schneewind O, Bremell T, Tarkowski A (2003) The role of Staphylococus aureus sortase A and sortase B in murine arthritis. Microbes Infect 5(9):775–780
Mishra A, Devarajan B, Reardon ME, Dwivedi P, Krishnan V, Cisar JO, Das A, Narayana SV, Ton-That H (2011) Two autonomous structural modules in the fimbrial shaft adhesin FimA mediate Actinomyces interactions with streptococci and host cells during oral biofilm development. Mol Microbiol 81(5):1205–1220
Mishra A, Wu C, Yang J, Cisar JO, Das A, Ton-That H (2010) The Actinomyces oris type 2 fimbrial shaft FimA mediates co-aggregation with oral streptococci, adherence to red blood cells and biofilm development. Mol Microbiol 77(4):841–854
Okahashi N, Nakata M, Terao Y, Isoda R, Sakurai A, Sumitomo T, Yamaguchi M, Kimura RK, Oiki E, Kawabata S, Ooshima T (2011) Pili of oral Streptococcus sanguinis bind to salivary amylase and promote the biofilm formation. Microb Pathog 50(3–4):148–154
Metcalf DG, Bowler PG (2013) Biofilm delays wound healing: a review of the evidence. Burns Trauma 1(1):5–12
Saville RM, Rakshe S, Haagensen JA, Shukla S, Spormann AM (2011) Energy-dependent stability of Shewanella oneidensis MR-1 biofilms. J Bacteriol 193(13):3257–3264
Gilmore KS, Srinivas P, Akins DR, Hatter KL, Gilmore MS (2003) Growth, development, and gene expression in a persistent Streptococcus gordonii biofilm. Infect Immun 71(8):4759–4766
Stanley NR, Britton RA, Grossman AD, Lazazzera BA (2003) Identification of catabolite repression as a physiological regulator of biofilm formation by Bacillus subtilis by use of DNA microarrays. J Bacteriol 185(6):1951–1957
Acknowledgements
This work was supported by a grant from the National Natural Science Foundation of China (Grant Nos. 31460672, 31760739) and Qinghai High Level Talent Innovation Program.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zhang, X., Ma, Y. & Ye, G. Morphological Observation and Comparative Transcriptomic Analysis of Clostridium perfringens Biofilm and Planktonic Cells. Curr Microbiol 75, 1182–1189 (2018). https://doi.org/10.1007/s00284-018-1507-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-018-1507-z