Abstract
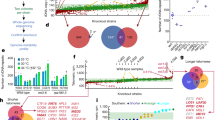
Mitochondrial dysfunction in Saccharomyces cerevisiae was selected as a marker of ion penetration following carbon ion beam (CIB) irradiation. Respiration-deficient mutants were screened. Following confirmation of negligible spontaneous mutation, eight genetically stable S. cerevisiae respiration-deficient mutant strains and a control strain were resequenced with ~ 200-fold read depth. Strategies were established to identify and validate the particular mutations induced by CIB irradiation. In the nuclear genome, CIB irradiation mainly caused base substitutions and some small (< 100 bp) insertions/deletions (indels), which were widely distributed across the chromosomes. Although mitochondrial dysfunction was selected as a screening marker, variants in the nuclear genome were detected at a high frequency (10−7) relative to spontaneous mutations (10−9). The transition to transversion ratio for base substitutions was 0.746, which was less than that of spontaneous mutations. In the mitochondrial genome, there were very large deletions including substantial gene areas, resulting in extremely low read coverage. Meanwhile, every mutant possessed a distinctive mutation pattern, for both the nuclear and the mitochondrial genome. Nuclear genomes contained scanty mitochondrial respiration-related genes that were potentially affected by verified mutations, suggesting that variants in the mitochondrial genome may be the main drivers of respiratory deficiencies. Our study confirmed the previous finding that heavy ion beam (HIB) irradiation mainly induces substantial base substitutions and some small indels but also yielded some novel findings, in particular, novel structural variants in the mitochondrial genomes. These data will enhance the understanding of HIB-induced damage and mutations and aid in the HIB-based microbial mutation breeding.









Similar content being viewed by others
References
Aokinakano M, Furusawa Y (2013) Misrepair of DNA double-strand breaks after exposure to heavy-ion beams causes a peak in the LET–RBE relationship with respect to cell killing in DT40 cells. J Radiat Res 54(6):1029–1035. https://doi.org/10.1093/jrr/rrt064
Bai H, Pekarek SD, Tichenor J, Eversman W, Buening DJ, Holbrook GR, Krefta RJ (2009) The Sequence Alignment-Map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Belfield EJ, Gan X, Mithani A, Brown C, Jiang C, Franklin K, Alvey E, Wibowo A, Jung M, Bailey K (2012) Genome-wide analysis of mutations in mutant lineages selected following fast-neutron irradiation mutagenesis of Arabidopsis thaliana. Genome Res 22(7):1306–1315. https://doi.org/10.1101/gr.131474.111
Birch-Machin MA, Swalwell H (2010) How mitochondria record the effects of UV exposure and oxidative stress using human skin as a model tissue. Mutagenesis 25(2):101–107. https://doi.org/10.1093/mutage/gep061
Boiteux S, Jinks-Robertson S (2013) DNA repair mechanisms and the bypass of DNA damage in Saccharomyces cerevisiae. Genetics 193(4):1025–1064. https://doi.org/10.1534/genetics.112.145219
Chen K, Wallis JW, McLellan MD, Larson DE, Kalicki JM, Pohl CS, McGrath SD, Wendl MC, Zhang Q, Locke DP (2009) BreakDancer: an algorithm for high-resolution mapping of genomic structural variation. Nat Methods 6(9):677–681. https://doi.org/10.1038/NMETH.1363
Downs JA, Lowndes NF, Jackson SP (2000) A role for Saccharomyces cerevisiae histone H2A in DNA repair. Nature 408(6815):1001–1004. https://doi.org/10.1038/35050000
Du Y, Li W, Yu L, Chen G, Liu Q, Luo S, Shu Q, Zhou L (2014) Mutagenic effects of carbon-ion irradiation on dry Arabidopsis thaliana seeds. Mutat Res-Gen Tox En 759(1):28–36. https://doi.org/10.1016/j.mrgentox.2013.07.018
Du Y, Luo S, Li X, Yang J, Cui T, Li W, Yu L, Feng H, Chen Y, Mu J (2017) Identification of substitutions and small insertion-deletions induced by carbon-ion beam irradiation in Arabidopsis thaliana. Front Plant Sci 8:1851. https://doi.org/10.3389/fpls.2017.01851
Du Y, Luo S, Yu L, Cui T, Chen X, Yang J, Li X, Li W, Wang J, Zhou L (2018) Strategies for identification of mutations induced by carbon-ion beam irradiation in Arabidopsis thaliana by whole genome re-sequencing. Mutat Res-Fund Mol M 807:21–30. https://doi.org/10.1016/j.mrfmmm.2017.12.001
Elahe A, Orlando TM, Léon S (2015) Biomolecular damage induced by ionizing radiation: the direct and indirect effects of low-energy electrons on DNA. Annu Rev Phys Chem 66(1):379–398. https://doi.org/10.1146/annurev-physchem-040513-103605
Feng H, Yu Z, Chu PK (2006) Ion implantation of organisms. Mater Sci Eng R 54(3):49–120. https://doi.org/10.1016/j.mser.2006.11.001
Foury F, Roganti T, Lecrenier N, Purnelle B (1998) The complete sequence of the mitochondrial genome of Saccharomyces cerevisiae. FEBS 440(3):325–331. https://doi.org/10.1016/S0014-5793(98)01467-7
Foyer CH, Noctor G, Hodges M (2011) Respiration and nitrogen assimilation: targeting mitochondria-associated metabolism as a means to enhance nitrogen use efficiency. J Exp Bot 62(4):1467–1482. https://doi.org/10.1093/jxb/erq453
Freel KC, Friedrich A, Schacherer J (2015) Mitochondrial genome evolution in yeasts: an all-encompassing view. FEMS Yeast Res 15(4):fov023. https://doi.org/10.1093/femsyr/fov023
Gdaniec Z, Ban B, Sowers LC, Fazakerley GV (1996) Methoxyamine-induced mutagenesis of nucleic acids - a proton NMR study of oligonucleotides containing N-4-methoxycytosine paired with adenine or guanine. Eur J Biochem 242(2):271–279. https://doi.org/10.1111/j.1432-1033.1996.0271r.x
Goodman MF, Hopkins RL, Lasken R, Mhaskar DN (1985) The biochemical basis of 5-bromouracil- and 2-aminopurine-induced mutagenesis. Basic Life Sci 31:409–423. https://doi.org/10.1007/978-1-4613-2449-2_25
Gualberto JM, Mileshina D, Wallet C, Niazi AK, Weberlotfi F, Dietrich A (2014) The plant mitochondrial genome: dynamics and maintenance. Biochimie 100(1):107–120. https://doi.org/10.1016/j.biochi.2013.09.016
Kazama Y, Ishii K, Hirano T, Wakana T, Yamada M, Ohbu S, Abe T (2017) Different mutational function of low- and high-linear energy transfer heavy-ion irradiation demonstrated by whole-genome resequencing of Arabidopsis mutants. Plant J 92(6):1020–1030. https://doi.org/10.1111/tpj.13738
Kiefer J, Egenolf R, Ikpeme S (2002) Heavy ion-induced DNA double-strand breaks in yeast. Radiat Res 157(2):141–148. https://doi.org/10.1667/0033-7587(2002)157[0141:hiidds]2.0.co;2
Kim SR, Lee KS, Choi JH, Ha SJ, Kweon DH, Seo JH, Jin YS (2010) Repeated-batch fermentations of xylose and glucose-xylose mixtures using a respiration-deficient Saccharomyces cerevisiae engineered for xylose metabolism. J Biotechnol 150(3):404–407. https://doi.org/10.1016/j.jbiotec.2010.09.962
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows–Wheeler transform. Bioinformatics 25(14):1754–1760. https://doi.org/10.1093/bioinformatics/btp324
Li X, Wang J, Tan Z, Ma L, Lu D, Li W, Wang J (2018) Cd resistant characterization of mutant strain irradiated by carbon-ion beam. J Hazard Mater 353:1–8. https://doi.org/10.1016/j.jhazmat.2018.03.036
Liu XB, Gu QY, Yu XB, Luo W (2012) Enhancement of butanol tolerance and butanol yield in Clostridium acetobutylicum mutant NT642 obtained by nitrogen ion beam implantation. J Microbiol 50(6):1024–1028. https://doi.org/10.1007/s12275-012-2289-9
Luo S, Zhou L, Li W, Du Y, Yu L, Feng H, Mu J, Chen Y (2016) Mutagenic effects of carbon ion beam irradiations on dry Lotus japonicus seeds. NuclInstrum Meth B 383:123–128. https://doi.org/10.1016/j.nimb.2016.06.021
Lynch M, Sung W, Morris K, Coffey N, Landry CR, Dopman EB, Dickinson WJ, Okamoto K, Kulkarni S, Hartl DL (2008) A genome-wide view of the spectrum of spontaneous mutations in yeast. P Natl Acad Sci US A 105(27):9272–9277. https://doi.org/10.1073/pnas.0803466105
Mao SH, Jin GM, Wei ZQ, Xie HM, Zhang H (2006) Screening and identification of respiration deficiency mutants of yeasts (Saccharomyces cerevisiae) induced by heavy iron irradiation. J Isotopes 19(1):44–47
Mao WJ, Liu ZZ, Zhu H, Zhu RR, Sun XY, Yao SD, Wang SL (2009) Selection of the respiration deficiency mutant yeast irritated by laser and optimization of the fermentation condition. J Rad Res Rad Proc 27(1):1–4
Matuo Y, Nishijima S, Hase Y, Sakamoto A, Tanaka A, Shimizu K (2006) Specificity of mutations induced by carbon ions in budding yeast Saccharomyces cerevisiae. Mutat Res-Fund Mol M 602(1–2):7–13. https://doi.org/10.1016/j.mrfmmm.2006.07.001
Naito K, Kusaba M, Shikazono N, Takano T, Tanaka A, Tanisaka T, Nishimura M (2005) Transmissible and nontransmissible mutations induced by irradiating Arabidopsis thaliana pollen with gamma-rays and carbon ions. Genetics 169(2):881–889. https://doi.org/10.1534/genetics.104.033654
Ortiz-Muñiz B, Carvajal-Zarrabal O, Aguilar B, Aguilar-Uscanga MG (2012) Improvement in ethanol production using respiratory deficient phenotype of a wild type yeast Saccharomyces cerevisiae ITV-01. Renew Energy 37(1):197–201. https://doi.org/10.1016/j.renene.2011.06.019
Ossowski S, Schneeberger K, Lucaslledó JI, Warthmann N, Clark RM, Shaw RG, Weigel D, Lynch M (2010) The rate and molecular spectrum of spontaneous mutations in Arabidopsis thaliana. Science 327(5961):92–94. https://doi.org/10.1126/science.1180677
Pitsikas P, Patapas JM, Cupples CG (2004) Mechanism of 2-aminopurine-stimulated mutagenesis in Escherichia coli. Mutat Res-Fund Mol M 550(1–2):25–32. https://doi.org/10.1016/j.mrfmmm.2004.01.008
Porro D, Smeraldi C, Martegani E, Ranzi BM, Alberghina L (1994) Flow-cytometric determination of the respiratory activity in growing Saccharomyces cerevisiae populations. Biotechnol Prog 10(2):193–197. https://doi.org/10.1021/bp00026a009
Ravanat JL, Douki T (2016) UV and ionizing radiations induced DNA damage, differences and similarities. Radiat Phys Chem 128:92–102. https://doi.org/10.1016/j.radphyschem.2016.07.007
Reuter JA, Spacek DV, Snyder MP (2015) High-throughput sequencing technologies. Mol Cell 58(4):586–597. https://doi.org/10.1016/j.molcel.2015.05.004
Salazar AN, Gorter dVAR, Marcel VDB, Wijsman M, Pilar DLTC, Brickwedde A, Brouwers N, Daran JMG, Abeel T (2017) Nanopore sequencing enables near-complete de novo assembly of Saccharomyces cerevisiae reference strain CEN.PK113-7D. FEMS Yeast Res 17(7). https://doi.org/10.1093/femsyr/fox074
Santivasi WL, Xia F (2014) Ionizing radiation-induced DNA damage, response, and repair. Antioxid Redox Signal 21(2):251–259. https://doi.org/10.1089/ars.2013.5668
Schillingtóth B, Sándor N, Kis E, Kadhim M, Sáfrány G, Hegyesi H (2011) Analysis of the common deletions in the mitochondrial DNA is a sensitive biomarker detecting direct and non-targeted cellular effects of low dose ionizing radiation. Mutat Res-Fund Mol M 716(1–2):33–39. https://doi.org/10.1016/j.mrfmmm.2011.07.018
Seifert EL, Fiehn O, Bezaire V, Bickel DR, Wohlgemuth G, Adams SH, Harper ME (2010) Long-chain fatty acid combustion rate is associated with unique metabolite profiles in skeletal muscle mitochondria. PLoS One 5(3):e9834. https://doi.org/10.1371/journal.pone.0009834
Shaughnessy DT, McAllister K, Worth L, Haugen AC, Meyer JN, Domann FE, Houten BV, Mostoslavsky R, Bultman SJ, Baccarelli AA (2014) Mitochondria, energetics, epigenetics, and cellular responses to stress. Environ Health Perspect 122(12):1271–1278. https://doi.org/10.1289/ehp.1408418
Solieri L (2010) Mitochondrial inheritance in budding yeasts: towards an integrated understanding. Trends Microbiol 18(11):521–530. https://doi.org/10.1016/j.tim.2010.08.001
Strope PK, Skelly DA, Kozmin SG, Mahadevan G, Stone EA, Magwene PM, Dietrich FS, McCusker JH (2015) The 100-genomes strains, an S. cerevisiae resource that illuminates its natural phenotypic and genotypic variation and emergence as an opportunistic pathogen. Genome Res 25(5):762–774. https://doi.org/10.1101/gr.185538.114
Szumiel I (2015) Ionizing radiation-induced oxidative stress, epigenetic changes and genomic instability: the pivotal role of mitochondria. Int J Radiat Biol 91(1):1–12. https://doi.org/10.3109/09553002.2014.934929
Taanman JW (1999) The mitochondrial genome: structure, transcription, translation and replication. B BA-Bioenergetics 1410(2):103–123. https://doi.org/10.1016/S0005-2728(98)00161-3
Tanaka A, Shikazono N, Hase Y (2010) Studies on biological effects of ion beams on lethality, molecular nature of mutation, mutation rate, and spectrum of mutation phenotype for mutation breeding in higher plants. J Radiat Res 51(3):223–233. https://doi.org/10.1269/jrr.09143
Terato H, Tanaka R, Nakaarai Y, Nohara T, Doi Y, Iwai S, Hirayama R, Furusawa Y, Ide H (2008) Quantitative analysis of isolated and clustered DNA damage induced by gamma-rays, carbon ion beams, and iron ion beams. J Radiat Res 49(2):133–146. https://doi.org/10.1269/jrr.07089
Wallace DC (1992) Diseases of the mitochondrial DNA. Annu Rev Biochem 61(61):1175–1212. https://doi.org/10.1146/annurev.bi.61.070192.005523
Wang Y, Xu C, Du LQ, Cao J, Liu JX, Su X, Zhao H, Fan FY, Wang B, Katsube T (2013) Evaluation of the comet assay for assessing the dose-response relationship of DNA damage induced by ionizing radiation. Int J Mol Sci 14(11):22449–22461. https://doi.org/10.3390/ijms141122449
Xu A, Yao J, Yu L, Lv S, Wang J, Yan B, Yu Z (2004) Mutation of Gluconobacter oxydans and Bacillus megaterium in a two-step process of L-ascorbic acid manufacture by ion beam. J Appl Microbiol 96(6):1317–1323. https://doi.org/10.1111/j.1365-2672.2004.02270.x
Xu TT, Bai ZZ, Wang LJ, He BF (2010) Breeding of D(-)-lactic acid high producing strain by low-energy ion implantation and preliminary analysis of related metabolism. Applied Biochem Biotech 160(2):314–321. https://doi.org/10.1007/s12010-008-8274-4
Zhang H, Lu D, Li X, Feng Y, Cui Q, Song X (2018a) Heavy ion mutagenesis combined with triclosan screening provides a new strategy for improving the arachidonic acid yield in Mortierella alpina. BMC Biotechnol 18(1):23. https://doi.org/10.1186/s12896-018-0437-y
Zhang MM, Cao GZ, Guo XP, Gao Y, Li WJ, Lu D (2018b) A comet assay for DNA damage and repair after exposure to carbon-ion beams or X-rays in Saccharomyces cerevisiae. Dose-Response 16(3):1–9. https://doi.org/10.1177/1559325818792467
Zhang W, Zhao G, Luo Z, Lin Y, Wang L, Guo Y, Wang A, Jiang S, Jiang Q, Gong J (2017) Engineering the ribosomal DNA in a megabase synthetic chromosome. Science 355(6329):eaaf3981. https://doi.org/10.1126/science.aaf3981
Zhao XT, Feng JB, Yu Wen LI, Luo Q, Yang XC, Xue LU, Chen DQ, Liu QJ (2012) Identification of two novel mitochondrial DNA deletions induced by ionizing radiation. Biomed Environ Sci 25(5):533–541. https://doi.org/10.3967/0895-3988.2012.05.006
Zhou X, Li N, Wang Y, Wang Y, Zhang X, Zhang H (2011) Effects of X-irradiation on mitochondrial DNA damage and its supercoiling formation change. Mitochondrion 11(6):886–892. https://doi.org/10.1016/j.mito.2011.07.005
Acknowledgements
The authors would like to thank the colleagues at HIRFL for providing high-quality carbon ion beam irradiation.
Funding
This work was funded by the Chinese Academy of Sciences Key Deployment Project (No. KFZD-SW-109), a Joint project of the Chinese Academy of Sciences and the Industrial Technology Research Institute (CAS-ITRI 201801) and the National Natural Science Fund of China (No. 11575259).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there are no conflicts of interest.
Human and animal rights
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 1039 kb)
Rights and permissions
About this article
Cite this article
Guo, X., Zhang, M., Gao, Y. et al. A genome-wide view of mutations in respiration-deficient mutants of Saccharomyces cerevisiae selected following carbon ion beam irradiation. Appl Microbiol Biotechnol 103, 1851–1864 (2019). https://doi.org/10.1007/s00253-019-09626-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-019-09626-0