Abstract
Bacterial symbionts are crucial to the biology of Bactrocera dorsalis. With larval diet (fruit host) being a key factor that determines microbiome composition and with B. dorsalis using more than 400 fruits as hosts, it is unclear if certain bacterial symbionts are preserved and are passed on to B. dorsalis progenies despite changes in larval diet. Here, we conducted a fly rearing experiment to characterize diet-induced changes in the microbiome of female B. dorsalis. In order to explicitly investigate the impacts of larval diet on the microbiome, including potential stable bacterial constituents of B. dorsalis, we performed 16S rRNA sequencing on the gut tissues of teneral female flies reared from four different host fruits (guava, mango, papaya, and rose apple) infested using a single cohort of wild B. dorsalis that emerged from tropical almond (mother flies). Although B. dorsalis-associated microbiota were predominantly shaped by the larval diet, some major bacterial species from the mother flies were retained in progenies raised on different larval diets. With some variation, Klebsiella (ASV 1 and 2), Morganella (ASV 3), and Providencia (ASV 6) were the major bacterial symbionts that were stable and made up 0.1–80% of the gut and ovipositor microbiome of female teneral flies reared on different host fruits. Our results suggest that certain groups of bacteria are stably associated with female B. dorsalis across larval diets. These findings provide a basis for unexplored research on symbiotic bacterial function in B. dorsalis and may aid in the development of novel management techniques against this devastating pest of horticultural importance.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bacterial symbionts can have crucial roles in the success of insects in their environment. They share common niches with insects and often engage in intricate ecological interactions that influence an insect’s overall fitness [1] by affecting their growth, fecundity, lifespan, and mate selection [2,3,4]. Additionally, bacteria may also mediate interactions with infectious disease and natural enemies [5, 6], impart temperature stress tolerance [7, 8], aid in the metabolism of xenobiotics [9, 10], and provide nutrients to their host insects [11, 12].
Bacterial symbionts can generally exist as obligate and facultative symbionts [13]. Obligate symbionts are essential for the host insect’s survival [14] and enable insects to survive adverse conditions [15], and in return, the host insect provides shelter and supplies nutrients to its bacterial partner [16]. Such symbiosis are cases of obligate symbiosis, as bot, the insect and its symbiotic bacteria suffer and even perish in the absence of each other [17]. Obligate symbionts are most often transmitted vertically from mother to offspring [18]. In contrast, facultative symbionts are not essential for the survival of the host insect but aid their hosts under certain conditions and are transmitted vertically or horizontally to the host’s offspring [13, 19]. An example of facultative symbionts is commensal bacteria, which can also endure in various niches in the absence of the host insect [20]. Previous studies have provided convincing evidence that commensal bacteria, despite being non-essential for the survival of host insect, contribute to important aspects of their host insect’s biology [3, 4, 7]. Thus, a thorough understanding of host-microbiome interactions also requires comprehensive knowledge of acquisition, transmission, host-associated bacterial communities, and the factors shaping them.
The acquisition and transmission of bacterial symbionts and pathogens in B. dorsalis often occur through contact with substrates with bacterial presence, such as fruit/leaf surfaces. However, the majority of bacterial symbionts are acquired within the larval feeding environment (host fruit or diet) that contains microbes deposited by the mother fly during oviposition. Although some studies have explored the dynamics of microbial communities in B. dorsalis [21], studies on bacteria that get retained by B. dorsalis even with the change of larval diet are limited.
Microbial communities in insects can be influenced by many factors with “larval diet” being a major contributor [21]. To explore factors shaping insect microbial communities, researchers often studied the association between host genetic divergence and diet. For example, the genetic correlation between host taxa and variation in the microbial community may suggest the role of genetic effects in shaping insect microbial communities, while the correlation between diet and microbial community composition may point to environmental effects. Although this approach has been applied in vertebrate systems [22,23,24], the factors that determine the microbiome composition in higher animals are not yet clear. This is partly due to the difficulty of controlling diet and other environmental factors and the complexity of microbial communities associated with vertebrates [25,26,27], which consist of thousands of taxa. In contrast, the gut microbiome of invertebrates, especially insects, is comprised of substantially fewer bacterial taxa [28, 29]. This makes insects as attractive models for disentangling complex host-microbial interactions [29, 30]. Furthermore, insects can serve as axenic and gnotobiotic models, and such experiments can widen our knowledge on insect-microbe interactions [31, 32].
The oriental fruit fly, Bactrocera dorsalis (Diptera: Tephritidae) is an important insect pest of horticultural crops around the globe [33]. This insect causes significant economic losses from fruit damage and export limitation due to quarantine issues [34,35,36]. Additionally, its utilization of a broad host range, temperature tolerance, and high mating success makes it a serious pest with extreme invasive potential [37,38,39], and these abilities may in part be attributed to their microbiome [4, 7]. Recently, there is growing evidence that there are key metabolic roles associated with bacterial symbionts found in a tephritid fly’s microbiome, and specific bacterial taxa may gain an advantage over others in different host fruits [21, 40, 41]. However, there is a possibility that more than one kind of bacterial taxa can fulfill a particular metabolic role (phenol degradation, cellulose degradation, etc.), thus prompting some plasticity in the microbiome composition of flies that emerged from different host fruits [21].
To understand the diversity and potentially stable constituents of bacterial communities associated with B. dorsalis, it is imperative to investigate teneral flies from different host fruits (larval diet) that have a known common wild fly mother. Therefore, we investigated the microbiome of teneral female flies from four different host fruits that were infested by a single cohort of wild flies that emerged from tropical almond (mother flies).
Materials and Methods
Collection and Rearing of Insects
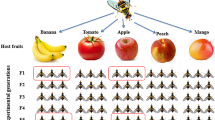
The mother culture of B. dorsalis was obtained from tropical almond fruits collected from Onekahakaha Beach Park, Hilo, HI, USA, in June–July 2022. Infested tropical almond fruits were transferred to the laboratory (25 ± 1 °C, 65% RH, and 16:8 h light and dark photoperiod) and placed on a 2-cm thick layer of sterile vermiculite (Vigoro, USA) to aid pupation. Pupae were sieved from the sterile vermiculite into a sterile Petri plate (90 mm dia.) and placed in rearing cages (30 × 30 × 30 cm, BugDorm.com) for adults to emerge. Emerged adult flies were provided with a mixture of honey and yeast hydrolysate (1:1 w/w) and water ad libitum for maturation and egg development. Mature gravid female flies (30 days old) from tropical almond fruits were provided with either guava, mango, papaya, or rose apple separately as oviposition substrates (Fig. 1). Organic fruits from local grocery stores were washed with a diluted soap solution, followed by 5% sodium hypochlorite solution for 1 min and rinsed in distilled water three times, and wiped dry using sterile tissues before providing them to wild B. dorsalis gravid female flies that emerged from tropical almond fruits (30 days old; 50 flies). After oviposition occurred (48 h), each fruit was placed into individual containers holding sterile vermiculite (2-cm thick layer) to aid pupation. The pupae that developed from the individual fruits were placed separately in rearing cages, and the newly emerged teneral female flies were used for further experiments.
Graphical representation of the experimental design. Mother flies were sourced from infested tropical almonds. A single cohort of mother flies from tropical almonds was allowed to oviposit into four different fruit hosts (larval diet; guava, mango, papaya, and rose apple). The teneral female flies that emerged from each fruit host were further used to study their gut and ovipositor microbiome
Dissection of Gut Tissues of Teneral Flies
Tissues of flies from tropical almond (mother flies; 30 days old), guava, mango, papaya, and rose apple (teneral female flies;1 day old; not fed with diet) were dissected using general aseptic dissection procedures from gut and ovipositor. Prior to dissection, female flies from each fruit were kept at 5 °C for 15 mins to slow down the movement and were individually surface sterilized with 5% sodium hypochlorite solution for 30 s followed by a wash with 70% ethanol for 1–2 min and then rinsed with sterilized distilled water for 3 min. Female flies (n = 10 per host fruit species) were dissected individually on an aseptic plate containing 2 ml sterilized phosphate-buffered saline (PBS, 137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4, pH 7.4) using sterilized forceps under a stereomicroscope. The forceps were sterilized intermediately during the dissection procedure by dipping in ethanol (70%) and flaming the tips using a spirit lamp. The whole gut tissue including the ovipositor was dissected, and later, the ovipositor along with the oviduct was separated from the whole gut tissue. Gut and ovipositor tissues from each fly were transferred separately into individual aseptic centrifuge tube containing 1 ml of DNA/RNA Shield (Zymo Research), thus making 10 replicates of tissues from flies that emerged from each selected fruit. The tubes containing sample tissues were stored at −80 °C until further use.
DNA Extraction and Microbiome Analysis
DNA was extracted using a ZymoBIOMICS DNA Extraction kit on a KingFisher Flex Magnetic Particle Processor (Thermo Scientific). Mock communities (Zymo) and empty wells were included in the extraction plates as positive and negative controls, respectively. 16S-rRNA metabarcoding was conducted as described previously [42]. The V4 sub-region of the 16S SSU rRNA was amplified using 515F (GTGCCAGCMGCCGCGGTAA) and 806R (GGACTACHVGGGTWTCTAAT). Reactions were performed in 25 µl volumes using Q5 Hot Start High-Fidelity Polymerase (New England Biolabs) with 0.2 µM of each primer. Reaction conditions were 98 ℃ for 30 s, 30 cycles of 98 ℃ for 30 s, 50 ℃ for 30 s, and 72 ℃ for 2 min, and a final extension at 72 ℃ for 10 min. PCR products were normalized using a Just-A-Plate normalization kit (Charm Biotech). Amplicon pools were sequenced at the ASGPB Genomics Core at the University of Hawai’i at Manoa using Illumina MiSeq V3 600 chemistry (Illumina, San Diego, CA, USA).
Processing of Sequence Data
Illumina 16S reads were processed with the “DADA2” (v. 1.24) pipeline to obtain amplicon sequence variants (ASVs) [43] implemented in R. Steps included filtering, dereplication, inference of sequence variants, mergers of paired-end reads, and chimera detection and removal. Taxonomy assignments were performed using “DECIPHER” (v. 2.24.0) [44] with a trained version of the Ribosomal Database Project (RDP) reference database (v 18) and ASVs assigned to chloroplast and mitochondria, or those which were unclassified at the domain level were removed from the dataset prior to statistical analyses. After initial processing and taxonomic assessment, we performed classifications of ASVs with the RDP naïve Bayesian classifier with the v18 database [45].
Statistical Analyses
Statistical analyses were performed using R 4.2.1 in RStudio [46, 47]. Microbiome data were analyzed using “vegan” [48] after processing with DADA2 [43]. Samples were rarefied to 8000 sequences for downstream analyses. ASV composition and membership were analyzed using Bray-Curtis and Jaccard distances, respectively. Distances were visualized and analyzed using non-metric multidimensional scaling (NMDS) and PERMANOVA implemented with adonis2. Since gut and ovipositor samples were from paired individuals, we used individuals as strata in the model. NMDS plots were made for the individual tissues (gut and ovipositor) to display differences in the initial fruit source. Pairwise multivariate analysis of Bray-Curtis and Jaccard distances was performed using the package “pairwiseAdonis” using a false discovery rate (FDR) correction [49]. Diversity metrics such as ASV richness, Shannon, and 1/Simpson metrics were computed in vegan and analyzed using a Kruskal-Wallis test with pairwise comparisons being performed with Dunn tests [50]. In order to identify potential core components of the oriental fruit fly microbiomes in our study, we evaluated occupancy-abundance distributions of the ASVs associated with the flies. ASVs were grouped by the individual fly specimen (gut and ovipositor averaged). When applying a stringent occupancy = 1, we observed no conserved ASVs across all individual samples. Therefore, we conducted occupancy-abundance analysis using a threshold based on contribution to Bray-Curtis similarity [51]. The 500 most prevalent ASVs were ranked according to their contribution to Bray-Curtis beta-diversity, with a cutoff of core ASVs by the last 2% increase in explanatory value by Bray-Curtis similarity [51]. Both core and ASVs were plotted against a Sloan neutral model constructed using the R package “tyRa” [51,52,53]. Heatmap of core ASVs was generated using “pheatmap” [54], with Bray-Curtis distances being used to cluster samples and visualization of ASV relative abundances with a Z-transformation of ASV relative abundances.
Results
Sequencing of Controls
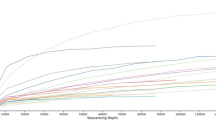
We included a negative kit control and positive mock communities through our extraction procedure and performed PCR alongside our experimental samples. The negative control yielded no sequences at the end of the pipeline. Sequencing of positive controls returned ASV numbers and taxonomical classifications to what was expected from the mock community (Fig. S1). Eight ASVs comprised >99% of the sequences in the mock community controls. Relative abundances of controls in the two extraction plates did not exhibit any marked differences from each other. There was some variation in the processed controls from the theoretic makeup, but that is unexpected given the biases inherent in PCR-based approaches.
Host Fruit Influences Microbial Community Composition
Larval diet had effects on bacterial ASV composition associated with B. dorsalis guts and ovipositors. Using Bray-Curtis distances observed, there were effects of larval diet (fruits) (PERMANOVA-F4,82 = 4.95, p < 0.001), but not the host tissue (F1,82 = 1.75, p = 0.122). We did observe interactive effects between host fruit and tissue source (F4,82 = 2.43, p = 0.002). Evaluating pairwise comparisons, most of the paired gut and ovipositor samples have similar responses with a couple of exceptions (Table S1). In all but one case (rose apple), there were no differences in gut and ovipositor memberships (p > 0.05). NMDS plots of separated gut and ovipositor samples demonstrated a triangular shape in the ordination (Fig. 2A, B) as well as some separation of samples originating from different host fruits, suggesting that the hosts yielded three different dominant ASV memberships. There were overlapping individuals from different host plants in both cases, indicating that, while the larval diet has an impact on the ASV composition, there is variation among the treatments. Jaccard distances using presence/absence data indicated additional differences in ASV memberships (Fig. 2C, D). PERMANOVA of Jaccard distances indicated that there was a significant impact of larval diet (F4,82 = 3.38, p < 0.001), adult tissue type (F1,82 = 5.91, p < 0.001), and their interaction (F4,82 = 1.84, p < 0.001) on ASV membership among samples. Compared to Bray-Curtis dissimilarities, pairwise Jaccard distances yielded far more differences between individuals, suggesting that sparsely populated aspects of the microbiome diverged between the female tissues. Additionally, Jaccard plots demonstrated a higher degree of structuring by host plant in both gut (Fig. 2C) and ovipositor (Fig. 2D) samples, indicating greater contribution of host plants to less-dominant aspects of the B. dorsalis adult female microbiome.
NMDS plots showing the impact of fruit source on female B. dorsalis 16S ASVs. Gut samples (A, C) and ovipositor samples (B, D) were analyzed separately for these analyses. NMDS plots were constructed with Bray-Curtis (top) and Jaccard (bottom) dissimilarities. Plots indicate some clustering of samples from different fruit sources, especially with gut tissues
Influence of larval diet on B. dorsalis ASV richness and diversity estimates analysis varied between tissue types (Fig. 3). For gut tissues, there was a significant effect of host plant on ASV richness (Fig. 2A; χ2 = 16.7, p = 0.002) and Shannon diversity (χ2 = 11.3, p = 0.023). Gut samples (Fig. 3A) collected from flies originating from rose apple reduced ASV richness than those from guava and mango and had less diversity than specimens collected from tropical almond. Bactrocera dorsalis ovipositor samples (Fig. 3B) did not differ in either richness (χ2 = 2.6, p = 0.627) or diversity (χ2 = 1.2, p = 0.886).
Evaluating Incidence of ASVs Across Samples
We used occupancy-abundance relationships using a Bray-Curtis similarity cutoff to evaluate potentially core ASVs across sample types (Fig. 4). A first-order cutoff (Fig. 4A; red line) yielded three potentially core ASVs, while 31 ASVs were determined to contribute to Bray-Curtis dissimilarity above a 2% cutoff (Fig. 4A; blue line). None of these core ASVs were present across all individuals sampled (Fig. 4B), with the highest incidence being ~85%. However, despite these ASVs not having universal occupancy, they were present in a specimen from each of the larval host fruits (Fig. 4C). Heat map clustering of the dominant ASVs in gut samples also demonstrated the sparseness of the B. dorsalis adult fly microbiome, with ~10 ASVs being the most dominant. As suggested by the ordination analysis, the gut samples were clustered into three main groups (left axis), generally following patterns indicated by host fruits. Samples were generally dominated by two to five ASVs, with the remaining community having relatively low abundances.
Determination and distribution of B. dorsalis ASVs as determined by occupancy-abundance analysis across all individual samples. Three core ASVs were determined using first-order differences (red line), and 36 were by contributions to Bray-Curtis distances (blue line) (A). Occupancy-abundance plot (B) displaying the 200 most prevalent ASVs across all samples, where an occupancy of 1 indicates presence across all sample types. ASVs that are shaded blue are those determined to be core components according to the Bray-Curtis cutoff method, with ASVs mapped to the Sloan neutral model. Heatmap (C) was constructed using only gut microbiome samples, with sample clustering being performed using Bray-Curtis distances between sample types and ASVs clustered using Euclidean distances. ASV relative abundances were z-score transformed before plotting
Taxonomic Consistency Across the Samples
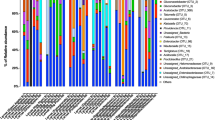
The three most prevalent ASVs in gut samples were classified as Klebsiella (ASV1 and 2), Morganella (ASV3), and Providencia (ASV6) (Fig. 4C). Because the ASVs classified as Klebsiella exhibited unique colonization patterns, it suggests that these are likely originating from separate strains that differentially associated with the samples. Evaluating the relative abundances of ASV taxa, we saw similar patterns. Overall, our results show that, on the genus level (Fig. 5A, 5B), the microbiome of flies from all fruits and tissue was dominated by Klebsiella (34–80%). Similarly, Providencia (0.15–48%) was also present in the gut and the ovipositor of flies from all selected fruits. In contrast, Morganella was present in the gut of flies from tropical almond (25%), mango (36%), and papaya (8%) but was virtually absent in guava and rose apple. However, they were present in the ovipositor of flies from all the fruits. The remaining genera, namely, unclassified Enterobacteria (0.6–23%), Enterococcus (0.8–6%), Acinetobacter (0–11%), Raoultella (0–6%), and Serratia (0–8%) were present in low abundance. It should be noted that although the major bacterial constituents in flies were similar, the overall abundance of other bacterial genera, in both gut and ovipositor of flies from guava, mango, papaya, and rose apple, differed from the microbiome of the mother culture (flies from tropical almond). It is also interesting to note that the three major taxa, Klebsiella, Providencia, and Morganella, dominated the microbiome and likely influenced other minor bacterial taxa. We could clearly see that the major bacterial taxa (Klebsiella, Providencia, and Morganella) replaced each other but did not allow other bacterial taxa to replace them although the flies emerged from different fruits (larval diet). This evidently proves that these three major taxa are crucial and may have an influence on the microbiome structure to some extent, other than the diet. However, further experiments are needed to confirm this aspect.
Relative abundance (%) of ASVs associated with B. dorsalis gut (A) and ovipositor (B). Classifications were conducted using the RDP classifier (v 2.13) using the v.18 training set. Taxonomy was determined at a 60% bootstrap threshold. Taxa with relative abundances < 2% across all specimens were grouped into the “Other” category
Discussion
Insect guts and ovipositors are complex environments where host and environmental factors can collectively influence microbial composition and function. Bactrocera dorsalis harbors a complex microbiome that affects its behavior and nutritional status [55,56,57]. In our study, we found high variability in the microbiome composition between individuals, but we found some bacterial ASVs and corresponding genera present in high frequency in all the individual flies tested. Similarly, in Drosophila, the microbiome composition can be highly variable between individuals [58], though some key bacterial species are typically present in higher frequency [29]. The goal of our study was to identify potential stable constituents of the B. dorsalis female fly microbiome.
We determined that the microbiome structure varied between individual teneral flies raised on different host fruits and between individual flies raised on the same fruit suggesting that the forces influencing the microbiome structure extend beyond diet. However, we found ASVs taxonomically classified as Klebsiella present in all teneral female flies irrespective of their origin, though at different relative abundances (Fig. 4). This supports other studies from all around the globe that have indicated Klebsiella to be an important aspect of tephritid fruit flies’ microbiome [[59] and references within]. ASVs/OTUs of Enterobacteriaceae [60,61,62,63] and particularly Klebsiella are highly dominant taxa [64,65,66] in most of the tephritid flies. Klebsiella spp. have been shown to improve larval development and can affect adult performance and behavior, aiding courtship and reproduction in tephritid flies [67,68,69,70]. They may also contribute to the fly fitness by increasing nitrogen availability through nitrogen fixation, an common trait of Klebsiella [71, 72]. Klebsiella is also known to provide carbon to the flies by pectinolysis, a trait of K. oxytoca [73], a dominant cultured bacterial species in tephritid fruit flies including B. dorsalis [3, 7, 74, 75]. This suggests that Klebsiella may have some integral functions in the life history of fruit flies and warrants further exploration.
The microbiome of the gut and ovipositor of flies raised on selected fruits was generally consistent except for the microbiome of flies raised on rose apple that was dominated by Providencia. However, Providencia was abundant in the gut (10–40% of relative abundance) in all teneral flies. Providencia is a gram-negative, non-spore forming bacteria that is an opportunistic pathogen to some insects [75] and has also been isolated from many fruit fly species including A. ludens [76, 77], A. obliqua, A. serpentina, A. striata [77], B. dorsalis [3], B. oleae [31], B. tryoni [78], Ceratitis capitata [79], Zeugodacus tau [80], Z. cucurbitae [64], and Drosophila melanogaster [81]. To date, there is no evidence of these bacteria causing infections in B. dorsalis including our unpublished data from colonies maintained by the USDA ARS (Mason unpublished) and feeding experiments (Kempraj and Cha unpublished). On the contrary, some studies have demonstrated positive effects of Providencia on the development of fruit flies and improving fly resistance to fungal infection [31]. The other bacterial taxa that were abundant in the flies’ gut and ovipositor tissues were Morganella. Salas et al. [82] observed M. morganii to cause infection in reared A. ludens larvae and was also detected in wild B. dorsalis [3], B. tryoni [78], and Z. tau [80]. Although potentially pathogenic, the reason for the abundance of these bacteria in the gut and ovipositor tissues in teneral flies is to be uncovered.
In our analysis, we observed some differences in how host fruit altered the ovipositor microbiome compared to that of the gut. Currently, we can only speculate what may drive these differences. Considering that the samples were surface sterilized, we do not think that external colonizers were the drivers of these differences. We suspect that differences in tissue structure may be the root cause. The ovipositor and junction between the gut and ovipositor are comprised of a cuticle, while the epithelial cells are lined with a mucosal-like peritrophic matrix. These substrates can favor different colonization frequencies and result in relatively distinct microbial communities for insects [83, 84]. Additionally, the gut is active and dynamic in regulating microbial populations where the ovipositor structures may not be. Further manipulative experiments are needed to understand these differences for B. dorsalis and other fruit flies.
The present study explored differences in the gut and ovipositor microbiome of teneral B. dorsalis females reared from wild origins on four different host fruits. Overall, we found that the larval diet does have an effect on the gut microbial composition of newly emerged flies. However, we also found that some bacterial genera, such as Klebsiella, Providencia, and Morganella, that were abundant in the wild mother fly were also abundant and preserved in the flies that were reared on different host fruits. This result is similar to other studies on humans and Drosophila, where different individuals could preserve and maintain specific microbes even after extensive dietary changes [85, 86]. Taken together, our results show that although there is a possibility of vertical transfer of specific bacterial constituents from the mother to progeny, larval diet (host fruit) had a profound impact on the microbiome structure overall [21]. It would be safe to consider that the genera Klebsiella, Providencia, and Morganella are the major bacterial constituents that are extremely stable in flies from all the selected fruits, but there are potentially different strains that could dominate the gut microbiome. This raises questions about how similar bacterial strains may compete and dominate in the guts of insects. Further exploration with other sequencing techniques alongside manipulative experiments is needed to determine the ecological drivers and ultimate effects. Overall, the knowledge provided here can aid us to understand B. dorsalis-microbiome interaction in an ecological context and shows that for stable interactions, the fly may have to conserve a few specific bacteria to possibly increase fitness. Moreover, these results develop the use of B. dorsalis as a model to study microbiota proliferation and colonization. Upon understanding the role of these specific bacteria in the life history of B. dorsalis, the knowledge may aid us to develop novel management strategies to control this devastating horticultural pest.
Data Availability
Data and R code has been deposited in public databases. Raw sequence reads are available at NCBI SRA (PRJNA1061202). Processed ASV sequences, tables, taxonomy, and associated R code have been deposited at USDA ARS Ag Data Commons (https://doi.org/10.15482/USDA.ADC/24969123)
References
McFall-Ngai M, Hadfield MG, Bosch TCG et al (2013) Animals in a bacterial world, a new imperative for the life sciences. P Natl Acad Sci USA 110:3229–3236. https://doi.org/10.1073/pnas.1218525110
Collins SM, Surette M, Bercik P (2012) The interplay between the intestinal microbiota and the brain. Nat Rev Microbiol 10:735–742. https://doi.org/10.1038/nrmicro2876
Sommer F, Bäckhed F (2013) The gut microbiota - masters of host development and physiology. Nat Rev Microbiol. 11:227–238. https://doi.org/10.1038/nrmicro2974
Damodaram KJP, Ayyasamy A, Kempraj V (2016) Commensal bacteria aid mate-selection in the fruit fly, Bactrocera dorsalis. Microb ecol 72:725–729. https://doi.org/10.1007/s00248-016-0819-4
Eleftherianos I, Atri J, Accetta J, Castillo JC (2013) Endosymbiotic bacteria in insects: guardians of the immune system? Front Physiol 4. https://doi.org/10.3389/fphys.2013.00046
Frago E, Mala M, Weldegergis BT et al (2017) Symbionts protect aphids from parasitic wasps by attenuating herbivore-induced plant volatiles. Nat Commun 8:1860. https://doi.org/10.1038/s41467-017-01935-0
Ayyasamy A, Kempraj V, Damodaram KJP (2021) Endosymbiotic bacteria aid to overcome temperature induced stress in the oriental fruit fly, Bactrocera dorsalis. Microb Ecol 82:783–792. https://doi.org/10.1007/s00248-021-01682-2
Mason CJ, Shikano I (2023) Hotter days, stronger immunity? Exploring the impact of rising temperatures on insect gut health and microbial relationships. Curr Opin Insect Sci. 59:101096. https://doi.org/10.1016/j.cois.2023.101096
van den Bosch TJM, Welte CU (2016) Detoxifying symbionts in agriculturally important pest insects. Microb Biotechnol 10:531–540. https://doi.org/10.1111/1751-7915.12483
Denise MD, Kaltenpoth M, Gershenzon J (2022) Demonstrating the role of symbionts in mediating detoxification in herbivores. Symbiosis 87:59–66. https://doi.org/10.1007/s13199-022-00863-y
Akman Gündüz E, Douglas AE (2009) Symbiotic bacteria enable insect to use a nutritionally inadequate diet. Proc R Soc B 276:987–991. https://doi.org/10.1098/rspb.2008.1476
Skidmore IH, Hansen AK (2017) The evolutionary development of plant-feeding insects and their nutritional endosymbionts. Insect Sci 24:910–928. https://doi.org/10.1111/1744-7917.12463
Ferrari J, Vavre F (2011) Bacterial symbionts in insects or the story of communities affecting communities. Phil Trans R Soc B 366:1389–1400. https://doi.org/10.1098/rstb.2010.0226
Renoz F, Lopes MR, Gaget K et al (2022) Compartmentalized into bacteriocytes but highly invasive: the puzzling case of the co-obligate symbiont Serratia symbiotica in the aphid Periphyllus lyropictus. Microbiol Spec 10:e00457-22. https://doi.org/10.1128/spectrum.00457-22
Douglas AE (2014) The molecular basis of bacterial–insect symbiosis. J Mol Bio 426:3830–3837. https://doi.org/10.1016/j.jmb.2014.04.005
Storelli G, Strigini M, Grenier T et al (2018) Drosophila perpetuates nutritional mutualism by promoting the fitness of its intestinal symbiont Lactobacillus plantarum. Cell metab 27:362–377. https://doi.org/10.1016/j.cmet.2017.11.011
Bennett GM, Moran NA (2015) Heritable symbiosis: the advantages and perils of an evolutionary rabbit hole. Proc Natl Acad Sci 112:10169–10176. https://doi.org/10.1073/pnas.1421388112
Bright M, Bulgheresi S (2010) A complex journey: transmission of microbial symbionts. Nat Rev Microbiol 8:218–230. https://doi.org/10.1038/nrmicro2262
Koga R, Meng XY, Tsuchida T, Fukatsu T (2012) Cellular mechanism for selective vertical transmission of an obligate insect symbiont at the bacteriocyte–embryo interface. Proc Natl Acad Sci 109:E1230–E1237. https://doi.org/10.1073/pnas.1119212109
Gilbert JA, Neufeld JD (2014) Life in a world without microbes. PLoS Biol 12:e1002020. https://doi.org/10.1371/journal.pbio.1002020
Jose PA, Yuval B, Jurkevitch E (2022) Maternal and host effects mediate the adaptative expansion and contraction of the microbiome during ontogeny in a holometabolous, polyphagous insect. Funct Ecol 37:929–946. https://doi.org/10.1111/1365-2435.14286
Ley RE, Hamady M, Lozupone C et al (2008) Evolution of mammals and their gut microbes. Science 320:1647–1651. https://doi.org/10.1126/science.1155725\
Ochman H, Worobey M, Kuo CH et al (2010) Evolutionary relationships of wild hominids recapitulated by gut microbial communities. PLoS Biol 8:e1000546. https://doi.org/10.1371/journal.pbio.1000546
Muegge BD, Kuczynski J, Knights D et al (2011) Diet drives convergence in gut microbiome functions across mammalian phylogeny and within humans. Science 332:970–974. https://doi.org/10.1126/science.1198719
Heitlinger E, Ferreira SC, Thierer D, Hofer H, East ML (2017) The intestinal eukaryotic and bacterial biome of spotted hyenas: the impact of social status and age on diversity and composition. Front Cell Infect Microbiol 7:262. https://doi.org/10.3389/fcimb.2017.00262
Gilbert JA, Blaser MJ, Caporaso JG, Jansson JK, Lynch SV, Knight R (2018) Current understanding of the human microbiome. Nat Med 24:392–400. https://doi.org/10.1038/nm.4517
Chu M, Zhang X (2022) Bacterial atlas of mouse gut microbiota. Cell Microbiol 2022:5968814. https://doi.org/10.1155/2022/5968814
Wong CNA, Ng P, Douglas AE (2011) Low-diversity bacterial community in the gut of the fruit fly Drosophila melanogaster. Enviro Microbiol 13:1889–1900. https://doi.org/10.1111/j.1462-2920.2011.02511.x
Broderick NA, Lemaitre B (2012) Gut-associated microbes of Drosophila melanogaster. Gut microbes 3:307–321. https://doi.org/10.4161/gmic.19896
Erkosar B, Storelli G, Defaye A, Leulier F (2013) Host-intestinal microbiota mutualism: “learning on the fly.” Cell host & microbe 13:8–14. https://doi.org/10.1016/j.chom.2012.12.004
Koskinioti P, Ras E, Augustinos AA, Beukeboom LW et al (2020) Manipulation of insect gut microbiota towards the improvement of Bactrocera oleae artificial rearing. Entomol Exp Appl 168:523–540. https://doi.org/10.1111/eea.12934
Chen B, Mason CJ, Peiffer M, Zhang D, Shao Y, Felton GW (2022) Enterococcal symbionts of caterpillars facilitate the utilization of a suboptimal diet. J Insect Physiol 138:104369. https://doi.org/10.1016/j.jinsphys.2022.104369
Drew RAI, Raghu S (2002) The fruit fly fauna (Diptera: Tephritidae: Dacinae) of the rainforest habitat of the Western Ghats, India. Raffles Bull Zool 50:327–352
Metcalf R, Metcalf E (1992) Fruit flies of the family Tephritidae. In: Metcalf R, Metcalf E (eds) Plant kairomones in insect ecology and control, 1st edn. Chapman and Hall, New York, pp 109–152
Verghese A, Madhura HS, Kamala Jayanthi PD, Stonehouse JM (2002) Fruit flies of economic significance in India with special reference to Bactrocera dorsalis (Hendel). In: Barnes NB (ed) Proceedings of the 6th international Fruit Flies Symposium, 1st edn. Scientific Publications, Stellenbosch, pp 317–324
Clark AR, Armstrong KF, Carmichael AE et al (2005) Invasive phytophagous pest arising through a recent tropical evolutionary radiation: The Bactrocera dorsalis Complex of Fruit Fly. Annu Rev Entomol 50:293–319. https://doi.org/10.1146/annurev.ento.50.071803.130428
Shelly TE (2000) Fecundity of female oriental fruit flies (Diptera: Tephritidae): effects of methyl eugenol-fed and multiple mates. Ann Entomol Soc Am 93:559–564. https://doi.org/10.1603/0013-8746(2000)093[0559:FOFOFF]2.0.CO;2
Stephens AEA, Kriticos DJ, Leriche A (2007) The current and future potential geographical distribution of the oriental fruit fly, Bactrocera dorsalis (Diptera: Tephritidae). Bull Entomol Res 97:369–378. https://doi.org/10.1017/S0007485307005044
Yuan M, Wang B, Song CB, Rong XL, Yin Y (2008) Effect of climate factors and host plants on population dynamics of Bactrocera dorsalis (Hendel) in Suzhou. J Anhui Agric Sci 22:9619–9621
Jose PA, Ben-Yosef M, Jurkevitch E, Yuval B (2019) Symbiotic bacteria affect oviposition behavior in the olive fruit fly Bactrocera oleae. J Insect Physiol 117:103917. https://doi.org/10.1016/j.jinsphys.2019.103917
Zaada DSY, Ben-Yosef M, Yuval B, Jurkevitch E (2019) The host fruit amplifies mutualistic interaction between Ceratitis capitata larvae and associated bacteria. BMC Biotechnol 19:92. https://doi.org/10.1186/s12896-019-0581-z
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Callahan BJ, McMurdie PJ, Rosen MJ (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13:581–583. https://doi.org/10.1038/nmeth.3869
Wright ES (2016) Using DECIPHER v2. 0 to analyze big biological sequence data in R. R Journal 8:352–359
Wang Q, Garrity GM, Tiedje JM, Cole CR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Posit Team (2023) RStudio: Integrated development environment for R. Posit Software, PBC, Boston, MA. http://www.posit.co/.
R Core Team (2023) R: A language and environment for statistical computing. Vienna, Austria. https://www.R-project.org/
Oksanen J, Simpson G, Blanchet F, et al. (2022). vegan: Community ecology package. R package version 2.6-4. https://CRAN.R-project.org/package=vegan
Martinez Arbizu P (2020) PairwiseAdonis: pairwise multilevel comparison using adonis. R package version 0.4.
Ogle D, Ogle MD (2017) Package ‘FSA’. CRAN Repos:1–206
Shade A, Stopnisek N (2019) Abundance-occupancy distributions to prioritize plant core microbiome membership. Curr Opin Microbiol 49:50–58. https://doi.org/10.1016/j.mib.2019.09.008
Sloan WT, Lunn M, Woodcock S, Head IM, Nee S, Curtis TP (2006) Quantifying the roles of immigration and chance in shaping prokaryote community structure. Environ Microbiol 8:732–740. https://doi.org/10.1111/j.1462-2920.2005.00956.x
Sprockett DD, Martin M, Costello EK, Burns AR, Holmes SP, Gurven MD, Relman DA (2020) Microbiota assembly, structure, and dynamics among Tsimane horticulturalists of the Bolivian Amazon. Nat Commun 11:3772. https://doi.org/10.1038/s41467-020-17541-6
Kolde R (2012) Pheatmap: pretty heatmaps. R package version 1.
Cheng D, Guo Z, Riegler M, Xi Z, Liang G, Xu Y (2017) Gut symbiont enhances insecticide resistance in a significant pest, the Oriental fruit fly Bactrocera dorsalis (Hendel). Microbiome 5:13. https://doi.org/10.1186/s40168-017-0236-z
Akami M, Andongma AA, Zhengzhong C et al (2019) Intestinal bacteria modulate the foraging behavior of the oriental fruit fly Bactrocera dorsalis (Diptera: Tephritidae). PLoS One 14:e0210109. https://doi.org/10.1371/journal.pone.0210109
Khan M, Seheli K, Bari MA, Sultana N, Khan SA, Sultana KF, Hossain MA (2019) Potential of a fly gut microbiota incorporated gel-based larval diet for rearing Bactrocera dorsalis (Hendel). BMC Biotechnol 19:94. https://doi.org/10.1186/s12896-019-0580-0
Ludington WB, Ja WW (2020) Drosophila as a model for the gut microbiome. PLoS Pathogens 16:e1008398. https://doi.org/10.1371/journal.ppat.1008398
Raza MF, Yao Z, Bai S, Cai Z, Zhang H (2020) Tephritidae fruit fly gut microbiome diversity, function and potential for applications. Bull Entomol Res 110:423–437. https://doi.org/10.1017/S0007485319000853
Morrow JL, Frommer M, Shearman DC, Riegler M (2015) The microbiome of field-caught and laboratory-adapted Australian tephritid fruit fly species with different host plant use and specialisation. Microb Ecol 70:498–508. https://doi.org/10.1007/s00248-015-0571-1
Zhao X, Zhang X, Chen Z, Wang Z, Lu Y, Cheng D (2018) The divergence in bacterial components associated with Bactrocera dorsalis across developmental stages. Front Microbiol 9:114. https://doi.org/10.3389/fmicb.2018.00114
Yong HS, Song SL, Eamsobhana P, Pasartvit A, Lim PE (2019) Differential abundance and core members of the bacterial community associated with wild male Zeugodacus cucurbitae fruit flies (Insecta: Tephritidae) from three geographical regions of Southeast Asia. Mol Biol Rep 46:3765–3776. https://doi.org/10.1007/s11033-019-04818-3
Naaz N, Choudhary JS, Choudhary A, Dutta A, Das B (2020) Developmental stage-associated microbiota profile of the peach fruit fly, Bactrocera zonata (Diptera: Tephritidae) and their functional prediction using 16S rRNA gene metabarcoding sequencing. 3 Biotech 10:390. https://doi.org/10.1007/s13205-020-02381-4
Choudhary JS, Naaz N, Prabhakar CS, Das B, Singh AK, Bhatt BP (2021) High taxonomic and functional diversity of bacterial communities associated with melon fly, Zeugodacus cucurbitae (Diptera: Tephritidae). Curr Microbiol 78:611–623. https://doi.org/10.1007/s00284-020-02327-2
Darrington M, Leftwich PT, Holmes NA et al (2021) Characterisation of the symbionts in the Mediterranean fruitfly gut. Microb Genom 8:000801. https://doi.org/10.1099/mgen.0.000801
Wang AL, Yao ZC, Zheng WW, Zhang HY (2014) Bacterial communities in the gut and reproductive organs of Bactrocera minax (Diptera: Tephritidae) based on 454 pyrosequencing. PLoS One 9:e106988. https://doi.org/10.1371/journal.pone.0106988
Ben Ami E, Yuval B, Jurkevitch E (2010) Manipulation of the microbiota of mass-reared Mediterranean fruit flies Ceratitis capitata (Diptera: Tephritidae) improves sterile male sexual performance. ISME J 4:28–37. https://doi.org/10.1038/ismej.2009.82
Gavriel S, Jurkevitch E, Gazit Y, Yuval B (2011) Bacterially enriched diet improves sexual performance of sterile male Mediterranean fruit flies. J Appl Entomol 135:564–573. https://doi.org/10.1111/j.1439-0418.2010.01605.x
Kyritsis GA, Augustinos AA, Caceres C, Bourtzis K (2017) Medfly gut microbiota and enhancement of the sterile insect technique: similarities and differences of Klebsiella oxytoca and Enterobacter sp. AA26 probiotics during the larval and adult stages of the VIENNA 8(D53+) genetic sexing strain. Front Microbiol 8:2064. https://doi.org/10.3389/fmicb.2017.02064
Behar A, Yuval B, Jurkevitch E (2008) Gut bacterial communities in the Mediterranean fruit fly (Ceratitis capitata) and their impact on host longevity. J Insect Physiol 54:1377–1383. https://doi.org/10.1016/j.jinsphys.2008.07.011
Ben-Yosef M, Pasternak Z, Jurkevitch E, Yuval B (2014) Symbiotic bacteria enable olive flies (Bactrocera oleae) to exploit intractable sources of nitrogen. J Evol Biol 27:2695–2705. https://doi.org/10.1111/jeb.12527
Blin C, Passet V, Touchon M, Rocha EPC, Brisse S (2017) Metabolic diversity of the emerging pathogenic lineages of Klebsiella pneumoniae. Environ Microbiol 19:1881–1898. https://doi.org/10.1111/1462-2920.13689
Sakazaki R, Tamura K, Kosako Y, Yoshizaki E (1989) Klebsiella ornithinolytica sp. nov formerly known as ornithine-positive Klebsiella oxytoca. Curr Microbiol 18:201–206. https://doi.org/10.1007/BF01570291
Behar A, Ben-Yosef M, Lauzon CR, Yuval B, Jurkevich E (2008) Structure and function of the bacterial community associated with the Mediterranean fruit fly. In: Bourtzis K, Miller TA (eds) Insect Symbiosis, 1st edn. CRC Press, Boca Raton, pp 251–272. https://doi.org/10.1201/9781420064117
O’Hara CM, Brenner FW, Miller JM (2000) Classification, identification, and clinical significance of Proteus, Providencia, and Morganella. Clin Microbiol Rev 13:534–546. https://doi.org/10.1128/cmr.13.4.534
Kuzina LV, Peloquin JJ, Vacek DC, Miller TA (2001) Isolation and identification of bacteria associated with adult laboratory Mexican fruit flies, Anastrepha ludens (Diptera: Tephritidae). Curr Microbiol 42:290–294. https://doi.org/10.1007/s002840110219
Ventura C, Briones-Roblero CI, Hernández E, Rivera-Orduña FN, Zúñiga G (2018) Comparative analysis of the gut bacterial community of four Anastrepha fruit flies (Diptera: Tephritidae) based on pyrosequencing. Curr Microbiol 75:966–976. https://doi.org/10.1007/s00284-018-1473-5
Majumder R, Sutcliffe B, Adnan SM et al (2020) Artificial larval diet mediates the microbiome of Queensland fruit fly. Front Microbiol 11:576156. https://doi.org/10.3389/fmicb.2020.576156
Guerfali MM, Djobbi W, Charaabi K et al (2018) Evaluation of Providencia rettgeri pathogenicity against laboratory Mediterranean fruit fly strain (Ceratitis capitata). PLoS One 13:e0196343. https://doi.org/10.1371/journal.pone.0196343
Noman MS, Shi G, Liu LJ, Li ZH (2021) Diversity of bacteria in different life stages and their impact on the development and reproduction of Zeugodacus tau (Diptera: Tephritidae). Insect Sci 28:363–376. https://doi.org/10.1111/1744-7917.12768
Galac MR, Lazzaro BP (2011) Comparative pathology of bacteria in the genus Providencia to a natural host, Drosophila melanogaster. Microbes Infect 13:673–683. https://doi.org/10.1186/1471-2164-13-612
Salas B, Conway HE, Schuenzel EL, Hopperstad K, Vitek C, Vacek DC (2017) Morganella morganii (Enterobacteriales: Enterobacteriaceae) is a lethal pathogen of Mexican fruit fly (Diptera: Tephritidae) larvae. Fla Entomol 100:743–751. https://doi.org/10.1653/024.100.0422
Andert J, Marten A, Brandl R, Brune A (2010) Inter- and intraspecific comparison of the bacterial assemblages in the hindgut of humivorous scarab beetle larvae (Pachnoda spp.). FEMS Microbiol Ecol 74:439–449. https://doi.org/10.1111/j.1574-6841.2010.00950.x
Zhang F, Sun XX, Zhang XC et al (2018) The interactions between gut microbiota and entomopathogenic fungi: a potential approach for biological control of Blattella germanica (L.). Pest Manag Sci 74:438–447
Pais IS, Valente RS, Sporniak M, Teixeira L (2018) Drosophila melanogaster establishes a species-specific mutualistic interaction with stable gut-colonizing bacteria. PLoS Biol 16:e2005710. https://doi.org/10.1371/journal.pbio.2005710
Leeming ER, Johnson AJ, Spector TD, Le Roy CI (2019) Effect of diet on the gut microbiota: rethinking intervention duration. Nutrients 11:2862. https://doi.org/10.3390/nu11122862
Acknowledgements
We thank Impana Lingesh and Tae-Hyung Kwon for assistance with collecting infested tropical almond fruits and Joanna Bloese and Cindy McCarty for providing laboratory space to rear out flies. We appreciate the comments.
Funding
This research was supported in part by the U.S. Department of Agriculture, Agricultural Research Service project numbers 2040-43000-018-000-D and 2040-22430-028-000-D. This research was supported in part by funding from the USDA-APHIS under the Interagency Agreement to DHC (#60-2040-3-003), Plant Protection Act 7721 funding to DHC (23-8130-1035-IA), and USDA-ARS Innovation Fund to DHC (#292-0142-021,002).
Author information
Authors and Affiliations
Contributions
VK and DHC conceived the project and design. VK, JA, and CM collected and processed flies and tissue samples. CM and VK collected and analyzed the data. VK and CM wrote the first draft of the manuscript. DHC secured funding and established the experimental framework. All authors contributed to the final version of the manuscript.
Corresponding authors
Ethics declarations
Disclosure
The findings and conclusions in this publication are those of the authors and should not be construed to represent any official USDA or U.S. government determination or policy. Mention of trade names or commercial products in this publication is solely for the purpose of providing specific information and does not imply recommendation or endorsement by the U.S. Department of Agriculture. USDA is an equal opportunity provider and employer.
Competing Interests
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kempraj, V., Auth, J., Cha, D.H. et al. Impact of Larval Food Source on the Stability of the Bactrocera dorsalis Microbiome. Microb Ecol 87, 46 (2024). https://doi.org/10.1007/s00248-024-02352-9
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00248-024-02352-9