Abstract
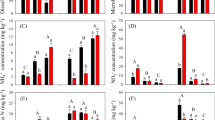
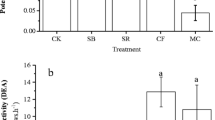
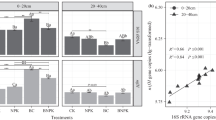
Preferable inorganic fertilization over the last decades has led to fertility degradation of black soil in Northeast China. However, how fertilization regimes impact denitrification and its related bacterial community in this soil type is still unclear. Here, taking advantage of a suit of molecular ecological tools in combination of assaying the potential denitrification (DP), we explored the variation of activity, community structure, and abundance of nirS and nirK denitrifiers under four different fertilization regimes, namely no fertilization control (N0M0), organic pig manure (N0M1), inorganic fertilization (N1M0), and combination of inorganic fertilizer and pig manure (N1M1). The results indicated that organic fertilization increased DP, but inorganic fertilization had no impacts. The increase of DP was mirrored by the shift of nirS denitrifiers’ community structure but not by that of nirK denitrifiers’. Furthermore, the change of DP coincided with the variation of abundances of both denitrifiers. Shifts of community structure and abundance of nirS and nirK denitrifiers were correlated with the change of soil pH, total nitrogen (TN), organic matter (OM), C:P, total phosphorus (TP), and available phosphorus (Olsen P). Our results suggest that the change of DP under these four fertilization regimes was closely related to the shift of denitrifying bacteria communities resulting from the variation of properties in the black soil tested.




Similar content being viewed by others
References
Attard E, Recous S, Chabbi A, De Berranger C, Guillaumaud N, Labreuche J, Philippot L, Schmid B, Le Roux X (2011) Soil environmental conditions rather than denitrifier abundance and diversity drive potential denitrification after changes in land uses. Glob Chang Biol 17:1975–1989
Báttard E, Rechová T, Vaněk D, Picek T, Šantrůčková H (2010) Effect of pH and dissolved organic matter on the abundance of nirK and nirS denitrifiers in spruce forest soil. Biogeochemistry 101:123–132
Bender SF, Plantenga F, Neftel A, Jocher M, Oberholzer HR, Köhl L, Giles M, Daniell TJ, van der Heijden MG (2014) Symbiotic relationships between soil fungi and plants reduce N2O emissions from soil. ISME J 8:1336–1345
Braker G, Fesefeldt A, Witzel KP (1998) Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl Environ Microbiol 64:3769–3775
Braker G, Schwarz J, Conrad R (2010) Influence of temperature on the composition and activity of denitrifying soil communities. FEMS Microbiol Ecol 73:134–148
Braker G, Dörsch P, Bakken LR (2012) Genetic characterization of denitrifier communities with contrasting intrinsic functional traits. FEMS Microbiol Ecol 79:542–554
Bremer C, Braker G, Matthies D, Reuter A, Engels C, Conrad R (2007) Impact of plant functional group, plant species, and sampling time on the composition of nirK-type denitrifier communities in soil. Appl Environ Microbiol 73:6876–6884
Chen Z, Luo X, Hu R, Wu M, Wu J, Wei W (2010) Impact of long-term fertilization on the composition of denitrifier communities based on nitrite reductase analyses in a paddy soil. Microb Ecol 60:850–861
Cui P, Fan F, Yin C, Li Z, Song A, Wan Y, Liang Y (2013) Urea-and nitrapyrin-affected N2O emission is coupled mainly with ammonia oxidizing bacteria growth in microcosms of three typical Chinese arable soils. Soil Biol Biochem 66:214–221
Culman SW, Bukowski R, Gauch HG, Cadillo-Quiroz H, Buckley DH (2009) T-REX: software for the processing and analysis of T-RFLP data. BMC Bioinformatics 10:171
Dambreville C, Hallet S, Nguyen C, Morvan T, Germon JC, Philippot L (2006) Structure and activity of the denitrifying community in a maize-cropped field fertilized with composted pig manure or ammonium nitrate. FEMS Microbiol Ecol 56:119–131
Dandie CE, Burton DL, Zebarth BJ, Henderson SL, Trevors JT, Goyer C (2008) Changes in bacterial denitrifier community abundance over time in an agricultural field and their relationship with denitrification activity. Appl Environ Microbiol 74:5997–6005
Dandie CE, Wertz S, Leclair CL, Goyer C, Burton DL, Patten CL, Zebarth BJ, Trevors JT (2011) Abundance, diversity and functional gene expression of denitrifier communities in adjacent riparian and agricultural zones. FEMS Microbiol Ecol 77:69–82
Enwall K, Philippot L, Hallin S (2005) Activity and composition of the denitrifying bacterial community respond differently to long-term fertilization. Appl Environ Microbiol 71:8335–8343
Enwall K, Throbäck IN, Stenberg M, Söderström M, Hallin S (2010) Soil resources influence spatial patterns of denitrifying communities at scales compatible with land management. Appl Environ Microbiol 76:2243–2250
Fan F, Yang Q, Li Z, Wei D, Cui X, Liang Y (2011) Impacts of organic and inorganic fertilizers on nitrification in a cold climate soil are linked to the bacterial ammonia oxidizer community. Microb Ecol 62:982–990
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci U S A 103:626–631
Gyaneshwar P, Kumar GN, Parekh L, Poole P (2002) Role of soil microorganisms in improving P nutrition of plants, food security in nutrient-stressed environments: exploiting plants’ genetic capabilities. Springer, pp 133-143
Hallin S, Jones CM, Schloter M, Philippot L (2009) Relationship between N-cycling communities and ecosystem functioning in a 50-year-old fertilization experiment. ISME J 3:597–605
He JZ, Zheng Y, Chen CR, He YQ, Zhang LM (2008) Microbial composition and diversity of an upland red soil under long-term fertilization treatments as revealed by culture-dependent and culture-independent approaches. J Soil Sediments 8:349–358
Henderson SL, Dandie CE, Patten CL, Zebarth BJ, Burton DL, Trevors JT, Goyer C (2010) Changes in denitrifier abundance, denitrification gene mRNA levels, nitrous oxide emissions, and denitrification in anoxic soil microcosms amended with glucose and plant residues. Appl Environ Microbiol 76:2155–2164
Henry S, Baudoin E, Lopez-Gutierrez JC, Martin-Laurent F, Brauman A, Philippot L (2004) Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. J Microbiol Methods 59:327–335
Heylen K, Gevers D, Vanparys B, Wittebolle L, Geets J, Boon N, De Vos P (2006) The incidence of nirS and nirK and their genetic heterogeneity in cultivated denitrifiers. Environ Microbiol 8:2012–2021
Hofstra N, Bouwman A (2005) Denitrification in agricultural soils: summarizing published data and estimating global annual rates. Nutr Cycl Agroecosyst 72:267–278
Jones CM, Hallin S (2010) Ecological and evolutionary factors underlying global and local assembly of denitrifier communities. ISME J 4:633–641
Jones CM, Stres B, Rosenquist M, Hallin S (2008) Phylogenetic analysis of nitrite, nitric oxide, and nitrous oxide respiratory enzymes reveal a complex evolutionary history for denitrification. Mol Biol Evol 25:1955–1966
Kandeler E, Deiglmayr K, Tscherko D, Bru D, Philippot L (2006) Abundance of narG, nirS, nirK, and nosZ genes of denitrifying bacteria during primary successions of a glacier foreland. Appl Environ Microbiol 72:5957–5962
Lashof DA, Ahuja DR (1990) Relative contributions of greenhouse gas emissions to global warming. Nature 344:529–531
Li P, Lang M (2014) Gross nitrogen transformations and related N2O emissions in uncultivated and cultivated black soil. Biol Fertil Soils 50:197–206
Liu B, Mørkved PT, Frostegård Å, Bakken LR (2010) Denitrification gene pools, transcription and kinetics of NO, N2O and N2 production as affected by soil pH. FEMS Microbiol Ecol 72:407–417
Morales SE, Cosart T, Holben WE (2010) Bacterial gene abundances as indicators of greenhouse gas emission in soils. ISME J 4:799–808
Oksanen J, Kindt R, Legendre P, O'Hara RB (2007) Vegan: community ecology package version 1.8–5. Available at http://r-forge.r-project.org/projects/vegan/
Petersen DG, Blazewicz SJ, Firestone M, Herman DJ, Turetsky M, Waldrop M (2012) Abundance of microbial genes associated with nitrogen cycling as indices of biogeochemical process rates across a vegetation gradient in Alaska. Environ Microbiol 14:993–1008
Philippot L, Hallin S, Schloter M (2007) Ecology of denitrifying prokaryotes in agricultural soil. Adv Agron 96:249–305
R Team C (2012) R: a language and environment for statistical computing
Ravishankara A, Daniel JS, Portmann RW (2009) Nitrous oxide (N2O): the dominant ozone-depleting substance emitted in the 21st century. Science 326:123–125
Šimek M, Cooper J (2002) The influence of soil pH on denitrification: progress towards the understanding of this interaction over the last 50 years. Eur J Soil Sci 53:345–354
Simek M, Cooper JE, Picek T, Santruckova H (2000) Denitrification in arable soils in relation to their physico-chemical properties and fertilization practice. Soil Biol Biochem 32:101–110
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Tang H, Yan K, Zhang L, Chi F, Li Q, Lian S, Wei D (2010) Diversity analysis of nitrite reductase genes (nirS) in black soil under different long-term fertilization conditions. Ann Microbiol 60:97–104
Throbäck IN, Enwall K, Jarvis Å, Hallin S (2004) Reassessing PCR primers targeting nirS, nirK and nosZ genes for community surveys of denitrifying bacteria with DGGE. FEMS Microbiol Ecol 49:401–417
Wakelin SA, Macdonald LM, O'Callaghan M, Forrester ST, Condron LM (2014) Soil functional resistance and stability are linked to different ecosystem properties. Austral Ecol 39:522–531
Xu XZ, Xu Y, Chen SC, Xu SG, Zhang HW (2010) Soil loss and conservation in the black soil region of Northeast China: a retrospective study. Environ Sci Pol 13:793–800
Yin C, Fan F, Li Z, Song A, Zhu P, Peng C, Liang Y (2012) Influences of long-term application of organic and inorganic fertilizers on the composition and abundance of nirS-type denitrifiers in black soil. Environ Sci 33:3967–3975 (in Chinese)
Yoshida M, Ishii S, Otsuka S, Senoo K (2009) Temporal shifts in diversity and quantity of nirS and nirK in a rice paddy field soil. Soil Biol Biochem 41:2044–2051
Yoshida M, Ishii S, Otsuka S, Senoo K (2010) nirK-harboring denitrifiers are more responsive to denitrification-inducing conditions in rice paddy soil than nirS-harboring bacteria. Microbes Environ 25:45–48
Yu WT, Xu YG, Bi ML, Ma Q, Zhou H (2010) Activity and composition of ammonia-oxidizing bacteria in an aquic brown soil as influenced by land use and fertilization. Pedosphere 20:789–798
Zhao LP, Wang HB, Liu Y, Wang YL, Liu SX, Wang Y (2006) Mechanism of fertility degradation of black soil in corn belt of Songliao Plain. Acta Pedol Sin 1:79–84 (in Chinese)
Zhou ZF, Zheng YM, Shen JP, Zhang LM, He JZ (2011) Response of denitrification genes nirS, nirK, and nosZ to irrigation water quality in a Chinese agricultural soil. Environ Sci Pollut Res 18:1644–1652
Zumft WG (1997) Cell biology and molecular basis of denitrification. Microbiol Mol Biol Rev 61:533–616
Acknowledgments
This work was jointly supported by the grants from National Key Basic Research Support Foundation of China (NKBRSF) (Approved No. 2015CB150502), National Natural Science Foundation of China (Approved No. 41171208) and the 12th Five-Year Key Programs for Science and Technology Development of China “Study of Key Technologies for Alleviating Obstacle Factors and Improving Productivity of Low-yield Cropland” (Approved No. 2012BAD05B06). Thanks are given to Dr. Steven Wakelin from AgResearch Ltd, Lincoln Science Centre, New Zealand, for his careful reading of this manuscript. Thanks are also given to two anonymous reviewers for their thoughtful comments that help improve the manuscript.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 648 kb)
Rights and permissions
About this article
Cite this article
Yin, C., Fan, F., Song, A. et al. Denitrification potential under different fertilization regimes is closely coupled with changes in the denitrifying community in a black soil. Appl Microbiol Biotechnol 99, 5719–5729 (2015). https://doi.org/10.1007/s00253-015-6461-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-015-6461-0