Abstract
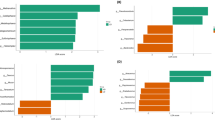
Present study described rumen microbiome of Indian cattle (Kankrej breed) to better understand the microbial diversity and largely unknown functional capacity of the rumen microbiome under different dietary treatments. Kankrej cattle were gradually adapted to a high-forage diet (four animals with dry forage and four with green forage) containing 50 % (K1), 75 % (K2) to 100 % (K3) forage, and remaining concentrate diet, each for 6 weeks followed by analysis of rumen fiber adherent and fiber-free metagenomic community by shotgun sequencing using ion torrent PGM platform and EBI-metagenomics annotation pipeline. Taxonomic analysis indicated that rumen microbiome was dominated by Bacteroidetes followed by Firmicutes, Fibrobacter, Proteobacteria, and Tenericutes. Functional analysis based on gene ontology classified all reads in total 157 categories based on their functional role in biological, molecular, and cellular component with abundance of genes associated with hydrolase activity, membrane, transport, transferase, and different metabolism (such as carbohydrate and protein). Statistical analysis using STAMP revealed significant differences (P < 0.05) between solid and liquid fraction of rumen (in 65 categories), between all three treatments (in 56 categories), and between green and dry roughage (17 categories). Diet treatment also exerted significant difference in environmental gene tags (EGTs) involved in metabolic pathways for production of volatile fatty acids. EGTs for butyrate production were abundant in K2, whereas EGTs for propionate production was abundant during K1. Principal component analysis also demonstrated that diet proportion, fraction of rumen, and type of forage affected rumen microbiome at taxonomic as well as functional level.





Similar content being viewed by others
References
Bergman EN (1990) Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol Rev 70:567–590
Bergman EN, Reid RS, Murray MG, Brockway JM, Whitelaw FG (1965) Interconversions and production of volatile fatty acids in the sheep rumen. Biochem J 97:53–58
Brulc JM, Antonopoulos DA, Miller ME, Wilson MK, Yannarell AC, Dinsdale EA, Edwards RE, Frank ED, Emerson JB, Wacklin P, Coutinhoj PM, Henrissatj B, Nelsoni KE, Whitea BA (2009) Gene-centric metagenomics of the fiber-adherent bovine rumen microbiome reveals forage specific glycoside hydrolases. Proc Natl Acad Sci U S A 106:1948–1953. doi:10.1073/pnas.0806191105
Cotta MA (1992) Interaction of ruminal bacteria in the production and utilization of maltooligosaccharides from starch. Appl Environ Microbiol 58:48–54
De Filippo C, Cavalieri D, Di Paola M, Ramazzotti M, Poullet JB, Massart S, Collini S, Pieraccini G, Lionetti P (2010) Impact of diet in shaping gut microbiota revealed by a comparative study in children from Europe and rural Africa. Proc Natl Acad Sci U S A 107:14691–14696. doi:10.1073/pnas.1005963107
de Menezes AB, Lewis E, O'Donovan M, O'Neill BF, Clipson N, Doyle EM (2011) Microbiome analysis of dairy cows fed pasture or total mixed ration diets. FEMS Microbiol Ecol 78:256–265. doi:10.1111/j.1574-6941.2011.01151.x
Dehority BA (1991) Effects of microbial synergism on fibre digestion in the rumen. Proc Nutr Soc 50:149–159
Fernando SC, Purvis HT 2nd, Najar FZ, Sukharnikov LO, Krehbiel CR, Nagaraja TG, Roe BA, Desilva U (2010) Rumen microbial population dynamics during adaptation to a high-grain diet. Appl Environ Microbiol 76:7482–7490. doi:10.1128/AEM. 00388-10
Firkins JL, Hristov AN, Hall MB, Varga GA, St-Pierre NR (2006) Integration of ruminal metabolism in dairy cattle. J Dairy Sci 89(Suppl 1):E31–E51. doi:10.3168/jds.S0022-0302(06)72362-1
Flint HJ (1997) The rumen microbial ecosystem—some recent developments. Trends Microbiol 5:483–488. doi:10.1016/S0966-842X(97)01159-1
Golder HM, Denman SE, McSweeney C, Celi P, Lean IJ (2014) Ruminal bacterial community shifts in grain-, sugar-, and histidine-challenged dairy heifers. J Dairy Sci 97:5131–5150. doi:10.3168/jds. 2014-8003
Gosalbes MJ, Durban A, Pignatelli M, Abellan JJ, Jimenez-Hernandez N, Perez-Cobas AE, Latorre A, Moya A (2011) Metatranscriptomic approach to analyze the functional human gut microbiota. PLoS One 6:e17447. doi:10.1371/journal.pone.0017447
Guan LL, Nkrumah JD, Basarab JA, Moore SS (2008) Linkage of microbial ecology to phenotype: correlation of rumen microbial ecology to cattle's feed efficiency. FEMS Microbiol Lett 288:85–91. doi:10.1111/j.1574-6968.2008.01343.x
Hammer O, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Hess M, Sczyrba A, Egan R, Kim TW, Chokhawala H, Schroth G, Luo S, Clark DS, Chen F, Zhang T, Mackie RI, Pennacchio LA, Tringe SG, Visel A, Woyke T, Wang Z, Rubin EM (2011) Metagenomic discovery of biomass-degrading genes and genomes from cow rumen. Science 331:463–467. doi:10.1126/science.1200387
Hook SE, Steele MA, Northwood KS, Wright AD, McBride BW (2011) Impact of high-concentrate feeding and low ruminal pH on methanogens and protozoa in the rumen of dairy cows. Microb Ecol 62:94–105. doi:10.1007/s00248-011-9881-0
Hunter S, Corbett M, Denise H, Fraser M, Gonzalez-Beltran A, Hunter C, Jones P, Leinonen R, McAnulla C, Maguire E, Maslen J, Mitchell A, Nuka G, Oisel A, Pesseat S, Radhakrishnan R, Rocca-Serra P, Scheremetjew M, Sterk P, Vaughan D, Cochrane G, Field D, Sansone S (2014) EBI metagenomics—a new resource for the analysis and archiving of metagenomic data. Nucleic Acids Res 42:D600–D606. doi:10.1093/nar/gkt961
Jami E, Mizrahi I (2012) Composition and similarity of bovine rumen microbiota across individual animals. PLoS One 7:e33306. doi:10.1371/journal.pone.0033306
Jami E, Israel A, Kotser A, Mizrahi I (2013) Exploring the bovine rumen bacterial community from birth to adulthood. ISME J 7:1069–1079. doi:10.1038/ismej.2013.2
Kong Y, Teather R, Forster R (2010) Composition, spatial distribution, and diversity of the bacterial communities in the rumen of cows fed different forages. FEMS Microbiol Ecol 74:612–622. doi:10.1111/j.1574-6941.2010.00977.x
Krause DO, Denman SE, Mackie RI, Morrison M, Rae AL, Attwood GT, McSweeney CS (2003) Opportunities to improve fiber degradation in the rumen: microbiology, ecology, and genomics. FEMS Microbiol Rev 27:663–693
Kurokawa K, Itoh T, Kuwahara T, Oshima K, Toh H, Toyoda A, Takami H, Morita H, Sharma VK, Srivastava TP, Taylor TD, Noguchi H, Mori H, Ogura Y, Ehrlich DS, Itoh K, Takagi T, Sakaki Y, Hayashi T, Hattori M (2007) Comparative metagenomics revealed commonly enriched gene sets in human gut microbiomes. DNA Res 14:169–181. doi:10.1093/dnares/dsm018
Lawrence P, Kenny DA, Earley B, Crews DH Jr, McGee M (2011) Grass silage intake, rumen and blood variables, ultrasonic and body measurements, feeding behavior, and activity in pregnant beef heifers differing in phenotypic residual feed intake. J Anim Sci 89:3248–3261. doi:10.2527/jas. 2010-3774
Lettat A, Hassanat F, Benchaar C (2013) Corn silage in dairy cow diets to reduce ruminal methanogenesis: effects on the rumen metabolically active microbial communities. J Dairy Sci 96:5237–5248. doi:10.3168/jds. 2012-6481
Li RW, Li C (2006) Butyrate induces profound changes in gene expression related to multiple signal pathways in bovine kidney epithelial cells. BMC Genomics 7:234. doi:10.1186/1471-2164-7-234
Li CJ, Li RW (2008) Butyrate induced cell cycle arrest in bovine cells through targeting gene expression relevant to DNA replication apparatus. Gene Regul Syst Biol 2:113–123
Macy JM, Probst I (1979) The biology of gastrointestinal bacteroides. Annu Rev Microbiol 33:561–594. doi:10.1146/annurev.mi.33.100179.003021
McCann JC, Wiley LM, Forbes TD, Rouquette FM, Tedeschi LO (2014) Relationship between the rumen microbiome and residual feed intake-efficiency of Brahman bulls stocked on bermudagrass pastures. PloS One 9:e91864 doi:10.1371/journal.pone.0091864
Matsui H, Ogata K, Tajima K, Nakamura M, Nagamine T, Aminov RI, Benno Y (2000) Phenotypic characterization of polysaccharidases produced by four Prevotella type strains. Curr Microbiol 41:45–49
Parks DH, Beiko RG (2010) Identifying biologically relevant differences between metagenomic communities. Bioinformatics 26:715–721. doi:10.1093/bioinformatics/btq041
Pitta DW, Pinchak E, Dowd SE, Osterstock J, Gontcharova V, Youn E, Dorton K, Yoon I, Min BR, Fulford JD, Wickersham TA, Malinowski DP (2010) Rumen bacterial diversity dynamics associated with changing from bermudagrass hay to grazed winter wheat diets. Microb Ecol 59:511–522. doi:10.1007/s00248-009-9609-6
Qin J, Li R, Raes J, Arumugam M, Burgdorf KS, Manichanh C, Nielsen T, Pons N, Levenez F, Yamada T, Daniel R, Mende LJ, Xu J, Li S, Li D, Cao J, Wang B, Liang H, Zheng H, Xie Y, Tap J, Lepage P, Bertalan M, Batto J, Hansen T, Le Paslier D, Linneberg A, Nielsen HB, Pelletier E, Renault P, Sicheritz-Ponten T, Turner K, Zhu H, Yu C, Li S, Jian M, Zhou Y, Li Y, Zhang X, Li S, Qin N, Yang H, Wang J, Brunak S, Doré J, Guarner F, Kristiansen K, Pedersen O, Parkhill J, Weissenbach J, Bork P, Ehrlich SD, Wang J (2010) A human gut microbial gene catalogue established by metagenomic sequencing. Nature 464:59–65. doi:10.1038/nature08821
Qu A, Brulc JM, Wilson MK, Law BF, Theoret JR, Joens LA, Konkel ME, Angly F, Dinsdale EA, Edwards RA, Nelson KE, White BA (2008) Comparative metagenomics reveals host specific metavirulomes and horizontal gene transfer elements in the chicken cecum microbiome. PLoS One 3:e2945. doi:10.1371/journal.pone.0002945
Reichardt N, Duncan SH, Young P, Belenguer A, McWilliam Leitch C, Scott KP, Flint HJ, Louis P (2014) Phylogenetic distribution of three pathways for propionate production within the human gut microbiota. ISME J 8:1323–1335. doi:10.1038/ismej.2014.14
Russell JB, Muck RE, Weimer PJ (2009) Quantitative analysis of cellulose degradation and growth of cellulolytic bacteria in the rumen. FEMS Microbiol Ecol 67:183–197. doi:10.1111/j.1574-6941.2008.00633.x
Sergeant MJ, Constantinidou C, Cogan TA, Bedford MR, Penn CW, Pallen MJ (2014) Extensive microbial and functional diversity within the chicken cecal microbiome. PLoS One 9:e91941. doi:10.1371/journal.pone.0091941
Singh KM, Ahir VB, Tripathi AK, Ramani UV, Sajnani M, Koringa PG, Jakhesara S, Pandya PR, Rank DN, Murty DS, Kothari RK, Joshi CG (2012) Metagenomic analysis of Surti buffalo (Bubalus bubalis) rumen: a preliminary study. Mol Biol Rep 39:4841–4848. doi:10.1007/s11033-011-1278-0
Stevenson DM, Weimer PJ (2007) Dominance of Prevotella and low abundance of classical ruminal bacterial species in the bovine rumen revealed by relative quantification real-time PCR. Appl Microbiol Biotechnol 75:165–174. doi:10.1007/s00253-006-0802-y
Tajima K, Aminov RI, Nagamine T, Matsui H, Nakamura M, Benno Y (2001) Diet-dependent shifts in the bacterial population of the rumen revealed with real-time PCR. Appl Environ Microbiol 67:2766–2774. doi:10.1128/AEM. 67.6.2766-2774.2001
Turnbaugh PJ (2009) A core gut microbiome in obese and lean twins. Nature 457:480–484. doi:10.1038/nature07540
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1031. doi:10.1038/nature05414
Turnbaugh PJ, Backhed F, Fulton L, Gordon JI (2008) Diet-induced obesity is linked to marked but reversible alterations in the mouse distal gut microbiome. Cell Host Microbe 3:213–223. doi:10.1016/j.chom.2008.02.015
Uden P, Rounsaville TR, Wiggans GR, Van Soest PJ (1982) The measurement of liquid and solid digesta retention in ruminants, equines and rabbits given timothy (Phleum pratense) hay. Br J Nutr 48:329–339
Verberkmoes NC, Russell AL, Shah M, Godzik A, Rosenquist M, Halfvarson J, Lefsrud MG, Apajalahti J, Tysk C, Hettich RL, Jansson JK (2009) Shotgun metaproteomics of the human distal gut microbiota. ISME J 3:179–189. doi:10.1038/ismej.2008.108
Walker AW, Duncan SH, Harmsen HJ, Holtrop G, Welling GW, Flint HJ (2008) The species composition of the human intestinal microbiota differs between particle-associated and liquid phase communities. Environ Microbiol 10:3275–3283. doi:10.1111/j.1462-2920.2008.01717.x
Wang X, Heazlewood SP, Krause DO, Florin TH (2003) Molecular characterization of the microbial species that colonize human ileal and colonic mucosa by using 16S rDNA sequence analysis. J Appl Microbiol 95:508–520
Whitford MF, Forster RJ, Beard CE, Gong J, Teather RM (1998) Phylogenetic analysis of rumen bacteria by comparative sequence analysis of cloned 16S rRNA genes. Anaerobe 4:153–163. doi:10.1006/anae.1998.0155
Wright AD, Klieve AV (2011) Does the complexity of the rumen microbial ecology preclude methane mitigation? Anim Feed Sci Technol 166–167:248–253
Acknowledgments
This work was supported by the Niche area of excellence program on Metagenomic analysis of ruminal microbes funded by Indian Council of Agriculture Research (ICAR), New Delhi, India
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 937 kb)
Rights and permissions
About this article
Cite this article
Patel, V., Patel, A.K., Parmar, N.R. et al. Characterization of the rumen microbiome of Indian Kankrej cattle (Bos indicus) adapted to different forage diet. Appl Microbiol Biotechnol 98, 9749–9761 (2014). https://doi.org/10.1007/s00253-014-6153-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-014-6153-1