Abstract
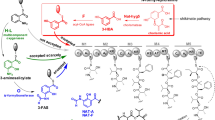
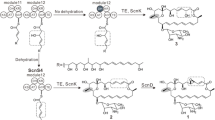
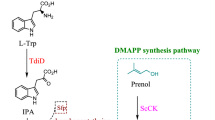
Polyene antibiotics, including amphotericin, nystatin, pimaricin, and tetramycin, are important antifungal agents. Increasing the production of polyenes and generation of their improved analogues based on the biosynthetic pathway engineering has aroused wide concern in application researches. Herein, tetramycin and nystatin, both of which share most of acyl-CoA precursors, are produced by Streptomyces hygrospinosus var. beijingensis CGMCC 4.1123. Thus, the intracellular malonyl-CoA is found to be insufficient for PKSs (polyketide synthases) extension of tetramycin by quantitative analysis in this wild-type strain. To circumvent this problem and increase tetramycin titer, the acyl-CoA competing biosynthetic gene cluster (BGC) of nystatin was disrupted, and the biosynthetic genes of malonyl-CoA from S. coelicolor M145 were integrated and overexpressed in nys-disruption mutant strain (SY02). Moreover, in order to specifically accumulate tetramycin B from A, two copies of tetrK and a copy of tetrF were introduced, resulting in elevating tetramycin B fermentration titer by 122% to 865 ± 8 mg/L than the wild type. In this optimized strain, a new tetramycin derivative, 12-decarboxy-12-methyl tetramycin B, was generated with a titer of 371 ± 26 mg/L through inactivation of a P450 monooxygenase gene tetrG. Compared with tetramycin B, the new compound exhibited higher antifungal activity against Saccharomyces cerevisiae and Rhodotorula glutinis, but lower hemolytic toxicity to erythrocyte. This research provided a good example of employing biosynthetic engineering strategies for fermentation titer improvement of polyene and development of the derivatives for medicinal applications.





Similar content being viewed by others
References
Aparicio JF, Barreales EG, Payero TD, Vicente CM, de Pedro A, Santos-Aberturas J (2016) Biotechnological production and application of the antibiotic pimaricin: biosynthesis and its regulation. Appl Microbiol Biotechnol 100(1):61–78. https://doi.org/10.1007/s00253-015-7077-0
Armando JW, Boghigian BA, Pfeifer BA (2012) LC-MS/MS quantification of short-chain acyl-CoA’s in Escherichia coli demonstrates versatile propionyl-CoA synthetase substrate specificity. Lett Appl Microbiol 54(2):140–148. https://doi.org/10.1111/j.1472-765X.2011.03184.x
Behrendorff JBYH, Huang W, Gillam EMJ (2015) Directed evolution of cytochrome P450 enzymes for biocatalysis: exploiting the catalytic versatility of enzymes with relaxed substrate specificity. Biochem J 467(1):1–15. https://doi.org/10.1042/BJ20141493
Blin K, Wolf T, Chevrette MG, Lu X, Schwalen CJ, Kautsar SA, Suarez Duran HG, de Los Santos ELC, Kim HU, Nave M, Dickschat JS, Mitchell DA, Shelest E, Breitling R, Takano E, Lee SY, Weber T, Medema MH (2017) antiSMASH 4.0-improvements in chemistry prediction and gene cluster boundary identification. Nucleic Acids Res 45(W1):W36–W41. https://doi.org/10.1093/nar/gkx319
Borgos SE, Tsan P, Sletta H, Ellingsen TE, Lancelin JM, Zotchev SB (2006) Probing the structure-function relationship of polyene macrolides: engineered biosynthesis of soluble nystatin analogues. J Med Chem 49(8):2431–2439. https://doi.org/10.1021/jm050895w
Brautaset T, Sekurova ON, Sletta H, Ellingsen TE, StrLm AR, Valla S, Zotchev SB (2000) Biosynthesis of the polyene antifungal antibiotic nystatin in Streptomyces noursei ATCC 11455: analysis of the gene cluster and deduction of the biosynthetic pathway. Chem Biol 7(6):395–403. https://doi.org/10.1016/S1074-5521(00)00120-4
Brautaset T, Sletta H, Nedal A, Borgos SE, Degnes KF, Bakke I, Volokhan O, Sekurova ON, Treshalin ID, Mirchink EP, Dikiy A, Ellingsen TE, Zotchev SB (2008) Improved antifungal polyene macrolides via engineering of the nystatin biosynthetic genes in Streptomyces noursei. Chem Biol 15(11):1198–1206. https://doi.org/10.1016/j.chembiol.2008.08.009
Caffrey P, De Poire E, Sheehan J, Sweeney P (2016) Polyene macrolide biosynthesis in streptomycetes and related bacteria: recent advances from genome sequencing and experimental studies. Appl Microbiol Biotechnol 100(9):3893–3908. https://doi.org/10.1007/s00253-016-7474-z
Cao B, Yao F, Zheng X, Cui D, Shao Y, Zhu C, Deng Z, You D (2012) Genome mining of the biosynthetic gene cluster of the polyene macrolide antibiotic tetramycin and characterization of a P450 monooxygenase involved in the hydroxylation of the tetramycin B polyol segment. ChemBioChem 13(15):2234–2242. https://doi.org/10.1002/cbic.201200402
Carmody M, Murphy B, Byrne B, Power P, Rai D, Rawlings B, Caffrey P (2005) Biosynthesis of amphotericin derivatives lacking exocyclic carboxyl groups. J Biol Chem 280(41):34420–34426. https://doi.org/10.1074/jbc.M506689200
Chen S, Mao X, Shen Y, Zhou Y, Li J, Wang L, Tao X, Yang L, Wang Y, Zhou X, Deng Z, Wei D (2009) Tailoring the P450 monooxygenase gene for FR-008/candicidin biosynthesis. Appl Environ Microbiol 75(6):1778–1781. https://doi.org/10.1128/AEM.00859-08
Cheron M, Cybulska B, Mazerski J, Grzybowska J, Czerwinski A, Borowski E (1988) Quantitative structure-activity relationships in amphotericin B derivatives. Biochem Pharmacol 37(5):827–836. https://doi.org/10.1016/0006-2952(88)90168-2
Choi SS, Katsuyama Y, Bai L, Deng Z, Ohnishi Y, Kim ES (2018) Genome engineering for microbial natural product discovery. Curr Opin Microbiol 45:53–60. https://doi.org/10.1016/j.mib.2018.02.007
Cui H, Ni X, Shao W, Su J, Su J, Ren J, Xia H (2015) Functional manipulations of the tetramycin positive regulatory gene ttmRIV to enhance the production of tetramycin A and nystatin A1 in Streptomyces ahygroscopicus. J Ind Microbiol Biotechnol 42(9):1273–1282. https://doi.org/10.1007/s10295-015-1660-3
Cui H, Ni X, Liu S, Wang J, Sun Z, Ren J, Su J, Chen G, Xia H (2016) Characterization of three positive regulators for tetramycin biosynthesis in Streptomyces ahygroscopicus. FEMS Microbiol Lett 363(12). https://doi.org/10.1093/femsle/fnw109
Diacovich L, Peiru S, Kurth D, Rodriguez E, Podesta F, Khosla C, Gramajo H (2002) Kinetic and structural analysis of a new group of Acyl-CoA carboxylases found in Streptomyces coelicolor A3(2). J Biol Chem 277(34):31228–31236. https://doi.org/10.1074/jbc.M203263200
Du YL, Chen SF, Cheng LY, Shen XL, Tian Y, Li YQ (2009) Identification of a novel Streptomyces chattanoogensis L10 and enhancing its natamycin production by overexpressing positive regulator ScnRII. J Microbiol 47(4):506–513. https://doi.org/10.1007/s12275-009-0014-0
Hamill RJ (2013) Amphotericin B formulations: a comparative review of efficacy and toxicity. Drugs 73(9):919–934. https://doi.org/10.1007/s40265-013-0069-4
Hasan H, Abd Rahim MH, Campbell L, Carter D, Abbas A, Montoya A (2018) Overexpression of acetyl-CoA carboxylase in Aspergillus terreus to increase lovastatin production. New Biotechnol 44:64–71. https://doi.org/10.1016/j.nbt.2018.04.008
Jiang H, Wang YY, Ran XX, Fan WM, Jiang XH, Guan WJ, Li YQ (2013) Improvement of natamycin production by engineering of phosphopantetheinyl transferases in Streptomyces chattanoogensis L10. Appl Environ Microbiol 79(11):3346–3454. https://doi.org/10.1128/AEM.00099-13
Khosla C, Gokhale RS, Jacobsen JR, Cane DE (1999) Tolerance and specificity of polyketide synthases. Annu Rev Biochem 68(1):219–253. https://doi.org/10.1146/annurev.biochem.68.1.219
Kong D, Lee MJ, Lin S, Kim ES (2013) Biosynthesis and pathway engineering of antifungal polyene macrolides in actinomycetes. J Ind Microbiol Biotechnol 40(6):529–543. https://doi.org/10.1007/s10295-013-1258-6
van der Lee TAJ, Medema MH (2016) Computational strategies for genome-based natural product discovery and engineering in fungi. Fungal Genet Biol 89:29–36. https://doi.org/10.1016/j.fgb.2016.01.006
Li L, Zhao Y, Ruan L, Yang S, Ge M, Jiang W, Lu Y (2015) A stepwise increase in pristinamycin II biosynthesis by Streptomyces pristinaespiralis through combinatorial metabolic engineering. Metab Eng 29:12–25. https://doi.org/10.1016/j.ymben.2015.02.001
Li L, Zheng G, Chen J, Ge M, Jiang W, Lu Y (2017) Multiplexed site-specific genome engineering for overproducing bioactive secondary metabolites in actinomycetes. Metab Eng 40:80–92. https://doi.org/10.1016/j.ymben.2017.01.004
Liu SP, Yuan PH, Wang YY, Liu XF, Zhou ZX, Bu QT, Yu P, Jiang H, Li YQ (2015) Generation of the natamycin analogs by gene engineering of natamycin biosynthetic genes in Streptomyces chattanoogensis L10. Microbiol Res 173:25–33. https://doi.org/10.1016/j.micres.2015.01.013
Lu C, Zhang X, Jiang M, Bai L (2016) Enhanced salinomycin production by adjusting the supply of polyketide extender units in Streptomyces albus. Metab Eng 35:129–137. https://doi.org/10.1016/j.ymben.2016.02.012
Mao X, Wang F, Zhang J, Chen S, Deng Z, Shen Y, Wei D (2009) The pH shift and precursor feeding strategy in a low-toxicity FR-008/candicidin derivative CS103 fermentation bioprocess by a mutant of Streptomyces sp. FR-008. Appl Biochem Biotechnol 159(3):673–686. https://doi.org/10.1007/s12010-008-8502-y
Matsumori N, Eiraku N, Matsuoka S, Oishi T, Murata M, Aoki T, Ide T (2004) An amphotericin B-ergosterol covalent conjugate with powerful membrane permeabilizing activity. Chem Biol 11(5):673–679. https://doi.org/10.1016/j.chembiol.2004.02.027
Meng L, Xiong Z, Chu J, Wang Y (2016) Enhanced production of avermectin by deletion of type III polyketide synthases biosynthetic cluster rpp in Streptomyces avermitilis. Lett Appl Microbiol 63(5):384–390. https://doi.org/10.1111/lam.12635
Nah HJ, Pyeon HR, Kang SH, Choi SS, Kim ES (2017) Cloning and heterologous expression of a large-sized natural product biosynthetic gene cluster in Streptomyces species. Front Microbiol 8(394). https://doi.org/10.3389/fmicb.2017.00394
Nedal A, Sletta H, Brautaset T, Borgos SE, Sekurova ON, Ellingsen TE, Zotchev SB (2007) Analysis of the mycosamine biosynthesis and attachment genes in the nystatin biosynthetic gene cluster of Streptomyces noursei ATCC 11455. Appl Environ Microbiol 73(22):7400–7407. https://doi.org/10.1128/AEM.01122-07
Niemirowicz K, Durnas B, Tokajuk G, Gluszek K, Wilczewska AZ, Misztalewska I, Mystkowska J, Michalak G, Sodo A, Watek M, Kiziewicz B, Gozdz S, Gluszek S, Bucki R (2016) Magnetic nanoparticles as a drug delivery system that enhance fungicidal activity of polyene antibiotics. Nanomedicine 12(8):2395–2404. https://doi.org/10.1016/j.nano.2016.07.006
Qi Z, Kang Q, Jiang C, Han M, Bai L (2015) Engineered biosynthesis of pimaricin derivatives with improved antifungal activity and reduced cytotoxicity. Appl Microbiol Biotechnol 99(16):6745–6752. https://doi.org/10.1007/s00253-015-6635-9
Qi Z, Zhou Y, Kang Q, Jiang C, Zheng J, Bai L (2017) Directed accumulation of less toxic pimaricin derivatives by improving the efficiency of a polyketide synthase dehydratase domain. Appl Microbiol Biotechnol 101(6):2427–2436. https://doi.org/10.1007/s00253-016-8074-7
Radics L, Incze M, Dornberger K, Thrum H (1982) Tetramycin B, a new polyene macrolide antibiotic : the structure of tetramycins A and B as studied by high-field NMR spectroscopy. Tetrahedron 38(1):183–189. https://doi.org/10.1016/0040-4020(82)85064-3
Ren J, Cui Y, Zhang F, Cui H, Ni X, Chen F, Li L, Xia H (2014) Enhancement of nystatin production by redirecting precursor fluxes after disruption of the tetramycin gene from Streptomyces ahygroscopicus. Microbiol Res 169(7–8):602–608. https://doi.org/10.1016/j.micres.2013.09.017
Robertsen HL, Weber T, Kim HU, Lee SY (2018) Toward systems metabolic engineering of streptomycetes for secondary metabolites production. Biotechnol J 13(1). https://doi.org/10.1002/biot.201700465
Rodriguez E, Banchio C, Diacovich L, Bibb MJ, Gramajo H (2001) Role of an essential acyl coenzyme a carboxylase in the primary and secondary metabolism of Streptomyces coelicolor A3(2). Appl Environ Microbiol 67(9):4166–4176. https://doi.org/10.1128/aem.67.9.4166-4176.2001
Tierrafría VH, Licona-Cassani C, Maldonado-Carmona N, Romero-Rodríguez A, Centeno-Leija S, Marcellin E, Rodríguez-Sanoja R, Ruiz-Villafán B, Nielsen L, Sánchez S (2016) Deletion of the hypothetical protein SCO2127 of Streptomyces coelicolor allowed identification of a new regulator of actinorhodin production. Appl Microbiol Biotechnol 100(21):9229–9237. https://doi.org/10.1007/s00253-016-7811-2
Wang TJ, Shan YM, Li H, Dou WW, Jiang XH, Mao XM, Liu SP, Guan WJ, Li YQ (2017) Multiple transporters are involved in natamycin efflux in streptomyces chattanoogensis L10. Mol Microbiol 103(4):713–728. https://doi.org/10.1111/mmi.13583
te Welscher YM, ten Napel HH, Balague MM, Souza CM, Riezman H, de Kruijff B, Breukink E (2008) Natamycin blocks fungal growth by binding specifically to ergosterol without permeabilizing the membrane. J Biol Chem 283(10):6393–6401. https://doi.org/10.1074/jbc.M707821200
te Welscher YM, van Leeuwen MR, de Kruijff B, Dijksterhuis J, Breukink E (2012) Polyene antibiotic that inhibits membrane transport proteins. Proc Natl Acad Sci U S A 109(28):11156–11159. https://doi.org/10.1073/pnas.1203375109
Funding
This work was supported by grants from the National Key Research and Development Program of China (No. 2018YFA0901900), the National Natural Science Foundation of China (Nos. 31770034, 21661140002, and 31700027), the Shanghai Municipal Council of Science and Technology (19ZR1475600), and the Startup Fund for Youngman Research at SJTU (17X100040064).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants performed by any of the authors.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 3515 kb)
Rights and permissions
About this article
Cite this article
Sheng, Y., Ou, Y., Hu, X. et al. Generation of tetramycin B derivative with improved pharmacological property based on pathway engineering. Appl Microbiol Biotechnol 104, 2561–2573 (2020). https://doi.org/10.1007/s00253-020-10391-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10391-8