Abstract
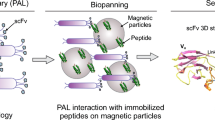
Infection with Helicobacter pylori may result in the emergence of gastric adenocarcinoma. Among various toxins assisting pathogenesis of H. pylori, the vacuolating cytotoxin A (VacA) is one of the most potent toxins known as the major cause of the peptic ulcer and gastric adenocarcinoma. To isolate single-chain variable fragments (scFvs) against two conserved regions of VacA, we capitalized on the phage display technology and a solution-phase biopanning (SPB). Characterization of scFvs was carried out by enzyme-linked immunosorbent assay (ELISA), immunoblotting, and surface plasmon resonance (SPR). Bioinformatics analyses were also performed in order to characterize the structural and functional properties of the isolated scFvs and the interaction(s) between the isolated antibodies (Ab)-antigen (Ag). After four rounds of biopanning, the positive colonies detected by scFv ELISA were harvested to extract the plasmids and perform sequencing. Of several colonies, three colonies showed high affinity to the VacA1 and two colonies for the VacA2. Further complementary examinations (e.g., sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), western blot, SPR, and flow cytometry) displayed the high affinity and specificity of the isolated scFvs to the VacA. Docking results revealed the interaction of the complementarity-determining regions (CDRs) with the VacA peptide. In conclusion, for the first time, we report on the isolation of several scFvs against conserved residues of VacA toxin with high affinity and specificity, which may be used as novel diagnostic/therapeutic tool in the H. pylori infection.









Similar content being viewed by others
References
Adler I, Muino A, Aguas S, Harada L, Diaz M, Lence A, Labbrozzi M, Muino JM, Elsner B, Avagnina A, Denninghoff V (2014) Helicobacter pylori and oral pathology: relationship with the gastric infection. World J Gastroenterol 20(29):9922–9935. https://doi.org/10.3748/wjg.v20.i29.9922
Aghebati-Maleki L, Bakhshinejad B, Baradaran B, Motallebnezhad M, Aghebati-Maleki A, Nickho H, Yousefi M, Majidi J (2016) Phage display as a promising approach for vaccine development. J Biomed Sci 23(1):66. https://doi.org/10.1186/s12929-016-0285-9
Ahmad ZA, Yeap SK, Ali AM, Ho WY, Alitheen NB, Hamid M (2012) scFv antibody: principles and clinical application. Clin Dev Immunol 2012:980250. https://doi.org/10.1155/2012/980250
Ayriss J, Woods T, Bradbury A, Pavlik P (2007) High-throughput screening of single-chain antibodies using multiplexed flow cytometry. J Proteome Res 6(3):1072–1082. https://doi.org/10.1021/pr0604108
Barderas R, Shochat S, Martinez-Torrecuadrada J, Altschuh D, Meloen R, Ignacio Casal J (2006) A fast mutagenesis procedure to recover soluble and functional scFvs containing amber stop codons from synthetic and semisynthetic antibody libraries. J Immunol Methods 312(1–2):182–189. https://doi.org/10.1016/j.jim.2006.03.005
Berdoz J, Corthesy B (2004) Human polymeric IgA is superior to IgG and single-chain Fv of the same monoclonal specificity to inhibit urease activity associated with Helicobacter pylori. Mol Immunol 41(10):1013–1022. https://doi.org/10.1016/j.molimm.2004.05.006
Bessede E, Dubus P, Megraud F, Varon C (2015) Helicobacter pylori infection and stem cells at the origin of gastric cancer. Oncogene 34(20):2547–2555. https://doi.org/10.1038/onc.2014.187
Brylinski M, Skolnick J (2008) A threading-based method (FINDSITE) for ligand-binding site prediction and functional annotation. Proc Natl Acad Sci U S A 105(1):129–134. https://doi.org/10.1073/pnas.0707684105
Capra JA, Laskowski RA, Thornton JM, Singh M, Funkhouser TA (2009) Predicting protein ligand binding sites by combining evolutionary sequence conservation and 3D structure. PLoS Comput Biol 5(12):e1000585. https://doi.org/10.1371/journal.pcbi.1000585
Capurso G, Carnuccio A, Lahner E, Panzuto F, Baccini F, Delle Fave G, Annibale B (2006) Corpus-predominant gastritis as a risk factor for false-negative 13C-urea breath test results. Aliment Pharmacol Ther 24(10):1453–1460. https://doi.org/10.1111/j.1365-2036.2006.03143.x
Carvalho MA, Machado NC, Ortolan EV, Rodrigues MA (2012) Upper gastrointestinal histopathological findings in children and adolescents with nonulcer dyspepsia with Helicobacter pylori infection. J Pediatr Gastroenterol Nutr 55(5):523–529. https://doi.org/10.1097/MPG.0b013e3182618136
Chey WD, Wong BC, Practice Parameters Committee of the American College of G (2007) American College of Gastroenterology guideline on the management of Helicobacter pylori infection. Am J Gastroenterol 102(8):1808–1825. https://doi.org/10.1111/j.1572-0241.2007.01393.x
Colovos C, Yeates TO (1993) Verification of protein structures: patterns of nonbonded atomic interactions. Protein Sci 2(9):1511–1519. https://doi.org/10.1002/pro.5560020916
Cover TL, Holland RL, Blanke SR (2016) Helicobacter pylori vacuolating toxin. In: Backert S, Yamaoka Y (eds) Helicobacter pylori research: from bench to bedside. Springer, Tokyo, pp 113–141
Dundas J, Ouyang Z, Tseng J, Binkowski A, Turpaz Y, Liang J (2006) CASTp: computed atlas of surface topography of proteins with structural and topographical mapping of functionally annotated residues. Nucleic Acids Res 34(Web Server):W116–W118. https://doi.org/10.1093/nar/gkl282
Eisenberg D, Luthy R, Bowie JU (1997) VERIFY3D: assessment of protein models with three-dimensional profiles. Methods Enzymol 277:396–404
Fahimi F, Tohidkia MR, Fouladi M, Aghabeygi R, Samadi N, Omidi Y (2017) Pleiotropic cytotoxicity of VacA toxin in host cells and its impact on immunotherapy. Bioimpacts 7(1):59–71. https://doi.org/10.15171/bi.2017.08
Fellouse FA, Pal G (2015) Methods for the construction of phage-displayed libraries In: Sidhu SS, Geyer CR (eds) Phage display in biotechnology and drug discovery. CRC Press
Figura N, Valassina M, Moretti E, Vindigni C, Collodel G, Iacoponi F, Giordano N, Roviello F, Marrelli D (2015) Histological variety of gastric carcinoma and Helicobacter pylori cagA and vacA polymorphism. Eur J Gastroenterol Hepatol 27(9):1017–1021. https://doi.org/10.1097/MEG.0000000000000414
Forsstrom B, Axnas BB, Rockberg J, Danielsson H, Bohlin A, Uhlen M (2015) Dissecting antibodies with regards to linear and conformational epitopes. PLoS One 10(3):e0121673. https://doi.org/10.1371/journal.pone.0121673
Gangwer KA, Mushrush DJ, Stauff DL, Spiller B, McClain MS, Cover TL, Lacy DB (2007) Crystal structure of the Helicobacter pylori vacuolating toxin p55 domain. Proc Natl Acad Sci U S A 104(41):16293–16298. https://doi.org/10.1073/pnas.0707447104
Garza-Gonzalez E, Perez-Perez GI, Maldonado-Garza HJ, Bosques-Padilla FJ (2014) A review of Helicobacter pylori diagnosis, treatment, and methods to detect eradication. World J Gastroenterol 20(6):1438–1449. https://doi.org/10.3748/wjg.v20.i6.1438
Georgieva Y, Konthur Z (2011) Design and screening of M13 phage display cDNA libraries. Molecules 16(2):1667–1681. https://doi.org/10.3390/molecules16021667
Hagymasi K, Tulassay Z (2014) Helicobacter pylori infection: new pathogenetic and clinical aspects. World J Gastroenterol 20(21):6386–6399. https://doi.org/10.3748/wjg.v20.i21.6386
Haque A, Tonks NK (2012) The use of phage display to generate conformation-sensor recombinant antibodies. Nat Protoc 7(12):2127–2143
Hensel F, Knorr C, Hermann R, Krenn V, Muller-Hermelink HK, Vollmers HP (1999) Mitogenic autoantibodies in Helicobacter pylori-associated stomach cancerogenesis. Int J Cancer 81(2):229–235
Jain AK, Agarwal A, Agrawal H, Agrawal GP (2012) Double-liposome-based dual-drug delivery system as vectors for effective management of peptic ulcer. J Liposome Res 22(3):205–214. https://doi.org/10.3109/08982104.2012.655284
Jain AK, Jain SK (2013) Development and characterization of nanolipobeads-based dual drug delivery system for H. pylori targeting. J Drug Target 21(6):593–603. https://doi.org/10.3109/1061186x.2013.784978
Jensen KB, Larsen M, Pedersen JS, Christensen PA, Alvarez-Vallina L, Goletz S, Clark BF, Kristensen P (2002) Functional improvement of antibody fragments using a novel phage coat protein III fusion system. Biochem Biophys Res Commun 298(4):566–573
Khoury GA, Tamamis P, Pinnaduwage N, Smadbeck J, Kieslich CA, Floudas CA (2014) Princeton_TIGRESS: protein geometry refinement using simulations and support vector machines. Proteins 82(5):794–814. https://doi.org/10.1002/prot.24459
Kohler G, Milstein C (1975) Continuous cultures of fused cells secreting antibody of predefined specificity. Nature 256(5517):495–497
Krieger E, Koraimann G, Vriend G (2002) Increasing the precision of comparative models with YASARA NOVA—a self-parameterizing force field. Proteins 47(3):393–402
Krieger E, Vriend G (2014) YASARA view—molecular graphics for all devices—from smartphones to workstations. Bioinformatics 30(20):2981–2982. https://doi.org/10.1093/bioinformatics/btu426
Kuo SH, Yeh KH, Chen LT, Lin CW, Hsu PN, Hsu C, Wu MS, Tzeng YS, Tsai HJ, Wang HP, Cheng AL (2014) Helicobacter pylori-related diffuse large B-cell lymphoma of the stomach: a distinct entity with lower aggressiveness and higher chemosensitivity. Blood Cancer J 4:e220. https://doi.org/10.1038/bcj.2014.40
Lim SL, Canavarro C, Zaw MH, Zhu F, Loke WC, Chan YH, Yeoh KG (2013) Irregular meal timing is associated with Helicobacter pylori infection and gastritis. ISRN Nutr 2013:714970. https://doi.org/10.5402/2013/714970
Liu JK (2014) The history of monoclonal antibody development—progress, remaining challenges and future innovations. Ann Med Surg (Lond) 3(4):113–116. https://doi.org/10.1016/j.amsu.2014.09.001
Oleastro M, Menard A (2013) The role of Helicobacter pylori outer membrane proteins in adherence and pathogenesis. Biology (Basel) 2(3):1110–1134. https://doi.org/10.3390/biology2031110
Palframan SL, Kwok T, Gabriel K (2012) Vacuolating cytotoxin A (VacA), a key toxin for Helicobacter pylori pathogenesis. Front Cell Infect Microbiol 2:92. https://doi.org/10.3389/fcimb.2012.00092
Papini E, Zoratti M, Cover TL (2001) In search of the Helicobacter pylori VacA mechanism of action. Toxicon 39(11):1757–1767
Parra RG, Schafer NP, Radusky LG, Tsai MY, Guzovsky AB, Wolynes PG, Ferreiro DU (2016) Protein Frustratometer 2: a tool to localize energetic frustration in protein molecules, now with electrostatics. Nucleic Acids Res 44(W1):W356–W360. https://doi.org/10.1093/nar/gkw304
Patel SK, Pratap CB, Jain AK, Gulati AK, Nath G (2014) Diagnosis of Helicobacter pylori: what should be the gold standard? World J Gastroenterol 20(36):12847–12859. https://doi.org/10.3748/wjg.v20.i36.12847
Pourakbari B, Ghazi M, Mahmoudi S, Mamishi S, Azhdarkosh H, Najafi M, Kazemi B, Salavati A, Mirsalehian A (2013) Diagnosis of Helicobacter pylori infection by invasive and noninvasive tests. Braz J Microbiol 44(3):795–798. https://doi.org/10.1590/S1517-83822013005000052
Pritchard DM, Crabtree JE (2006) Helicobacter pylori and gastric cancer. Curr Opin Gastroenterol 22(6):620–625. https://doi.org/10.1097/01.mog.0000245539.50765.f6
Reichert JM (2012) Marketed therapeutic antibodies compendium. MAbs 4(3):413–415. https://doi.org/10.4161/mabs.19931
Rieder G, Fischer W, Haas R (2005) Interaction of Helicobacter pylori with host cells: function of secreted and translocated molecules. Curr Opin Microbiol 8(1):67–73. https://doi.org/10.1016/j.mib.2004.12.004
Roy A, Kucukural A, Zhang Y (2010) I-TASSER: a unified platform for automated protein structure and function prediction. Nat Protoc 5(4):725–738. https://doi.org/10.1038/nprot.2010.5
Roy A, Yang J, Zhang Y (2012) COFACTOR: an accurate comparative algorithm for structure-based protein function annotation. Nucleic Acids Res 40(Web Server issue):W471–W477. https://doi.org/10.1093/nar/gks372
Salama NR, Hartung ML, Muller A (2013) Life in the human stomach: persistence strategies of the bacterial pathogen Helicobacter pylori. Nat Rev Microbiol 11(6):385–399. https://doi.org/10.1038/nrmicro3016
Schindler CE, de Vries SJ, Zacharias M (2015) Fully blind peptide-protein docking with pepATTRACT. Structure 23(8):1507–1515. https://doi.org/10.1016/j.str.2015.05.021
Schrodinger, LLC (2015) The PyMOL Molecular Graphics System, version 1.8
Siddiqui MZ (2010) Monoclonal antibodies as diagnostics; an appraisal. Indian J Pharm Sci 72(1):12–17. https://doi.org/10.4103/0250-474X.62229
Sidhu SS (2001) Engineering M13 for phage display. Biomol Eng 18(2):57–63
Tikunova NV, Morozova VV (2009) Phage display on the base of filamentous bacteriophages: application for recombinant antibodies selection. Acta Nat 1(3):20–28
Tohidkia MR, Barar J, Asadi F, Omidi Y (2012) Molecular considerations for development of phage antibody libraries. J Drug Target 20(3):195–208. https://doi.org/10.3109/1061186X.2011.611517
Trott M, Weibeta S, Antoni S, Koch J, von Briesen H, Hust M, Dietrich U (2014) Functional characterization of two scFv-Fc antibodies from an HIV controller selected on soluble HIV-1 Env complexes: a neutralizing V3- and a trimer-specific gp41 antibody. PLoS One 9(5):e97478. https://doi.org/10.1371/journal.pone.0097478
Verma R, Boleti E, George AJ (1998) Antibody engineering: comparison of bacterial, yeast, insect and mammalian expression systems. J Immunol Methods 216(1–2):165–181
Wang X, Campoli M, Ko E, Luo W, Ferrone S (2004) Enhancement of scFv fragment reactivity with target antigens in binding assays following mixing with anti-tag monoclonal antibodies. J Immunol Methods 294(1–2):23–35. https://doi.org/10.1016/j.jim.2004.08.005
Wiederstein M, Sippl MJ (2007) ProSA-web: interactive web service for the recognition of errors in three-dimensional structures of proteins. Nucleic Acids Res 35(Web Server):W407–W410. https://doi.org/10.1093/nar/gkm290
Wu S, Zhang Y (2007) LOMETS: a local meta-threading-server for protein structure prediction. Nucleic Acids Res 35(10):3375–3382. https://doi.org/10.1093/nar/gkm251
Yan R, Xu D, Yang J, Walker S, Zhang Y (2013) A comparative assessment and analysis of 20 representative sequence alignment methods for protein structure prediction. Sci Rep 3:2619. https://doi.org/10.1038/srep02619
Yang J, Roy A, Zhang Y (2013) Protein-ligand binding site recognition using complementary binding-specific substructure comparison and sequence profile alignment. Bioinformatics 29(20):2588–2595. https://doi.org/10.1093/bioinformatics/btt447
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12(1):7–8. https://doi.org/10.1038/nmeth.3213
Yang J, Zhang Y (2015) I-TASSER server: new development for protein structure and function predictions. Nucleic Acids Res 43(W1):W174–W181. https://doi.org/10.1093/nar/gkv342
Zhang Y (2008) I-TASSER server for protein 3D structure prediction. BMC Bioinformatics 9:40. https://doi.org/10.1186/1471-2105-9-40
Zhang Y, Skolnick J (2004) SPICKER: a clustering approach to identify near-native protein folds. J Comput Chem 25(6):865–871. https://doi.org/10.1002/jcc.20011
Zhao A, Tohidkia MR, Siegel DL, Coukos G, Omidi Y (2016) Phage antibody display libraries: a powerful antibody discovery platform for immunotherapy. Crit Rev Biotechnol 36(2):276–289. https://doi.org/10.3109/07388551.2014.958978
Acknowledgments
The authors wish to thank all the members of the cell culture facility at the Research Center for Pharmaceutical Nanotechnology at Tabriz University of Medical Sciences for their help. The authors also thanks Dr. Morteza Eskandani, Mr. Abolfazl Barzegari, Ms. Farzaneh Fathi, Dr. Morteza Milani, Dr. Mohammadreza Sadeghi and Dr. Ailar Nakhlband for their assistance during this research.
Funding
The study was funded by postgraduate grant of the Research Center for Pharmaceutical Nanotechnology (RCPN) at Tabriz University of Medical Sciences (RCPN-93005; recipient Dr. M.R. Tohidkia). This work was a joint project for a postgraduate thesis between the RCPN and the School of Advanced Biomedical Sciences at Tabriz University of Medical Sciences.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
The present study does not contain any experiments in relation to either human participants or animal models by any of the authors.
Electronic supplementary material
ESM 1
(DOCX 24.5 MB)
Rights and permissions
About this article
Cite this article
Fahimi, F., Sarhaddi, S., Fouladi, M. et al. Phage display-derived antibody fragments against conserved regions of VacA toxin of Helicobacter pylori. Appl Microbiol Biotechnol 102, 6899–6913 (2018). https://doi.org/10.1007/s00253-018-9068-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9068-4