Abstract
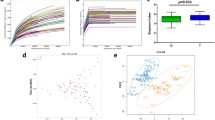
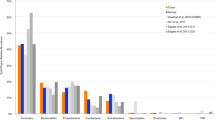
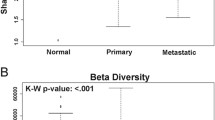
Tongue squamous cell carcinoma (TSCC) is the most common oral cavity malignancy. The role of the microbial community in TSCC development and progression is unclear. In the present study, 23 patients with TSCC were recruited. Tissue DNA was extracted from cancer and paracancerous normal tissues from all participants. Next-generation 16S rDNA amplicon sequencing and functional prediction were applied for taxonomic analysis. Alpha diversity measurements using the Shannon and Simpson diversity indexes indicated a significant increase in the microbiotic diversity of cancer samples (Shannon index: P = 0.001, Simpson index: P = 0.015); otherwise, no differences were found when using observed operational taxonomic units (OTUs) and Chao1 index (observed OTUs: P = 0.261, Chao1 index: P = 0.054). The dominant phyla of the microbiota included Bacteroidetes, Proteobacteria, Firmicutes, Actinobacteria, and Fusobacteria. Multivariate analysis of variance (Adonis) and nonparametric analysis of similarities (ANOSIM) based on unweighted unifrac distances demonstrated differences in the bacterial community structure between the two groups (P = 0.001 for Adonis, P = 0.001 for ANOSIM). Compared with the normal samples, Neisseria, Streptococcus, and Actinomyces levels decreased significantly in cancer samples. Co-occurrence network analysis implied that the bacterial community in cancer was more conserved than that in normal tissue. Matched-pair analysis of cancer and control samples revealed a significant alteration in the relative abundance of specific taxa. These findings will enrich our knowledge of the association between the oral microbial community and TSCC. Further experiments should investigate the potential carcinogenic mechanism of microbial community alterations in TSCC.
Key points
• Microbial community role in tongue squamous cell carcinoma.
• Significant alteration of microbiome found between cancer and normal tissues.
• Microbial community alteration and potential carcinogenic mechanism.






Similar content being viewed by others
Data availability
All data generated or analyzed during this study are included in this published article.
Code availability
Not applicable.
References
Al-Hebshi NN, Nasher AT, Maryoud MY, Homeida HE, Chen T, Idris AM, Johnson NW (2017) Inflammatory bacteriome featuring Fusobacterium nucleatum and Pseudomonas aeruginosa identified in association with oral squamous cell carcinoma. Sci Rep 7:1834. https://doi.org/10.1038/s41598-017-02079-3
Anderson AC (2014) Tim-3: an emerging target in the cancer immunotherapy landscape. Cancer Immunol Res 2(5):393–398. https://doi.org/10.1158/2326-6066.Cir-14-0039
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336. https://doi.org/10.1038/nmeth.f.303
Chen X, Winckler B, Lu M, Cheng H, Yuan Z, Yang Y, Jin L, Ye W (2015) Oral microbiota and risk for esophageal squamous cell carcinoma in a high-risk area of china. PLoS ONE 10(12):e0143603. https://doi.org/10.1371/journal.pone.0143603
Chen T, Yu WH, Izard J, Baranova OV, Lakshmanan A, Dewhirst FE (2010) The Human Oral Microbiome Database: a web accessible resource for investigating oral microbe taxonomic and genomic information. Database(Oxford) 2010:baq013. https://doi.org/10.1093/database/baq013
Fazlollahi M, Lee TD, Andrade J, Oguntuyo K, Chun Y, Grishina G, Grishin A, Bunyavanich S (2018) The nasal microbiome in asthma. J Allergy Clin Immunol 142:834-843.e2. https://doi.org/10.1016/j.jaci.2018.02.020
Flemer B, Warren RD, Barrett MP, Cisek K, Das A, Jeffery IB, Hurley E, O’Riordain M, Shanahan F, O’Toole PW (2018) The oral microbiota in colorectal cancer is distinctive and predictive. Gut 67(8):1454–1463. https://doi.org/10.1136/gutjnl-2017-314814
Fouhy F, Deane J, Rea MC, O’Sullivan Ó, Ross RP, O’Callaghan G, Plant BJ, Stanton C (2015) The effects of freezing on faecal microbiota as determined using MiSeq sequencing and culture-based investigations. PLoS ONE 10:e0119355. https://doi.org/10.1371/journal.pone.0119355
Gaibani P, Caroli F, Nucci C, Sambri V (2010) Major surface protein complex of Treponema denticola induces the production of tumor necrosis factor alpha, interleukin-1 beta, interleukin-6 and matrix metalloproteinase 9 by primary human peripheral blood monocytes. J Periodontal Res 45(3):361–366. https://doi.org/10.1111/j.1600-0765.2009.01246.x
Galvão-Moreira LV, da Cruz MC (2016) Oral microbiome, periodontitis and risk of head and neck cancer. Oral Oncol 53:17–19. https://doi.org/10.1016/j.oraloncology.2015.11.013
Gao X, Zhou J, Sun X, Li X, Zhou Y (2018) Diversity analysis of subgingival microbial bacteria in peri-implantitis in Uygur population. Medicine 97(5):e9774. https://doi.org/10.1097/md.0000000000009774
Hamidi B, Wallace K, Vasu C, Alekseyenko AV (2019) W∗d -test: robust distance-based multivariate analysis of variance. Microbiome 7:51. https://doi.org/10.1186/s40168-019-0659-9
Jo AR, Baek KJ, Shin JE, Choi Y (2014) Mechanisms of IL-8 suppression by Treponema denticola in gingival epithelial cells. Immunol Cell Biol 92(2):139–147. https://doi.org/10.1038/icb.2013.80
Kakabadze MZ, Paresishvili T, Karalashvili L, Chakhunashvili D, Kakabadze Z (2020) Oral microbiota and oral cancer: review. Oncol Rev 14:476. https://doi.org/10.4081/oncol.2020.476
Katz J, Onate MD, Pauley KM, Bhattacharyya I, Cha S (2011) Presence of Porphyromonas gingivalis in gingival squamous cell carcinoma. Int J Oral Sci 3:209–215. https://doi.org/10.4248/IJOS11075
Keir ME, Butte MJ, Freeman GJ, Sharpe AH (2008) PD-1 and its ligands in tolerance and immunity. Annu Rev Immunol 26:677–704. https://doi.org/10.1146/annurev.immunol.26.021607.090331
Kumar M, Nanavati R, Modi TG, Dobariya C (2016) Oral cancer: etiology and risk factors: a review. J Cancer Res and Ther 12(2):458–463. https://doi.org/10.4103/0973-1482.186696
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA, Clemente JC, Burkepile DE, Vega Thurber RL, Knight R, Beiko RG, Huttenhower C (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821. https://doi.org/10.1038/nbt.2676
Lee WH, Chen HM, Yang SF, Liang C, Peng CY, Lin FM, Tsai LL, Wu BC, Hsin CH, Chuang CY, Yang T, Yang TL, Ho SY, Chen WL, Ueng KC, Huang HD, Huang CN, Jong YJ (2017) Bacterial alterations in salivary microbiota and their association in oral cancer. Sci Rep 7:16540. https://doi.org/10.1038/s41598-017-16418-x
Li Y, Tan X, Zhao X, Xu Z, Dai W, Duan W, Huang S, Zhang E, Liu J, Zhang S, Yin R, Shi X, Lu Z, Pan Y (2020) Composition and function of oral microbiota between gingival squamous cell carcinoma and periodontitis. Oral Oncol 107:104710. https://doi.org/10.1016/j.oraloncology.2020.104710
Mager DL, Haffajee AD, Devlin PM, Norris CM, Posner MR, Goodson JM (2005) The salivary microbiota as a diagnostic indicator of oral cancer: a descriptive, non-randomized study of cancer-free and oral squamous cell carcinoma subjects. J Transl Med 3:27. https://doi.org/10.1186/1479-5876-3-27
Mukherjee PK, Wang H, Retuerto M, Zhang H, Burkey B, Ghannoum MA, Eng C (2017) Bacteriome and mycobiome associations in oral tongue cancer. Oncotarget 8:97273–97289. https://doi.org/10.18632/oncotarget.21921
Parsonnet J, Friedman GD, Vandersteen DP, Chang Y, Vogelman JH, Orentreich N, Sibley RK (1991) Helicobacter pylori infection and the risk of gastric carcinoma. N Engl J Med 325:1127–1131. https://doi.org/10.1056/NEJM199110173251603
Peters BA, Wu J, Pei Z, Yang L, Purdue MP, Freedman ND, Jacobs EJ, Gapstur SM, Hayes RB, Ahn J (2017) Oral microbiome composition reflects prospective risk for esophageal cancers. Cancer Res 77(23):6777–6787. https://doi.org/10.1158/0008-5472.Can-17-1296
Reyon D, Tsai SQ, Khayter C, Foden JA, Sander JD, Joung JK (2012) FLASH assembly of TALENs for high-throughput genome editing. Nat Biotechnol 30:460–465. https://doi.org/10.1038/nbt.2170
Rognes T, Flouri T, Nichols B, Quince C, Mahe F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Samim D, Méan M, Clair C, Marques-Vidal P (2018) A 10-year observational study on the trends and determinants of smoking status. PLoS ONE 13:e0200010. https://doi.org/10.1371/journal.pone.0200010
Sasaki M, Yamaura C, Ohara-Nemoto Y, Tajika S, Kodama Y, Ohya T, Harada R, Kimura S (2005) Streptococcus anginosus infection in oral cancer and its infection route. Oral Dis 11:151–156. https://doi.org/10.1111/j.1601-0825.2005.01051.x
Schmidt BL, Kuczynski J, Bhattacharya A, Huey B, Corby PM, Queiroz EL, Nightingale K, Kerr AR, DeLacure MD, Veeramachaneni R, Olshen AB, Albertson DG (2014) Changes in abundance of oral microbiota associated with oral cancer. PLoS ONE 9:e98741. https://doi.org/10.1371/journal.pone.0098741
Shibahara T (2017) Oral cancer diagnosis and therapy. Clin Calcium 27(10):1427–1433. https://pubmed.ncbi.nlm.nih.gov/28947694/
Simonson LG, Goodman CH, Bial JJ, Morton HE (1988) Quantitative relationship of Treponema denticola to severity of periodontal disease. Infect Immun 56(4):726–728. https://doi.org/10.1128/iai.56.4.726-728.1988
Walters KE, Martiny JBH (2020) Alpha-, beta-, and gamma-diversity of bacteria varies across habitats. PLoS ONE 15:e0233872. https://doi.org/10.1371/journal.pone.0233872
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Wang J, Sun F, Lin X, Li Z, Mao X, Jiang C (2018) Cytotoxic T cell responses to Streptococcus are associated with improved prognosis of oral squamous cell carcinoma. Exp Cell Res 362(1):203–208. https://doi.org/10.1016/j.yexcr.2017.11.018
Willis JR, González-Torres P, Pittis AA, Bejarano LA, Cozzuto L, Andreu-Somavilla N, Alloza-Trabado M, Valentín A, Ksiezopolska E, Company C, Onywera H, Montfort M, Hermoso A, Iraola-Guzmán S, Saus E, Labeeuw A, Carolis C, Hecht J, Ponomarenko J, Gabaldón T (2018) Citizen science charts two major “stomatotypes” in the oral microbiome of adolescents and reveals links with habits and drinking water composition. Microbiome 6(1):218. https://doi.org/10.1186/s40168-018-0592-3
Yost S, Stashenko P, Choi Y, Kukuruzinska M, Genco CA, Salama A, Weinberg EO, Kramer CD, Frias-Lopez J (2018) Increased virulence of the oral microbiome in oral squamous cell carcinoma revealed by metatranscriptome analyses. Int J Oral Sci 10:32. https://doi.org/10.1038/s41368-018-0037-7
Zhang L, Liu Y, Zheng HJ, Zhang CP (2019) The oral microbiota may have influence on oral cancer. Front Cell Infect Microbiol 9:476. https://doi.org/10.3389/fcimb.2019.00476
Zhao H, Chu M, Huang Z, Yang X, Ran S, Hu B, Zhang C, Liang J (2017) Variations in oral microbiota associated with oral cancer. Sci Rep 7:11773. https://doi.org/10.1038/s41598-017-11779-9
Zitvogel L, Ayyoub M, Routy B, Kroemer G (2016) Microbiome and anticancer immunosurveillance. Cell 165(2):276–287. https://doi.org/10.1016/j.cell.2016.03.001
Funding
This work was supported by the National Key Research and Development Program of China (Grant: 2018YFC2000505) and the Applied Research Program of Capital Clinical Features (Grant Z18110001718172).
Author information
Authors and Affiliations
Contributions
PY, HTX, and ZYL conceived the study. YL, PY, YJC, HY, and HTX analyzed data. PY and YL wrote the manuscript. All authors contributed to literature searches. All authors have read and approved the manuscript.
Corresponding authors
Ethics declarations
Ethics approval
This study was approved by the Ethics Committee of Beijing Hospital.
Consent to participate
All recruited patients have signed their consent form to participate.
Consent for publication
All co-authors have given their consent to publish this manuscript.
Conflict of interest
Theauthors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Ye, P., Liu, Y., Cai, YJ. et al. Microbial community alteration in tongue squamous cell carcinoma. Appl Microbiol Biotechnol 105, 8457–8467 (2021). https://doi.org/10.1007/s00253-021-11593-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-021-11593-4