Abstract
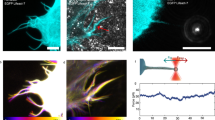
Filopodia are actin-built finger-like dynamic structures that protrude from the cell cortex. These structures can sense the environment and play key roles in migration and cell–cell interactions. The growth-retraction cycle of filopodia is a complex process exquisitely regulated by intra- and extra-cellular cues, whose nature remains elusive. Filopodia present wide variation in length, lifetime and growth rate. Here, we investigate the features of filopodia patterns in fixed prostate tumor cells by confocal microscopy. Analysis of almost a thousand filopodia suggests the presence of two different populations: one characterized by a narrow distribution of lengths and the other with a much more variable pattern with very long filopodia. We explore a stochastic model of filopodial growth which takes into account diffusion and reactions involving actin and the regulatory proteins formin and capping, and retrograde flow. Interestingly, we found an inverse dependence between the filopodial length and the retrograde velocity. This result led us to propose that variations in the retrograde velocity could explain the experimental lengths observed for these tumor cells. In this sense, one population involves a wider range of retrograde velocities than the other population, and also includes low values of this velocity. It has been hypothesized that cells would be able to regulate retrograde flow as a mechanism to control filopodial length. Thus, we propound that the experimental filopodia pattern is the result of differential retrograde velocities originated from heterogeneous signaling due to cell–substrate interactions or prior cell–cell contacts.






Similar content being viewed by others
References
Aberle H (2019) Axon guidance and collective cell migration by substrate-derived attractants. Front Mol Neurosci 12:148
Anderson TW, Vaughan AN, Cramer LP (2008) Retrograde flow and myosin ii activity within the leading cell edge deliver f-actin to the lamella to seed the formation of graded polarity actomyosin ii filament bundles in migrating fibroblasts. Mol Biol Cell 19(11):5006–5018
Arjonen A, Kaukonen R, Mattila E, Rouhi P, Högnäs G, Sihto H, Miller BW, Morton JP, Bucher E, Taimen P et al (2014) Mutant p53-associated myosin-x upregulation promotes breast cancer invasion and metastasis. J Clin Investig 124(3):1069–1082
Atilgan E, Wirtz D, Sun SX (2006) Mechanics and dynamics of actin-driven thin membrane protrusions. Biophys J 90(1):65–76
BenSaïda A (2020) Shapiro-wilk and shapiro-francia normality tests . https://www.mathworks.com/matlabcentral/fileexchange/13964-shapiro-wilk-and-shapiro-francia-normality-tests. MATLAB Central File Exchange. Retrieved 15 July 2020
Bornschlögl T (2013) How filopodia pull: what we know about the mechanics and dynamics of filopodia. Cytoskeleton 70(10):590–603
Bornschlögl T, Romero S, Vestergaard CL, Joanny JF, Van Nhieu GT, Bassereau P (2013) Filopodial retraction force is generated by cortical actin dynamics and controlled by reversible tethering at the tip. Proc Natl Acad Sci 110(47):18928–18933
Breitsprecher D, Koestler SA, Chizhov I, Nemethova M, Mueller J, Goode BL, Small JV, Rottner K, Faix J (2011) Cofilin cooperates with fascin to disassemble filopodial actin filaments. J Cell Sci 124(19):3305–3318
Campbell R (2018) notBoxPlot. https://github.com/raacampbell/notBoxPlot, GitHub. Retrieved 18 Dec 2018
Cohen M, Georgiou M, Stevenson NL, Miodownik M, Baum B (2010) Dynamic filopodia transmit intermittent delta-notch signaling to drive pattern refinement during lateral inhibition. Dev Cell 19(1):78–89
Daniels D (2010) Effect of capping protein on a growing filopodium. Biophys J 98(7):1139–1148
Dominguez R, Holmes KC (2011) Actin structure and function. Anuu Rev Biophys 40:169–186
Erban R, Flegg MB, Papoian GA (2014) Multiscale stochastic reaction-diffusion modeling: application to actin dynamics in filopodia. Bull Math Biol 76(4):799–818
Flegg MB, Chapman SJ, Erban R (2011) The two-regime method for optimizing stochastic reaction-diffusion simulations. J R Soc Interface 9:859–868
Flegg MB, Chapman SJ, Zheng L, Erban R (2014) Analysis of the two-regime method on square meshes. SIAM J Sci Comput 36(3):B561–B588
Gallo G, Letourneau PC (2004) Regulation of growth cone actin filaments by guidance cues. J Neurobiol 58(1):92–102
Goode BL, Eck MJ (2007) Mechanism and function of formins in the control of actin assembly. Annu Rev Biochem 76:593–627
Heckman CA, Plummer H III (2013) Filopodia as sensors. Cell Signal 25(11):2298–2311
Hoelzle MK, Svitkina T (2012) The cytoskeletal mechanisms of cell-cell junction formation in endothelial cells. Mol Biol Cell 23(2):310–323
Huang S (2009) Non-genetic heterogeneity of cells in development: more than just noise. Development 136(23):3853–3862
Husainy AN, Morrow AA, Perkins TJ, Lee JM (2010) Robust patterns in the stochastic organization of filopodia. BMC Cell Biol 11(1):86
Jacquemet G, Hamidi H, Ivaska J (2015) Filopodia in cell adhesion, 3d migration and cancer cell invasion. Curr Opin Cell Biol 36:23–31
Jacquemet G, Baghirov H, Georgiadou M, Sihto H, Peuhu E, Cettour-Janet P, He T, Perälä M, Kronqvist P, Joensuu H et al (2016) L-type calcium channels regulate filopodia stability and cancer cell invasion downstream of integrin signalling. Nat Commun 7(1):1–17
Jacquemet G, Paatero I, Carisey AF, Padzik A, Orange JS, Hamidi H, Ivaska J (2017) Filoquant reveals increased filopodia density during breast cancer progression. J Cell Biol 216(10):3387–3403
Jacquemet G, Stubb A, Saup R, Miihkinen M, Kremneva E, Hamidi H, Ivaska J (2019) Filopodome mapping identifies p130cas as a mechanosensitive regulator of filopodia stability. Curr Biol 29(2):202–216
Jang KJ, Kim MS, Feltrin D, Jeon NL, Suh KY, Pertz O (2010) Two distinct filopodia populations at the growth cone allow to sense nanotopographical extracellular matrix cues to guide neurite outgrowth. PLoS One 5(12):e15966
Jontes JD, Buchanan J, Smith SJ (2000) Growth cone and dendrite dynamics in zebrafish embryos: early events in synaptogenesis imaged in vivo. Nat Neurosci 3(3):231–237
Kapustina M, Vitriol E, Elston TC, Loew LM, Jacobson K (2010) Modeling capping protein frap and cali experiments reveals in vivo regulation of actin dynamics. Cytoskeleton 67(8):519–534
Kovar DR, Wu JQ, Pollard TD (2005) Profilin-mediated competition between capping protein and formin cdc12p during cytokinesis in fission yeast. Mol Biol Cell 16(5):2313–2324
Kovar DR, Harris ES, Mahaffy R, Higgs HN, Pollard TD (2006) Control of the assembly of atp-and adp-actin by formins and profilin. Cell 124(2):423–435
Lan Y, Papoian GA (2008) The stochastic dynamics of filopodial growth. Biophys J 94(10):3839–3852
Lin CH, Espreafico EM, Mooseker MS, Forscher P (1996) Myosin drives retrograde f-actin flow in neuronal growth cones. Neuron 16(4):769–782
Liou YR, Torng W, Kao YC, Sung KB, Lee CH, Kuo PL (2014) Substrate stiffness regulates filopodial activities in lung cancer cells. PLoS One 9(2):e89767
Marchenko OO, Das S, Yu J, Novak IL, Rodionov VI, Efimova N, Svitkina T, Wolgemuth CW, Loew LM (2017) A minimal actomyosin-based model predicts the dynamics of filopodia on neuronal dendrites. Mol Biol Cell 28(8):1021–1033
Mattila PK, Lappalainen P (2008) Filopodia: molecular architecture and cellular functions. Nat Rev Mol Biol Cell 9(6):446–454
McCroskery S, Chaudhry A, Lin L, Daniels MP (2006) Transmembrane agrin regulates filopodia in rat hippocampal neurons in culture. Mol Cell Neurosci 33(1):15–28
McGrath JL, Tardy Y, Dewey C Jr, Meister J, Hartwig J (1998) Simultaneous measurements of actin filament turnover, filament fraction, and monomer diffusion in endothelial cells. Biophys J 75(4):2070–2078
McMillen LM, Vavylonis D (2016) Model of turnover kinetics in the lamellipodium: implications of slow-and fast-diffusing capping protein and arp2/3 complex. Phys Biol 13(6):066009
Medeiros NA, Burnette DT, Forscher P (2006) Myosin ii functions in actin-bundle turnover in neuronal growth cones. Nat Cell Biol 8(3):216–226
Miller J, Fraser SE, McClay D (1995) Dynamics of thin filopodia during sea urchin gastrulation. Development 121(8):2501–2511
Mogilner A, Rubinstein B (2005) The physics of filopodial protrusion. Biophys J 89(2):782–795
Oldenbourg R, Katoh K, Danuser G (2000) Mechanism of lateral movement of filopodia and radial actin bundles across neuronal growth cones. Biophys J 78(3):1176–1182
Paez A, Vazquez E, Gueron G (2017) Heme oxygenase 1 governs the cytoskeleton at filopodia: pulling the brakes on the migratory capacity of prostate tumoral cells. Cell Death Discov 3(1):1–2
Paul NR, Allen JL, Chapman A, Morlan-Mairal M, Zindy E, Jacquemet G, Fernandez del Ama L, Ferizovic N, Green DM, Howe JD et al (2015) \(\alpha \)5\(\beta \)1 integrin recycling promotes arp2/3-independent cancer cell invasion via the formin fhod3. J Cell Biol 210(6):1013–1031
Peckham M (2016) How myosin organization of the actin cytoskeleton contributes to the cancer phenotype. Biochem Soc Trans 44(4):1026–1034
Peskin CS, Odell GM, Oster GF (1993) Cellular motions and thermal fluctuations: the brownian ratchet. Biophys J 65(1):316–324
Pollard TD, Blanchoin L, Mullins RD (2000) Molecular mechanisms controlling actin filament dynamics in nonmuscle cells. Annu Rev Biophys Biomol Struct 29(1):545–576
Roos C, Terlaky T, Vial JP (2005) Interior point methods for linear optimization. Springer Science & Business Media, Boston, MA
Saha T, Rathmann I, Viplav A, Panzade S, Begemann I, Rasch C, Klingauf J, Matis M, Galic M (2016) Automated analysis of filopodial length and spatially resolved protein concentration via adaptive shape tracking. Mol Biol Cell 27(22):3616–3626
Sanders TA, Llagostera E, Barna M (2013) Specialized filopodia direct long-range transport of shh during vertebrate tissue patterning. Nature 497(7451):628–632
Sasaki AT, Chun C, Takeda K, Firtel RA (2004) Localized ras signaling at the leading edge regulates pi3k, cell polarity, and directional cell movement. J Cell Biol 167(3):505–518
Shekhar S, Kerleau M, Kühn S, Pernier J, Romet-Lemonne G, Jégou A, Carlier MF (2015) Formin and capping protein together embrace the actin filament in a ménage à trois. Nat Commun 6(1):1–12
Shibue T, Brooks MW, Weinberg RA (2013) An integrin-linked machinery of cytoskeletal regulation that enables experimental tumor initiation and metastatic colonization. Cancer Cell 24(4):481–498
Sinnar SA, Antoku S, Saffin JM, Cooper JA, Halpain S (2014) Capping protein is essential for cell migration in vivo and for filopodial morphology and dynamics. Mol Biol Cell 25(14):2152–2160
Steffen A, Faix J, Resch GP, Linkner J, Wehland J, Small JV, Rottner K, Stradal TE (2006) Filopodia formation in the absence of functional wave-and arp2/3-complexes. Mol Biol Cell 17(6):2581–2591
Svitkina TM, Bulanova EA, Chaga OY, Vignjevic DM, Kojima SI, Vasiliev JM, Borisy GG (2003) Mechanism of filopodia initiation by reorganization of a dendritic network. J Cell Biol 160(3):409–421
Tatavarty V, Das S, Yu J (2012) Polarization of actin cytoskeleton is reduced in dendritic protrusions during early spine development in hippocampal neuron. Mol Biol Cell 23(16):3167–3177
Vasioukhin V, Bauer C, Yin M, Fuchs E (2000) Directed actin polymerization is the driving force for epithelial cell-cell adhesion. Cell 100(2):209–219
Vitriol EA, McMillen LM, Kapustina M, Gomez SM, Vavylonis D, Zheng JQ (2015) Two functionally distinct sources of actin monomers supply the leading edge of lamellipodia. Cell Rep 11(3):433–445
Wolff K, Barrett-Freeman C, Evans MR, Goryachev AB, Marenduzzo D (2014) Modelling the effect of myosin x motors on filopodia growth. Phys Biol 11(1):016005
Zhuravlev PI, Papoian GA (2009) Molecular noise of capping protein binding induces macroscopic instability in filopodial dynamics. Proc Natl Acad Sci 106(28):11570–11575. https://doi.org/10.1073/pnas.0812746106. https://www.pnas.org/content/106/28/11570
Zhuravlev PI, Papoian GA (2011) Protein fluxes along the filopodium as a framework for understanding the growth-retraction dynamics: the interplay between diffusion and active transport. Cell Adhesion Mig 5(5):448–456
Acknowledgements
We thank Luis Diambra for helpful discussions on the method to reconstruct the experimental distributions. We thank Carla Pallavicini for valuable advise on image analysis.
Author information
Authors and Affiliations
Contributions
LB and NG designed study, DS, LB and NG analyzed data, DS, AP, GG, LB and NG performed research, DS, LB and NG wrote the paper. LB and NG contributed equally to this study.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest or competing interests.
Ethics approval
Not applicable.
Funding
We acknowledge support from the Agencia Nacional de Promoción Científica y Tecnológica (PICT 2015-0370 and PICT-RAICES-2018-02639), Argentina.
Consent to participate
All the authors consent to participate.
Consent for publication
All the authors consent for publication.
Code availability
All the codes used in this work are available on request to the corresponding authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
We acknowledge support from the Agencia Nacional de Promoción Científica y Tecnológica (PICT 2015-0370 and PICT-RAICES-2018-02639), Argentina.
Rights and permissions
About this article
Cite this article
Senra, D., Páez, A., Gueron, G. et al. Following the footprints of variability during filopodial growth. Eur Biophys J 49, 643–659 (2020). https://doi.org/10.1007/s00249-020-01473-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-020-01473-6