Abstract
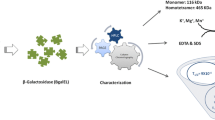
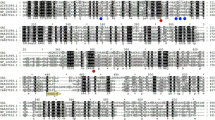
A novel β-galactosidase gene (galM) was cloned from an aquatic habitat metagenome. The analysis of its translated sequence (GalM) revealed its phylogenetic closeness towards Verrucomicrobia sp. The sequence comparison and homology structure analysis designated it a member of GH42 family. The three-dimensional homology model of GalM depicted a typical (β/α)8 TIM-barrel containing the catalytic core. The gene (galM) was expressed in a heterologous host, Escherichia coli, and the purified protein (GalM) was subjected to biochemical characterization. It displayed β-galactosidase activity in a wide range of pH (2.0 to 9.0) and temperature (4 to 60 °C). The heat exposed protein showed considerable stability at 40 and 50 °C, with the half-life of about 100 h and 35 h, respectively. The presence of Na, Mg, K, Ca, and Mn metals was favorable to the catalytic efficiency of GalM, which is a desirable catalytic feature, as these metals exist in milk. It showed remarkable tolerance of glucose and galactose in the reaction. Furthermore, GalM discerned transglycosylation activity that is useful in galacto-oligosaccharides’ production. These biochemical properties specify the suitability of this biocatalyst for milk and whey processing applications.
Key points
• A novel β-galactosidase gene was identified and characterized from an aquatic habitat.
• It was active in extreme acidic to mild alkaline pH and at cold to moderate temperatures.
• The β-galactosidase was capable to hydrolyze lactose in milk and whey.






Similar content being viewed by others

Data availability
The sequence of galM gene is available in the NCBI GenBank under the accession number OL412003.
References
Ambrogi V, Bottacini F, Cao L, Kuipers, B, Schoterman M, Van Sinderen D (2021) Galacto-oligosaccharides as infant prebiotics: production, application, bioactive activities and future perspectives. Crit Rev in Food Sci Nutr 1-14. https://doi.org/10.1080/10408398.2021.1953437
Cieśliński H, Kur J, Białkowska A, Baran I, Makowski K, Turkiewicz M, (2005) Cloning, expression, and purification of a recombinant cold-adapted β-galactosidase from Antarctic bacterium Pseudoalteromonas sp. 22b. Protein Expr Purif 39(1): 27–34. https://doi.org/10.1016/j.pep.2004.09.002
Coker JA, Sheridan PP, Loveland-Curtze J, Gutshall KR, Brenchley AAJ, JE, (2003) Biochemical characterization of a β-galactosidase with a low temperature optimum obtained from an Antarctic Arthrobacter isolate. J Bacteriol Res 185(18):5473–5482. https://doi.org/10.1128/JB.185.18.5473-5482.2003
Damin BIS, Kovalski FC, Fischer J, Piccin JS, Dettmer A (2021) Challenges and perspectives of the β-galactosidase enzyme. Appl Microbiol Biotechnol 105(13):5281–5298. https://doi.org/10.1007/s00253-021-11423-7
Dekker PJ, Koenders D, Bruins MJ (2019) Lactose-free dairy products: market developments, production, nutrition and health benefits. Nutrients 11(3):551. https://doi.org/10.3390/nu11030551
Deng Y, Misselwitz B, Dai N, Fox M (2015) Lactose intolerance in adults: biological mechanism and dietary management. Nutrients 7(9):8020–8035. https://doi.org/10.3390/nu7095380
Erich S, Kuschel B, Schwarz T, Ewert J, Böhmer N, Niehaus F, Eck J, Lutz-Wahl S, Stressler T, Fischer L (2015) Novel high-performance metagenome β-galactosidases for lactose hydrolysis in the dairy industry. J Biotechnol 210:27–37. https://doi.org/10.1016/j.jbiotec.2015.06.411
Fan F, Zhang L, Zhou X, Mu F, Shi G (2021) A sensitive fluorescent probe for β-galactosidase activity detection and application in ovarian tumor imaging. J Mater Chem B 9(1):170–175. https://doi.org/10.1039/D0TB02269A
Fan Y, Hua X, Zhang Y, Feng Y, Shen Q, Dong J, Zhao W, Zhang W, Jin Z, Yang R (2015) Cloning, expression and structural stability of a cold-adapted β-galactosidase from Rahnella sp. R3. Protein Expr Purif 115:158–164. https://doi.org/10.1016/j.pep.2015.07.001
Fan Y, Yi J, Hua X, Feng Y, Yang R, Zhang Y (2016) Structure analysis of a glycosides hydrolase family 42 cold adapted β-galactosidase from Rahnella sp. R3. RSC Adv 6(44):37362–37369. https://doi.org/10.1039/C6RA04529D
Gänzle MG, Haase G, Jelen P (2008) Lactose: crystallization, hydrolysis and value-added derivatives. Int Dairy J 18(7):685–694. https://doi.org/10.1016/j.idairyj.2008.03.003
Heine RG, AlRefaee F, Bachina P, De Leon JC, Geng L, Gong S, Madrazo JA, Ngamphaiboon J, Rogacion OC, JM, (2017) Lactose intolerance and gastrointestinal cow’s milk allergy in infants and children–common misconceptions revisited. World Allergy Organ J 10(1):1–8. https://doi.org/10.1186/s40413-017-0173-0
Hildebrandt P, Wanarska M, Kur J, (2009) A new cold-adapted β-D-galactosidase from the Antarctic Arthrobacter sp. 32c–gene cloning, overexpression, purification and properties. BMC Microbiol 9(1):1–11. https://doi.org/10.1186/1471-2180-9-151
Hoyoux A, Jennes I, Dubois P, Genicot S, Dubail F, François JM, Baise E, Feller G, Gerday C (2001) Cold-adapted β-galactosidase from the Antarctic psychrophile Pseudoalteromonas haloplanktis. Appl Environ Microbiol 67(4):1529–1535. https://doi.org/10.1128/aem.67.4.1529-1535.2001
Hu JM, Li H, Cao LX, Wu PC, Zhang CT, Sang SL, Zhang XY, Chen MJ, Lu JQ. Liu YH (2007) Molecular cloning and characterization of the gene encoding cold-active β-galactosidase from a psychrotrophic and halotolerant Planococcus sp. L4. J Agric Food Chem, 55(6):2217–2224. https://doi.org/10.1021/jf062910r
Kim CS, Ji ES, Oh DK (2003) Expression and characterization of Kluyveromyces lactis β-galactosidase in Escherichia coli. Biotechnol Lett 25(20):1769–1774. https://doi.org/10.1023/A:1026092029785
Kubo H, Murayama Y, Ogawa S, Matsumoto T, Yubakami M, Ohashi T, Kubota T, Okamoto K, Kamiya M, Urano Y, Otsuji E (2021) β-Galactosidase is a target enzyme for detecting peritoneal metastasis of gastric cancer. Sci Rep 11(1):1–10. https://doi.org/10.1038/s41598-021-88982-2
Kumari M, Padhi S, Sharma S, Phukon LC, Singh SP, Rai AK (2021). Biotechnological potential of psychrophilic microorganisms as the source of cold-active enzymes in food processing applications. 3 Biotech 11: 479. https://doi.org/10.1007/s13205-021-03008-y
Liu Y, Wu Z, Zeng X, Weng P, Zhang X, Wang C (2021) A novel cold-adapted phospho-beta-galactosidase from Bacillus velezensis and its potential application for lactose hydrolysis in milk. Int J Biol Macromol 166:760–770. https://doi.org/10.1016/j.ijbiomac.2020.10.233
Lomer MC, Sanderson PGC, JD, (2008) lactose intolerance in clinical practice–myths and realities. Aliment Pharmacol Ther 27(2):93–103. https://doi.org/10.1111/j.1365-2036.2007.03557.x
Lu L, Guo L, Wang K, Liu Y, Xiao M (2020) β-Galactosidases: a great tool for synthesizing galactose-containing carbohydrates. Biotechnol Adv 39:107465. https://doi.org/10.1016/j.biotechadv.2019.107465
Mangiagalli M, Brocca S, Orlando M, Lotti M (2020) The “cold revolution”. Present and future applications of cold-active enzymes and ice-binding proteins. N Biotechnol 55:5–11. https://doi.org/10.1016/j.nbt.2019.09.003
Mangiagalli M, Lapi M, Maione SO, M, Brocca S, Pesce A, Barbiroli A, Camilloni, C, Pucciarelli S, Lotti M, Nardini M, (2021) The co-existence of cold activity and thermal stability in an Antarctic GH42 β-galactosidase relies on its hexameric quaternary arrangement. FEBS J 288(2):546–565. https://doi.org/10.1111/febs.15354
Mangiagalli M, Lotti M (2021) Cold-Active β-Galactosidases: Insight into Cold Adaption Mechanisms and Biotechnological Exploitation. Mar Drugs 19(1):43. https://doi.org/10.3390/md19010043
Movahedpour A, Ahmadi N, Ghalamfarsa F, Ghesmati Z, Khalifeh M, Maleksabet A, Shabaninejad Z, Taheri-Anganeh M, Savardashtaki A (2021) β-Galactosidase: from its source and applications to its recombinant form. Biotechnol Appl Biochem. https://doi.org/10.1002/bab.2137
Mulualem DM, Agbavwe C, Ogilvie LA, Jones BV, Kilcoyne M, O’Byrne C, Boyd A (2021) Metagenomic identification, purification and characterization of the Bifidobacterium adolescentis BgaC β-galactosidase. Appl Microbiol Biotechnol 105(3):1063–1078. https://doi.org/10.1007/s00253-020-11084-y
Nakagawa T, Fujimoto Y, Ikehata R, Miyaji T, Tomizuka N (2006) Purification and molecular characterization of cold-active β-galactosidase from Arthrobacter psychrolactophilus strain F2. Appl Microbiol Biotechnol 72(4):720–725. https://doi.org/10.1007/s00253-006-0339-0
Nakagawa T, Ikehata R, Myoda T, Miyaji T, Tomizuka N (2007) Overexpression and functional analysis of cold-active β-galactosidase from Arthrobacter psychrolactophilus strain F2. Protein Expr Purif 54(2):295–299. https://doi.org/10.1016/j.pep.2007.03.010
Pennington JA, Wilson DB, Young BE, Vanderveen JRD, JE, (1987) Mineral content of market samples of fluid whole milk. J Am Diet Assoc 87(8):1036–1042. https://doi.org/10.1016/S0002-8223(21)03238-7
Ratajczak AE, Rychter AM, Zawada A, Dobrowolska A, Krela-Kaźmierczak I (2021) Lactose intolerance in patients with inflammatory bowel diseases and dietary management in prevention of osteoporosis. Nutrition 82:111043. https://doi.org/10.1016/j.nut.2020.111043
Rhimi, M, Aghajari N, Jaouadi B, Juy M, Boudebbouze S, Maguin E, Haser R, Bejar S (2009) Exploring the acidotolerance of β-galactosidase from Lactobacillus delbrueckii subsp. bulgaricus: an attractive enzyme for lactose bioconversion. Res Microbiol 160(10):775–784. https://doi.org/10.1016/j.resmic.2009.09.004
Rhimi M, Boisson A, Dejob M, Boudebouze S, Maguin E, Haser R, Aghajari N (2010) Efficient bioconversion of lactose in milk and whey: immobilization and biochemical characterization of a β-galactosidase from the dairy Streptococcus thermophilus LMD9 strain. Res Microbiol 161(7):515–525. https://doi.org/10.1016/j.resmic.2010.04.011
Rodrıguez ER.́, Alaejos MS. Romero CD, (2001) Mineral concentrations in cow’s milk from the Canary Island. J Food Compost Anal 14(4):419–430. https://doi.org/10.1006/jfca.2000.0986
Saqib S, Akram A, Halim S, Tassaduq R (2017) Sources of β-galactosidase and its applications in food industry. 3 Biotech 7(1):79. https://doi.org/10.1007/s13205-017-0645-5
Schmidt M, Stougaard P (2010) Identification, cloning and expression of a cold-active β-galactosidase from a novel Arctic bacterium, Alkalilactibacillus ikkense. Environ Technol 31(10):1107-1114. https://doi.org/10.1080/09593331003677872
Sharma SK, Leblanc RM (2017) Biosensors based on β-galactosidase enzyme: recent advances and perspectives. Anal Biochem 535:1–11. https://doi.org/10.1016/j.ab.2017.07.019
Sheridan PP, Brenchley JE (2000) Characterization of a salt-tolerant family 42 β-galactosidase from a psychrophilic Antarctic Planococcus isolate. Appl Environ Microbiol 66(6):2438–2444. https://doi.org/10.1128/aem.66.6.2438-2444.2000
Silanikove N, Leitner G, Merin U (2015) The interrelationships between lactose intolerance and the modern dairy industry: global perspectives in evolutional and historical backgrounds. Nutrients 7(9):7312–7331. https://doi.org/10.3390/nu7095340
Thakur M, Sharma N, Rai AK. Singh SP (2021) A novel cold-active type I pullulanase from a hot-spring metagenome for effective debranching and production of resistant starch. Bioresour Technol 320:124288. https://doi.org/10.1016/j.biortech.2020.124288
Tzortzis G, Vulevic J (2009) Galacto-oligosaccharide prebiotics. Prebiotics Probiotics Sci Technol :207. https://doi.org/10.1007/978-0-387-79058-9_7
Ugidos-Rodríguez S, Matallana-González MC, Sánchez-Mata MC (2018) Lactose malabsorption and intolerance: a review. Food Funct 9(8):4056–4068. https://doi.org/10.1039/c8fo00555a
Vasileva N, Ivanov Y, Damyanova S, Kostova I, Godjevargova T (2016) Hydrolysis of whey lactose by immobilized β-galactosidase in a bioreactor with a spirally wound membrane. Int J Biol Macromol 82:339–346. https://doi.org/10.1016/j.ijbiomac.2015.11.025
Vincent V, Aghajari N, Pollet N, Boisson A, Boudebbouze S, Haser R, Maguin E, Rhimi M (2013) The acid tolerant and cold-active β-galactosidase from Lactococcus lactis strain is an attractive biocatalyst for lactose hydrolysis. Antonie Leeuwenhoek 103(4):701–712. https://doi.org/10.1007/s10482-012-9852-6
Wang K, Li G, Yu SQ, Zhang CT, Liu YH (2010) A novel metagenome-derived β-galactosidase: gene cloning, overexpression, purification and characterization. Appl Microbiol Biotechnol 88(1):155–165. https://doi.org/10.1007/s00253-010-2744-7
Wang L, Mou Y, Guan B, Hu Y, Zhang Y, Zeng J, Ni Y (2020) Genome sequence of the psychrophilic Cryobacterium sp. LW097 and characterization of its four novel cold-adapted β-galactosidases. Int J Biol Macromol 163:2068–2083. https://doi.org/10.1016/j.ijbiomac.2020.09.100
Wierzbicka-Woś A, Cieśliński H, Wanarska M, Kozłowska-Tylingo K, Hildebrandt P. Kur J (2011) A novel cold-active β-D-galactosidase from the Paracoccus sp. 32d-gene cloning, purification and characterization. Microb Cell Fact 10(1):1–12. https://doi.org/10.1186/1475-2859-10-108
Xavier JR, Ramana KV, Sharma RK (2018) β-Galactosidase: biotechnological applications in food processing. J Food Biochem 42(5):2564. https://doi.org/10.1111/jfbc.12564
Xu K, Tang X, Gai Y, Mehmood MA, Xiao X, Wang F (2011) Molecular characterization of cold-inducible β-galactosidase from Arthrobacter sp. ON14 isolated from Antarctica. J Microbiol Biotechnol, 21(3):236–242. https://dx.doi.org/https://doi.org/10.4014/jmb.1009.09010
Yañez-Ñeco CV, Cervantes FV, Amaya-Delgado L, Ballesteros AO, Plou FJ, Arrizon J (2021) Synthesis of β (1→ 3) and β (1→ 6) galactooligosaccharides from lactose and whey using a recombinant β-galactosidase from Pantoea anthophila. Electron J Biotechnol 49:14–21. https://doi.org/10.1016/j.ejbt.2020.10.004
Yao C, Sun J, Wang W, Zhuang Z, Hao LJ, J, (2019) A novel cold-adapted β-galactosidase from Alteromonas sp. ML117 cleaves milk lactose effectively at low temperature. Process Biochem 82:94–101. https://doi.org/10.1016/j.procbio.2019.04.016
Acknowledgements
The authors acknowledge the Department of Biotechnology (DBT), Govt. of India, for financial support. M. T. acknowledges the CSIR fellowship. S. P. S. and A. K. R. acknowledge the DBT research project, BT/PR31829/NDB/39/649/2019.
Author information
Authors and Affiliations
Contributions
M. T. performed all the experiments, analyzed the data, and drafted the manuscript. S. P. S. conceived, designed, and supervised the study, analyzed the data, and drafted the manuscript. A. K. R. contributed to study design, data analysis, and reviewed the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Competing interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Thakur, M., Kumar Rai, A. & Singh, S.P. An acid-tolerant and cold-active β-galactosidase potentially suitable to process milk and whey samples. Appl Microbiol Biotechnol 106, 3599–3610 (2022). https://doi.org/10.1007/s00253-022-11970-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-11970-7