Abstract
Background
The interplay between regulatory T cells (Tregs) and neighboring cells, which is pivotal for anti-tumor immunity and closely linked to patient prognosis, remains to be fully elucidated.
Methods
Tissue microarrays of 261 operable NSCLC patients were stained by multiplex immunofluorescence (mIF) assay, and the interaction between Tregs and neighboring cells in the tumor microenvironment (TME) was evaluated. Employing various machine learning algorithms, we developed a spatial immune signature to predict the prognosis of NSCLC patients. Additionally, we explored the interplay between programmed death-1/programmed death ligand-1 (PD-1/PD-L1) interactions and their relationship with Tregs.
Results
Survival analysis indicated that the interplay between Tregs and neighboring cells in the invasive margin (IM) and tumor center was associated with recurrence in NSCLC patients. We integrated the intersection of the three algorithms to identify four crucial spatial immune features [P(CD8+Treg to CK) in IM, P(CD8+Treg to CD4) in IM, N(CD4+Treg to CK) in IM, N(CD4+Tcon to CK) in IM] and employed these characteristics to establish SIS, an independent prognosticator of recurrence in NSCLC patients [HR = 2.34, 95% CI (1.53, 3.58), P < 0.001]. Furthermore, analysis of cell interactions demonstrated that a higher number of Tregs contributed to higher PD-L1+ cells surrounded by PD-1+ cells (P < 0.001) with shorter distances (P = 0.004).
Conclusion
We dissected the cell interplay network within the TME, uncovering the spatial architecture and intricate interactions between Tregs and neighboring cells, along with their impact on the prognosis of NSCLC patients.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
T cells, functioning as immune supervisors and effectors, play a critical role in anti-tumor immunity by engaging in intricate interactions with other cells [1,2,3]. Through cytokines or direct contact with other cells within the tumor microenvironment (TME), T cells orchestrate the modulation of immune cell activities and lead to the death of tumor cells [4,5,6]. The interplay between T cells and other cells residing in the TME has been identified as a crucial point in the eradication of tumor cells, with mounting research indicating its intimate correlation with patient prognosis [7,8,9]. Therefore, mapping the architecture of T cells in the TME has greatly gained importance.
Regulatory T cells (Tregs) constitute an important T cell subset and exert a vital role in immune regulation, and their interaction with surrounding cells can significantly influence patient prognosis [10,11,12,13]. Recent studies have utilized cell-to-cell distance to investigate these interactions. For example, Nagl et al. found that shorter distances between Tregs and CD8+ T cells correlated with improved prognosis in patients with esophageal cancer [14]. Additionally, Feichtenbeiner et al. proposed that cell interactions occur within a specific range, as the proximity of Tregs to CD8+ T cells within a distance range of 30–110 μm was associated with favorable prognosis in gastric cancer patients [15]. However, in the context of non-small cell lung cancer (NSCLC), the leading cause of cancer-related death, an understanding of the reciprocal interactions between Tregs and surrounding cells, is scarce. Furthermore, Tregs are supposed to influence the efficacy of anti-PD-(L)1 immunotherapy covering all stages of NSCLC with unclear mechanisms [16,17,18]. Therefore, unraveling the intricate web of interactions between Tregs and neighboring cells contributes to exploring fresh avenues for precise therapeutics in NSCLC patients.
Eventually, in pursuit of a deeper comprehension of the latent biological mechanisms underlying the prognostic impacts of varied spatial arrangements, we delved into the intricate interplay between Tregs and neighboring cells and unveiled novel spatial parameters intricately linked to the recurrence of NSCLC. Subsequently, we formulated a predictive instrument, the spatial immune signature (SIS), to predict recurrence-free survival (RFS) across different stages, demonstrating robust and consistent performance. Additionally, in combination with clinical staging, the SIS held the potential for clinical applicability. Furthermore, we explored the correlation between PD-1/PD-L1 interaction and Tregs, aiming to offer novel insights into treatment strategies for NSCLC patients.
Methods
Samples and preparation of tissue microarrays (TMAs)
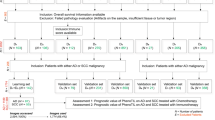
This study included patients with NSCLC who received radical surgery at the Shandong Cancer Hospital between January 2014 and October 2018. We implemented the eighth of the lung cancer classification for the identification of stages and the assessment of prognosis. The tissue samples of each patient who had not received neoadjuvant therapy were preserved using the formalin-fixed, paraffin-embedded (FFPE) method and only those with sufficient tissue remaining were used for TMA preparation. The two different cores of each TMA were chosen from two representative areas of the same patients containing the invasion margin (IM) and tumor center (TC). IM was delineated as the zone with a 1 mm radius, centered on the demarcation between malignant tissue and uninvolved tissue [19,20,21]. A total of 261 patients were ultimately included in the study excluding missing tissue and poor staining quality (Supplementary file 1: Fig. S1). Recurrence-free survival (RFS) was defined as the duration between the initial diagnosis and the first occurrence of recurrence or the conclusion of the follow-up period. This study was approved by the Ethics Committee of Shandong Cancer Hospital.
Multiplex immunofluorescence staining and imaging processing
The expression intensity and spatial distribution of FOXP3, CD4, CD8, and CK were visualized by mIF staining in Panel 1, and FOXP3, PD-1, PD-L1, and CK were shown in Panel 2. The TMA blocks containing the IM and TC areas were sectioned into 3-μm-thick slices. TMA slides were melted, subjected to 12 h of dehydration at 60 °C, and then underwent deparaffinization with xylene and graded alcohol rehydration. Next, the slides were placed in heat-induced antigen retrieval using either ethylene diamine tetra acid buffer (pH 8.0) or citric acid buffer (pH 6.0) in a microwave oven. Antibody concentrations and staining sequences were pre-optimized for this experiment. Following continuous incubation with primary antibodies and horseradish peroxidase-conjugated secondary antibodies, tyramide signal amplification (TSA) was performed. After each round of TSA, slides were subjected to heat for antigen retrieval and antibody stripping. After all staining series, the multi-stained slides were counterstained with 4',6-diamidino-2-phenylindole (DAPI) and scanned via the Vectra Polaris scanning system. Two expert pathologists, who were unaware of the patient information, evaluated all NSCLC samples and excluded inappropriate staining.
The stained slides were scanned by Vectra Polaris Scanner System, and the subsequent multispectral images were processed using Inform software 2.4.8. The inform software constructed a spectral library using single-stained slides for each tissue fluorescence spectrum and extracted the phenotyping algorithm from all spectrally unmixed images. Subsequently, the acquired images were recognized tumor or stroma compartments based on pan-CK labeled epithelial cell markers and analyzed cell segmentation according to the combination of the nuclear signals and cell morphology features. Detailed information about all primary antibodies and equipment used are provided in Supplementary file 1: Tables S1 and S2
Establishment of spatial immune characteristic parameters
We utilized mIF images and automated imaging processing system to identify various cell types and quantitatively analyzed cell density and spatial distribution within the TME (Supplementary file 2). Assuming A represented a specific cell type and B denoted another cell type, D(A) represented the density of A cells and D(B) represented the density of B cells in tissue site. Immune feature parameters of the mean nearest neighbor distance (mNND) and proximity score were established to explain the cell–cell interaction in the TME deciphered by the R software (version 3.6.3) phenoptrReports package, which was defined as follows:
The variable n signified the total cell count. The mNND calculated the average distance between all A cells and their closest neighboring B cells. Additionally, the expression \({d}_{min}\left({A}_{th}-B\right)\) referred to the minimum distance from each A cell to other B cells. A smaller value of \({N}_{(A to B)}\) indicated a stronger interaction between these cell types.
The proximity score was computed as the mean count of A cells within a specific radius surrounding all B cells throughout the tissue site. Additionally, N denoted the total cell count, and r indicated the specified radius. n(Cth \(\stackrel{r}{\to }\) A) represented the quantity of A cells within the r-um radius encircling each B cell. Therefore, an elevated proximity score for A cells signified an increased B cell density close to A cells within a defined distance, implying enhanced intratumoral infiltration.
Selection of crucial features and construction of a spatial immune signature
To obtain a more refined and stable model, the cohort of 261 NSCLC patients was randomly classified into training (N = 183) and calibration (N = 78) groups, maintaining a ratio of 7:3. Crucially, we meticulously maintained the equilibrium of clinical characteristics within the training and validation cohorts (Supplementary Table S3). In the training cohort, we adapted three types of machine learning algorithms, including least absolute shrinkage and selection operator (LASSO) regression, extreme gradient boosting (XGBoost), and random forest (RF), to select the most important relevant immune features in R (version 3.6.3). Afterward, the intersecting immune parameters among the LASSO, XGBoost, and RF were considered the most critical immune characteristics and were visualized. Then, the intersection-based importance outcomes and multivariate Cox regression coefficients were combined to construct the spatial immune signature (SIS). The final formula was calculated as follows: SIS = 1.25 × [P(CD8+Treg-CD4) in IM] + 0.70 × [P(CD8+Treg-CK) in IM] – 0.67 × [N(CD4+Treg-CK) in IM] – 0.56 × [N(CD4+Tcon-CK) in IM]. The performance of the SIS was evaluated using receiver-operating characteristic (ROC) curves and the associated areas under the ROC curve (AUCs).
Statistical analysis
Categorical data differences were assessed using the chi-square test and Fisher’s exact test. Continuous variable comparisons between two groups were conducted via the Mann–Whitney U test. Differences in RFS between the two groups were estimated using the log-rank test and the Kaplan–Meier method. The ROC and AUCs were implemented to evaluate the classification performance in the RFS of the model. The optimal cut-off value was determined using X-tile software (version 3.6.1). Data analysis and visualization were performed using SPSS (version 26.0), R (version 3.6.3), and GraphPad Prism (version 8.0). Statistical tests were two-sided, and a P value < 0.05 was considered significant.
Results
Patients clinicopathological characteristics
A total of 261 patients with stage I-III NSCLC who were recruited met the inclusion criteria among whom 103 (39.5%) cases relapsed during the follow-up time. The recurrence rates at 1, 3, and 5 years were 14.6%, 42.1%, and 62.5%, respectively. The majority of the cases were comprised of male (64.4%), lung adenocarcinoma (65.9%), and stage I (57.5%) patients among 261 patients with NSCLC, 165 (63.2%) patients received adjuvant chemotherapy, and 37 (14.2%) patients underwent adjuvant radiotherapy (Table 1).
Overview of the distribution of Tregs in the tumor microenvironment
mIF staining was used to visualize the real immune landscape, and inForm software was used to digitize the spatial features of diverse cells in whole TMA sections (Fig. 1a). We analyzed the composition of various cells in panel 1 (Fig. 1b) and identified Tregs based on the specific marker FOXP3. The results showed that Tregs were predominantly composed of CD4+Tregs (IM, 91.4%; TC, 93.3%) and CD8+Tregs (IM, 8.6%; TC, 6.7%) and a considerable proportion of CD4+T cells and CD8+T cells expressed FOXP3 in IM and TC (Fig. 1c, e). Then, we employed the survival analysis to explore the relationship between CD4+Tregs or CD8+Tregs and recurrence. We found the D(CD4+Tregs) in TC (65.03 vs. not reached, P = 0.011) and D(CD8+Tregs) in IM and TC (IM, 68.00 vs. not reached, P = 0.018; TC, 62.03 vs. not reached, P = 0.001) were associated with the better prognosis of NSCLC patients via the log-rank test and the Kaplan–Meier method (Supplementary file 1: Fig. S2). The results from the univariate Cox analysis are similar to the aforementioned findings (Supplementary file 1: Table S4). Of note, in multivariate Cox regression, the interaction of PD-L1/PD-1 appears no significant relationship with the recurrence of NSCLC after adjusting for clinicopathological factors (including age, gender, histological type, smoking index and stage).
Interaction between Tregs and neighboring cells was associated with the RFS of patients with NSCLC
Firstly, we developed the mNND parameter to quantify the strength of cell–cell interactions, with a shorter mNND indicating a more robust interaction between cells (Fig. 2a). Patients with more advanced clinical stages showed shorter distances between CD4+Tregs and tumor cells in IM and TC, and CD4+T cells in IM, while the distance between CD4+Tregs and CD8+T cells in IM and TC was prolonged. The distance between CD8+Tregs and tumor cells or CD4+T cells in IM and TC decreased with the advancing TNM staging, while the distance between CD8+Tregs and CD8+T cells in IM and TC increased (all P < 0.05; Supplementary file 1: Table S5). In addition, we observed no significant difference in the interaction between CD4+Tregs and tumor cells or T cells compared with CD4+T conventional cells (Tcons) in TME (Supplementary file 1: Fig. S3). Additionally, CD8+Tregs were observed to be more distant from tumor cells, while being closer to CD4+T cells and CD8+T cells in IM and TC (Fig. 2b–d). Moreover, the distance between CD4+Tregs (P < 0.001) or CD8+Tregs (P = 0.027) to tumor cells was longer in IM than in TC (Supplementary file 1: Fig. S4). Subsequently, we examined the impact of interactions between Tregs and surrounding cells on RFS in patients with NSCLC (Fig. 2e–p). The results demonstrated that shorter distances between CD4+Tregs and tumor cells in IM and TC (IM, 55.50 vs. not reached, P < 0.001; TC, 33.10 vs. not reached, P < 0.001), or between CD4+Tregs and CD4+ T cells in IM (94.47 vs. not reached, P = 0.023), were associated with poorer patient outcomes, which were in accordance with the mNND from the CD8+Treg to tumor cells in IM and TC (IM, 55.50 vs. not reached, P = 0.001; TC, 65.03 vs. not reached, P = 0.001), and to CD4+T cells in IM (20.30 vs. not reached, P = 0.001). Conversely, a shorter distance between CD4+Tregs (IM, not reached vs. 65.03, P = 0.031; TC, not reached vs. 58.90, P = 0.023) or CD8+Tregs (IM, not reached vs. 62.50, P = 0.011; TC, not reached vs. 94.47, P = 0.044) and CD8+T cells in IM and TC were correlated with longer RFS.
The association of mNND between Tregs and neighboring cells with RFS in NSCLC patients. a Illustration of the mNND involving Tregs and neighboring cells. b–d The discrepancy of the mNND from CD8+Tregs and CD8+Tcons to neighboring cells in IM and TC. e–p Kaplan–Meier curves of RFS based on the mNND of Tregs to neighboring cells. Comparison statistical analysis was determined by the Mann–Whitney U test, and cumulative RFS were calculated by log-rank test
Secondly, the proximity score was also applied to reflect the interactions between cells. A higher proximity score indicates that the core cell is surrounded by more neighboring cells, implying stronger interactions between the central cell and its surroundings (Fig. 3a). With the TNM staging, CD4+Tregs or CD8+Tregs were surrounded with more tumor cells in IM and TC and CD4+T cells in IM. In contrast, CD4+Tregs surrounded with more CD8+T cells in IM and TC and CD8+Tregs surrounded with more CD8+T cells in TC (all P < 0.05; Supplementary file 1: Table S6). We compared the spatial characteristics between Tregs and Tcons and the results showed that CD4+Tregs in TC and CD8+Tregs in both IM and TC were surrounded by more tumor cells (Fig. 3b, c). Moreover, CD8+Tregs had more CD4+T cells and CD8+T cells in IM and TC (Fig. 3d, e). Interestingly, the proximity score of CD4+Tregs to tumor cells, CD4+T cells, and CD8+T cells were similar to CD4+Tcons (Supplementary file 1: Fig. S5). And there were more tumor cells surrounded CD4+Tregs (P = 0.005) and CD8+Tregs (P = 0.007) in TC than in IM (Supplementary file 1: Fig. S6). Similarly, we analyzed the relationship between the proximity score of Tregs to surrounding cells and patient prognosis. The survival analysis suggested that a higher number of tumor cells around CD4+Tregs (IM, 51.33 vs. no reached, P = 0.020; TC, 39.93 vs. not reached, P < 0.001) and CD8+Tregs (IM, 46.93 vs. not reached, P = 0.002; TC, 45.03 vs. not reached, P < 0.001) showed a lower RFS both in IM and TC (Fig. 3f–i). In contrast, increased CD4+T cells around CD4+Tregs (not reached vs. 54.03, P = 0.005) and CD8+Tregs (not reached vs. 46.80, P = 0.003) in TC were associated with better patient prognosis. Interestingly, in IM, patients with a higher amount of CD4+T cells around CD8+Tregs had a statistical correlation with worse RFS (39.93 vs. not reached, P = 0.001; Fig. 3j–m). Additionally, a higher number of CD8+T cells around CD4+Tregs in both IM and TC (IM, not reached vs. 29.93, P < 0.001; TC, not reached vs. 26.23, P < 0.001), as well as around CD8+Tregs in TC (not reached vs. 65.03, P = 0.009), were all correlated with longer RFS in patients (Fig. 3n–q).
The association of proximity between Tregs and neighboring cells with RFS in NSCLC patients. a Illustration of the proximity score involving Tregs and neighboring cells. b–e The discrepancy of the proximity score from Tregs and Tcons to neighboring cells in IM and TC. f–q Kaplan–Meier curves of RFS based on the proximity score of Tregs to neighboring cells. Comparison statistical analysis was determined by the Mann–Whitney U test and cumulative RFS was calculated by log-rank test. NS, no significance
Tregs were correlated with the interplay between PD-1/PD-L1
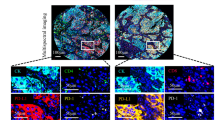
Our above findings revealed a close association between Tregs and tumor cells concerning the prognosis of NSCLC patients and current research unveiled the interaction of PD-1/PD-L1 influencing the proliferation and function of FOXP3+ Treg cells [22,23,24,25], which could offer novel insights to enhance the efficacy of anti-PD-1/PD-L1 antibody treatments. Using another mIF panel (Fig. 4a), our investigation revealed a direct relationship wherein an elevated quantity of FOXP3+ cells corresponded to an increased abundance of PD-1 + cells and PD-L1 + cells (Supplementary file 1: Fig. S7). Notably, we compared the differences in spatial interactions between PD-1/PD-L1 within the high FOXP3 + cells group and the low FOXP3+ cells group. Our results manifested that high FOXP3+ cells contributed higher proximity score of PD-L1 + cells to PD-1 + cells (P < 0.001) and shorter mNND of PD-L1 + cells to PD-1 + cells (P = 0.004), which reflects the strong interplay between the PD-1/PD-L1 (Fig. 4b, c). We examined the impact of interactions between PD-L1 and PD-1 cells on RFS in patients with NSCLC. The Kaplan–Meier analysis showed that the low N(PD-L1 to PD-1) in TC and high P(PD-L1 to PD-1) in TC (46.93 vs. not reached, P < 0.001) were associated with favorable prognosis in patients with NSCLC (Fig. 4d–g). By univariate Cox regression, we found the high P(PD-L1 to PD-1) in TC predicted longer RFS [HR = 0.68 (0.46, 0.99), P = 0.043, Supplementary file 1: Table S7].
Discrepancy of the interplay of PD-1/PD-L1 between FOXP3 high expression group and FOXP3 low expression group. a Representative composite and single-stained images of Panel 2. b mNND from PD-L1+ cells to PD-1+ cells in IM and TC. c Proximity score from PD-L1+ cells to PD-1+ cells in IM and TC. d–g Kaplan–Meier curves of RFS based on the interaction of PD-L1/PD-1. r equated 30 μm. Significance (P value) was determined by the Mann–Whitney U test
Spatial immune signature (SIS) contributed to predicting recurrence in patients with NSCLC
To comprehensively ascertain the effects of immune indicators on prognosis, we constructed spatial immune signature by three machine learning algorithms, including LASSO, XGBoost, and RF (Fig. 5a–c). We incorporated parameters including cell density and interaction of Tregs into the feature selection process of the model (Supplementary file 1:Table S8). The intersection of ten optimal variables with absolute values of nonzero coefficients selected by LASSO, and the top ten importance features extracted, respectively, by XGBoost and RF identified four critical immune parameters [P(CD8+Treg to CD4) in IM, P(CD8+Treg to CK) in IM, N(CD4+Treg to CK) in IM and N(CD4+Tcon to CK) in IM], which were applied to model construction (Fig. 5d). Univariate Cox regression manifested the higher P(CD8+Treg to CD4) in IM [HR = 1.87, 95%CI (1.25, 2.78), P = 0.002], P(CD8+Treg-CK) in IM [HR = 1.99, 95% CI (1.29, 3.06), P = 0.002] and shorter N(CD4+Treg-CK) in IM [HR = 0.45, 95% CI (0.30, 0.66), P < 0.001] and N(CD4+Tcon-CK) in IM [HR = 0.40, 95% CI (0.26, 0.63), P < 0.001] were associated with worse outcomes (Table 2).
Construction and validation of the spatial immune significance (SIS). a–c The important feature map was selected by three machine learning algorithms (Lasso, XGBoost and RF). d Venn diagram identified the four most critical spatial immune features. e–f ROC and AUCs of SIS for prediction of recurrence risk at 1, 3, and 5 years in the training (e), validation (f), and entire cohort (g). h–j Kaplan–Meier curves of RFS based on different SIS groups in the training (h), validation (i), and entire cohort. (j). Significance (P value) was determined using a Log-rank test
Then, the cases were split into a high-SIS group and a low-SIS group. Survival analysis showed that in the training (15.70 vs. not reached, P < 0.001), validation (55.50 vs. not reached, P = 0.003), and entire cohorts (27.70 vs. not reached, P < 0.001), the higher SIS group had a shorter RFS than the low-SIS group, respectively (Fig. 5e, f). Moreover, a multivariate Cox regression model of RFS was built based on NSCLC staging and SIS. Both NSCLC staging [HR = 2.26, 95% CI (1.51, 3.39), P < 0.001] and SIS [HR = 2.34, 95% CI (1.53, 3.58), P < 0.001] were significantly associated with RFS in NSCLC (Table 3). Additionally, we estimated its effectiveness by time-dependent ROC analysis. In the training set, the AUC values for 1-, 3-, and 5-year RFS were 0.797, 0.747, and 0.698, respectively, while in the validation set, they were 0.642, 0.685, and 0.704, respectively (Fig. 5h, i). Additionally, in terms of the entire dataset, the AUCs for predicting recurrence at 1, 3, and 5 years were 0.715, 0.695, and 0.676, respectively (Fig. 5j). Notably, the SIS system exhibited superior predictive capability compared to TNM staging and single indicators (Supplementary file 1: Figs. S8, S9). Collectively, the SIS consistently demonstrated stable and promising performance in distinguishing between NSCLC patients who experienced recurrence and those who remained recurrence-free after undergoing radical resection.
Discussion
Through the application of mIF in combination with spatial assays driven by artificial intelligence, we identified a spectrum of cell phenotypes and explored spatial interaction patterns within the TME across a cohort of 261 NSCLC cases. Furthermore, we established an SIS to predict the prognosis of NSCLC patients. Additionally, we utilized an alternative staining panel to investigate the impact of Tregs on the interaction between PD-1/PD-L1. Our findings provided valuable insights into the spatial architecture and diversity of Tregs within the TME of NSCLC, emphasizing the relationship between Tregs and surrounding cells, which might have implications for clinical outcomes and therapeutic strategies.
Our findings revealed that CD4+Tregs in TC as well as CD8+Tregs in IM and TC exhibited weaker interactions with surrounding tumor cells. Conversely, CD8+Tregs in IM and TC demonstrated stronger interactions with adjacent CD4+ T cells and CD8+ T cells. This phenomenon could potentially be attributed to the dynamic nature of the tumor microenvironment, as the immune milieu might be in an activated state of anti-tumor immunity during specimen collection. Subsequently, we conducted a survival analysis on the spatial characteristics of Tregs. The results revealed that strong interactions between Tregs and neighboring tumor cells in IM and TC were associated with adverse patient prognosis. Conversely, robust interactions between Tregs and adjacent CD4+T cells in TC and CD8+ T cells in IM and TC were correlated with favorable patient outcomes. These findings were consistent with prior research, where Tregs were known to foster an immunosuppressive niche that promoted tumor growth. Tregs could secrete inhibitory cytokines, such as interleukin-10 and transforming growth factor-β, attenuate the activity of nature killer (NK) cells and cytotoxic T cells and dampen the body’s anti-tumor immune surveillance [26,27,28,29,30,31]. Additionally, studies have shown that Tregs could exert direct cytolytic effects on NK cells and cytotoxic T cells through the action of granzyme B and perforin [31, 32]. Notably, strong interactions between CD4+Tregs in IM and TC and neighboring CD4+ T cells were associated with poor patient prognosis. This could potentially be attributed to the CD4+ T cells in this region being Th2 skewed, which migrates quickly and expresses elevated levels of anti-apoptotic signals, contributing to unfavorable patient outcomes [33,34,35,36]. These results underscored the significance of a precise analysis of short-range cell interactions in investigating the immunoregulatory role of Tregs.
Immunotherapies targeting NSCLC have garnered considerable attention in recent years, resulting in substantial advancements in patient outcomes. However, despite the considerable enthusiasm surrounding the success of these drugs, a significant proportion of patients fail to derive benefit from these treatments [37]. The expression of PD-L1 has limited predictive efficacy as an immunotherapy biomarker. Recent research on the interaction between PD-L1 and PD-1 has garnered significant attention. Sánchez-Magraner et al. reported that the interaction of the PD-1/PD-L1 but not the expression of PD-L1 is highly predictive of patient efficacy of immune treatment in NSCLC [38]. However, in our study, the interaction between PD-1 and PD-L1 in patients who did not receive immunotherapy appears to be unrelated to patient recurrence, suggesting that PD-L1/PD-1 interaction may only serve as a predictive marker of efficacy rather than prognosis. Therefore, future research should seek other more valuable biomarkers, aiming to guide patient stratification and treatment.
The application of immunotherapy has brought significant benefits to patients with NSCLC, and the impact of anti-PD-1/PD-L1 therapy on Tregs has also become a focal point of current research. Previous studies have demonstrated that the interaction between PD-1 and PD-L1 can promote the conversion of Th1 cells into inducible Tregs and selectively downregulate the expression of aminopeptidases, enhancing and sustaining FOXP3 expression on the surface of Tregs [39,40,41], which bolsters their immunosuppressive function. Our findings validated that the interaction between PD-1/PD-L1 is significantly stronger in the high FOXP3+ cells group, suggesting that this interaction fosters the proliferation and differentiation of Tregs, which is in line with prior research. Furthermore, in murine models, Yoshida et al. observed that ICI treatment blocking the PD-1/PD-L1 pathway effectively reduced the number of Tregs within tumors and increased tumor-infiltrating lymphocytes (TILs) [42]. Similarly, Toor et al. reported decreased FOXP3 expression on Treg cells’ surfaces in melanoma patients undergoing immunotherapy, consequently weakening the immunosuppressive function of Tregs [43]. However, Kamada et al. found that in patients receiving immunotherapy, tumor-infiltrating Tregs were significantly enhanced, leading to rapid tumor progression. The distinct regulatory effects of anti-PD-1/PD-L1 drugs on Tregs underscores the need for more in vivo and in vitro experiments to explore the impact of PD-1/PD-L1 interactions on Tregs [18]. This could potentially serve as a breakthrough in identifying efficacy markers for anti-PD-1/PD-L1 antibody therapy and target combination treatments contributing to precise therapeutic strategies for cancer patients.
Then, we created an SIS system that embraced diverse spatial immune features related to cell interactions, effectively portraying the intricate TME landscape and its influence on complex biological pathways. The SIS emerged as an autonomous and formidable prognostic marker for recurrence among patients with stages I-III NSCLC. It demonstrated notable precision in predicting recurrence outcomes across various TNM stages, aiding in the identification of individuals at heightened risk of early relapse. The SIS system put forth in the present investigation possessed a range of merits in contrast to alternative predictive frameworks. Firstly, the formulation of the SIS system integrated spatial immune features depicted through multi-color fluorescence staining which enabled a more faithful restoration of the TME and showcased commendable accuracy compared to features selected through solitary staining methods often employed in other studies. Secondly, due to the remarkable performance in clinical diagnosis and prognostication, diverse machine learning algorithms have gained extensive usage in predicting novel biomarkers and acquiring fresh insights. A combination of various machine learning offers an impartial approach to predicting patients’ clinical conditions [44,45,46].
Our study has several limitations. Firstly, our study lacked markers for other immune cell types within the TME, such as B cells, macrophages, and NK cells that could contribute to excavating more cell interaction network information to guide prediction and treatment. Second, we rarely detected the non-immune components in the TME to analyze these cells’ interaction with Tregs. Thirdly, our study was conducted in a single institute and external validation of the SIS system was lacking. Future investigations in multi-center settings involving diverse populations will be required to evaluate the robustness and credibility of our observations.
Future investigations ought to prioritize capturing the dynamic cellular composition and spatial architecture throughout the course of treatment, encompassing scenarios such as neoadjuvant and adjuvant immunotherapies. Moreover, the application of spatial transcriptomic and in vitro as well as in vivo experiments is imperative to meticulously unravel the intricate reconfiguration of the heterogeneous TME in response to diverse therapeutic interventions. In the context of the SIS system, forthcoming research endeavors should encompass a substantial sample size and treatment-related data to comprehensively assess its utility in predicting recurrence for patients afflicted with NSCLC.
Conclusion
We established a framework for scrutinizing the cellular interaction networks within the TME using mIF images, which are capable of comprehending the biological events where spatial interactions among cells determine functionality. The spatial landscape that mixes distributed patterns among Tregs, tumor cells, CD4+T cells, and CD8+T cells were underscored to help in the stratification of patients who might be at risk of early recurrence.
Availability of data and material
The datasets used and/or analyzed during the current study are available from the corresponding author upon reasonable request.
Abbreviations
- AUCs:
-
Associated areas under the receiver-operating characteristic curve
- CI:
-
Confidence interval
- FFPE:
-
Formalin-fixed, paraffin-embedded
- HR:
-
Hazard ratio
- IM:
-
Invasive margin
- LASSO:
-
Least absolute shrinkage and selection operator
- LUAD:
-
Lung adenocarcinoma
- LUSC:
-
Lung squamous cell carcinoma
- mIF:
-
Multiplex immunofluorescence
- mNND:
-
Mean nearest neighbor distance
- NK:
-
Nature killer
- NSCLC:
-
Non-small cell lung cancer
- PD-1:
-
Programmed death-1
- PD-L1:
-
Programmed death ligand-1
- RF:
-
Random forest
- RFS:
-
Recurrence-free survival
- ROC:
-
Receiver-operating characteristic
- ROI:
-
Region of interest
- SIS:
-
Spatial immune signature
- TC:
-
Tumor center
- Tcons:
-
Conventional T cells
- TMA:
-
Tissue microarray
- TME:
-
Tumor microenvironment
- Tregs:
-
Regulatory T cells
- XGBoost:
-
Extreme gradient boosting
References
Dunn GP, Old LJ, Schreiber RD (2004) The immunobiology of cancer immunosurveillance and immunoediting. Immunity 21:137–148. https://doi.org/10.1016/j.immuni.2004.07.017
Galon J, Angell HK, Bedognetti D, Marincola FM (2013) The continuum of cancer immunosurveillance: prognostic, predictive, and mechanistic signatures. Immunity 39:11–26. https://doi.org/10.1016/j.immuni.2013.07.008
Yu P, Fu YX (2006) Tumor-infiltrating T lymphocytes: friends or foes? Lab Investig 86:231–245. https://doi.org/10.1038/labinvest.3700389
Oliveira G, Wu CJ (2023) Dynamics and specificities of T cells in cancer immunotherapy. Nat Rev Cancer 23:295–316. https://doi.org/10.1038/s41568-023-00560-y
Alcover A, Alarcón B, Di Bartolo V (2018) Cell biology of T cell receptor expression and regulation. Annu Rev Immunol 36:103–125. https://doi.org/10.1146/annurev-immunol-042617-053429
Ostroumov D, Fekete-Drimusz N, Saborowski M, Kühnel F, Woller N (2018) CD4 and CD8 T lymphocyte interplay in controlling tumor growth. Cellular Mol Life Sci 75:689–713. https://doi.org/10.1007/s00018-017-2686-7
Borst J, Ahrends T, Bąbała N, Melief CJM, Kastenmüller W (2018) CD4(+) T cell help in cancer immunology and immunotherapy. Nat Rev Immunol 18:635–647. https://doi.org/10.1038/s41577-018-0044-0
Bedoui S, Heath WR, Mueller SN (2016) CD4(+) T-cell help amplifies innate signals for primary CD8(+) T-cell immunity. Immunol Rev 272:52–64. https://doi.org/10.1111/imr.12426
Vyas M, Requesens M, Nguyen TH, Peigney D, Azin M, Demehri S (2022) Natural killer cells suppress cancer metastasis by eliminating circulating cancer cells. Front Immunol 13:1098445. https://doi.org/10.3389/fimmu.2022.1098445
Piccirillo CA, Thornton AM (2004) Cornerstone of peripheral tolerance: naturally occurring CD4+CD25+ regulatory T cells. Trends Immunol 25:374–380. https://doi.org/10.1016/j.it.2004.04.009
Jiang M, Wu C, Zhang L et al (2021) FOXP3-based immune risk model for recurrence prediction in small-cell lung cancer at stages I-III. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-002339
Kinoshita T, Muramatsu R, Fujita T et al (2016) Prognostic value of tumor-infiltrating lymphocytes differs depending on histological type and smoking habit in completely resected non-small-cell lung cancer. Annals Oncol 27:2117–2123. https://doi.org/10.1093/annonc/mdw319
Moreno Ayala MA, Campbell TF, Zhang C, Dahan N, Bockman A, Prakash V, Feng L, Sher T, DuPage M (2023) CXCR3 expression in regulatory T cells drives interactions with type I dendritic cells in tumors to restrict CD8(+) T cell antitumor immunity. Immunity 56:1613–30.e5. https://doi.org/10.1016/j.immuni.2023.06.003
Nagl S, Haas M, Lahmer G, Büttner-Herold M, Grabenbauer GG, Fietkau R, Distel LV (2016) Cell-to-cell distances between tumor-infiltrating inflammatory cells have the potential to distinguish functionally active from suppressed inflammatory cells. Oncoimmunology 5:e1127494. https://doi.org/10.1080/2162402x.2015.1127494
Feichtenbeiner A, Haas M, Büttner M, Grabenbauer GG, Fietkau R, Distel LV (2014) Critical role of spatial interaction between CD8+ and Foxp3+ cells in human gastric cancer: the distance matters. Cancer Immunol Immunother 63:111–119. https://doi.org/10.1007/s00262-013-1491-x
Kumagai S, Koyama S, Itahashi K et al (2022) Lactic acid promotes PD-1 expression in regulatory T cells in highly glycolytic tumor microenvironments. Cancer Cell 40:201–18.e9. https://doi.org/10.1016/j.ccell.2022.01.001
Kumagai S, Togashi Y, Kamada T et al (2020) The PD-1 expression balance between effector and regulatory T cells predicts the clinical efficacy of PD-1 blockade therapies. Nat Immunol 21:1346–1358. https://doi.org/10.1038/s41590-020-0769-3
Kamada T, Togashi Y, Tay C et al (2019) PD-1(+) regulatory T cells amplified by PD-1 blockade promote hyperprogression of cancer. Proc Natl Acad Sci USA 116:9999–10008. https://doi.org/10.1073/pnas.1822001116
Mlecnik B, Bindea G, Kirilovsky A et al (2016) The tumor microenvironment and Immunoscore are critical determinants of dissemination to distant metastasis. Sci Transl Med. 8:32726. https://doi.org/10.1126/scitranslmed.aad6352
Donnem T, Kilvaer TK, Andersen S et al (2016) Strategies for clinical implementation of TNM-Immunoscore in resected nonsmall-cell lung cancer. Ann Oncol 27:225–232. https://doi.org/10.1093/annonc/mdv560
Carstens JL, Correa de Sampaio P, Yang D, Barua S, Wang H, Rao A, Allison JP, LeBleu VS, Kalluri R (2017) Spatial computation of intratumoral T cells correlates with survival of patients with pancreatic cancer. Nat Commun 8:15095. https://doi.org/10.1038/ncomms15095
Kornepati AVR, Vadlamudi RK, Curiel TJ (2022) Programmed death ligand 1 signals in cancer cells. Nat Rev Cancer 22:174–189. https://doi.org/10.1038/s41568-021-00431-4
Freeman GJ, Long AJ, Iwai Y et al (2000) Engagement of the PD-1 immunoinhibitory receptor by a novel B7 family member leads to negative regulation of lymphocyte activation. J Exp Med 192:1027–1034. https://doi.org/10.1084/jem.192.7.1027
Zou W, Wolchok JD, Chen L (2016) PD-L1 (B7–H1) and PD-1 pathway blockade for cancer therapy: mechanisms, response biomarkers, and combinations. Sci Transl Med 8:3284. https://doi.org/10.1126/scitranslmed.aad7118
Gong J, Chehrazi-Raffle A, Reddi S, Salgia R (2018) Development of PD-1 and PD-L1 inhibitors as a form of cancer immunotherapy: a comprehensive review of registration trials and future considerations. J Immunother Cancer 6:8. https://doi.org/10.1186/s40425-018-0316-z
Saito T, Nishikawa H, Wada H et al (2016) Two FOXP3(+)CD4(+) T cell subpopulations distinctly control the prognosis of colorectal cancers. Nat Med 22:679–684. https://doi.org/10.1038/nm.4086
Scott EN, Gocher AM, Workman CJ, Vignali DAA (2021) Regulatory T cells: barriers of immune infiltration into the tumor microenvironment. Front Immunol 12:702726. https://doi.org/10.3389/fimmu.2021.702726
deLeeuw RJ, Kost SE, Kakal JA, Nelson BH (2012) The prognostic value of FoxP3+ tumor-infiltrating lymphocytes in cancer: a critical review of the literature. Clin Cancer Res 18:3022–3029. https://doi.org/10.1158/1078-0432.Ccr-11-3216
Sawant DV, Yano H, Chikina M et al (2019) Adaptive plasticity of IL-10(+) and IL-35(+) T(reg) cells cooperatively promotes tumor T cell exhaustion. Nat Immunol 20:724–735. https://doi.org/10.1038/s41590-019-0346-9
Strauss L, Bergmann C, Szczepanski M, Gooding W, Johnson JT, Whiteside TL (2007) A unique subset of CD4+CD25highFoxp3+ T cells secreting interleukin-10 and transforming growth factor-beta1 mediates suppression in the tumor microenvironment. Clin Cancer Res 13:4345–4354. https://doi.org/10.1158/1078-0432.Ccr-07-0472
Nishikawa H, Koyama S (2021) Mechanisms of regulatory T cell infiltration in tumors: implications for innovative immune precision therapies. J Immunother Cancer. https://doi.org/10.1136/jitc-2021-002591
Cao X, Cai SF, Fehniger TA, Song J, Collins LI, Piwnica-Worms DR, Ley TJ (2007) Granzyme B and perforin are important for regulatory T cell-mediated suppression of tumor clearance. Immunity 27:635–646. https://doi.org/10.1016/j.immuni.2007.08.014
Frafjord A, Buer L, Hammarström C, Aamodt H, Woldbæk PR, Brustugun OT, Helland Å, Øynebråten I, Corthay A (2021) the immune landscape of human primary lung tumors is Th2 skewed. Front Immunol 12:764596. https://doi.org/10.3389/fimmu.2021.764596
Zhang X, Brunner T, Carter L et al (1997) Unequal death in T helper cell (Th)1 and Th2 effectors: Th1, but not Th2, effectors undergo rapid Fas/FasL-mediated apoptosis. J Exp Med 185:1837–1849. https://doi.org/10.1084/jem.185.10.1837
Fang Y, Yu S, Ellis JS, Sharav T, Braley-Mullen H (2010) Comparison of sensitivity of Th1, Th2, and Th17 cells to Fas-mediated apoptosis. J Leukoc Biol 87:1019–1028. https://doi.org/10.1189/jlb.0509352
Huang D, Chen X, Zeng X et al (2021) Targeting regulator of G protein signaling 1 in tumor-specific T cells enhances their trafficking to breast cancer. Nat Immunol 22:865–879. https://doi.org/10.1038/s41590-021-00939-9
Acúrcio RC, Pozzi S, Carreira B et al (2022) Therapeutic targeting of PD-1/PD-L1 blockade by novel small-molecule inhibitors recruits cytotoxic T cells into solid tumor microenvironment. J Immunother Cancer. https://doi.org/10.1136/jitc-2022-004695
Sánchez-Magraner L, Gumuzio J, Miles J et al (2023) Functional engagement of the PD-1/PD-L1 complex but not PD-L1 expression is highly predictive of patient response to immunotherapy in non-small-cell lung cancer. J Clin Onco 41:2561–2570. https://doi.org/10.1200/jco.22.01748
Duraiswamy J, Freeman GJ, Coukos G (2013) Therapeutic PD-1 pathway blockade augments with other modalities of immunotherapy T-cell function to prevent immune decline in ovarian cancer. Can Res 73:6900–6912. https://doi.org/10.1158/0008-5472.Can-13-1550
Jacobs JF, Idema AJ, Bol KF et al (2009) Regulatory T cells and the PD-L1/PD-1 pathway mediate immune suppression in malignant human brain tumors. Neuro Oncol 11:394–402. https://doi.org/10.1215/15228517-2008-104
Amarnath S, Mangus CW, Wang JC et al (2011) The PDL1-PD1 axis converts human TH1 cells into regulatory T cells. Sci Transl Med 3:111ra20. https://doi.org/10.1126/scitranslmed.3003130
Yoshida K, Okamoto M, Sasaki J et al (2020) Anti-PD-1 antibody decreases tumour-infiltrating regulatory T cells. BMC Cancer 20:25. https://doi.org/10.1186/s12885-019-6499-y
Toor SM, Syed Khaja AS, Alkurd I, Elkord E (2018) In-vitro effect of pembrolizumab on different T regulatory cell subsets. Clin Exp Immunol 191:189–197. https://doi.org/10.1111/cei.13060
Wang Z, Wang Y, Yang T et al (2021) Machine learning revealed stemness features and a novel stemness-based classification with appealing implications in discriminating the prognosis, immunotherapy and temozolomide responses of 906 glioblastoma patients. Brief Bioinform. https://doi.org/10.1093/bib/bbab032
Lai Y, Lin P, Lin F, Chen M, Lin C, Lin X, Wu L, Zheng M, Chen J (2022) Identification of immune microenvironment subtypes and signature genes for Alzheimer’s disease diagnosis and risk prediction based on explainable machine learning. Front Immunol 13:1046410. https://doi.org/10.3389/fimmu.2022.1046410
Errington N, Iremonger J, Pickworth JA et al (2021) A diagnostic miRNA signature for pulmonary arterial hypertension using a consensus machine learning approach. EBioMedicine 69:103444. https://doi.org/10.1016/j.ebiom.2021.103444
Acknowledgements
Not applicable.
Funding
This work was supported by grants from the National Natural Science Foundation of China (Grant No. 8217286 and No. 82373424.). The Natural Science Foudation of shandong Province (Grant No. ZR2021LZL005; No. ZR2019LZL019), the Department of Science & Technology of Shandong Province (Grant N0. 2021CXGC011102), and the Start-up Fund of Shandong Cancer Hospital (Grant N0. 2020PYA04).
Author information
Authors and Affiliations
Contributions
SC, GY, LY, WZ, and XS designed the study and developed the hypotheses. LY, WZ, and XS designed and performed multiplex immunofluorescence experiments. SC, GY, MH, and CL conducted data analysis. LY and XS mentored the data analysis. SC and XS interpreted the data. LY, WZ, and JS reviewed the pathological samples from all patients. SC, GY, LY, WZ, and FS assisted with the enrollment, evaluation, and clinical annotation of the patients described herein. SC wrote the manuscript. MH, CL, LY, WZ, LX, and XS revised the manuscript critically. LX and XS provided financial support. All authors reviewed and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethics approval and consent to participate
This study was approved by the Ethics Review Committee of Shandong Cancer Hospital (SDTHEC2022007013) and complied with the provisions of the Declaration of Helsinki.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cai, S., Yang, G., Hu, M. et al. Spatial cell interplay networks of regulatory T cells predict recurrence in patients with operable non-small cell lung cancer. Cancer Immunol Immunother 73, 189 (2024). https://doi.org/10.1007/s00262-024-03762-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00262-024-03762-x