Abstract
One of the most prominent manifestations of climate change is the changing Arctic sea-ice regime with a reduction in the summer sea-ice extent and a shift from thicker, perennial multiyear ice towards thinner, first-year ice. These changes in the physical environment are likely to impact microbial communities, a key component of Arctic marine food webs and biogeochemical cycles. During the Norwegian young sea ICE expedition (N-ICE2015) north of Svalbard, seawater samples were collected at the surface (5 m), subsurface (20 or 50 m), and mesopelagic (250 m) depths on 9 March, 27 April, and 16 June 2015. In addition, several physical and biogeochemical data were recorded to contextualize the collected microbial communities. Through the massively parallel sequencing of the small subunit ribosomal RNA amplicon and metagenomic data, this work allows studying the Arctic’s microbial community structure during the late winter to early summer transition. Results showed that, at compositional level, Alpha- (30.7%) and Gammaproteobacteria (28.6%) are the most frequent taxa across the prokaryotic N-ICE2015 collection, and also the most phylogenetically diverse. Winter to early summer trends were quite evident since there was a high relative abundance of thaumarchaeotes in the under-ice water column in late winter while this group was nearly absent during early summer. Moreover, the emergence of Flavobacteria and the SAR92 clade in early summer might be associated with the degradation of a spring bloom of Phaeocystis. High relative abundance of hydrocarbonoclastic bacteria, particularly Alcanivorax (54.3%) and Marinobacter (6.3%), was also found. Richness showed different patterns along the depth gradient for prokaryotic (highest at mesopelagic depth) and protistan communities (higher at subsurface depths). The microbial N-ICE2015 collection analyzed in the present study provides comprehensive new knowledge about the pelagic microbiota below drifting Arctic sea-ice. The higher microbial diversity found in late winter/early spring communities reinforces the need to continue with further studies to properly characterize the winter microbial communities under the pack-ice.






Similar content being viewed by others
Abbreviations
- N-ICE2015 :
-
Norwegian young sea ICE expedition 2015
- SSU rRNA :
-
small subunit ribosomal RNA
- FYI :
-
first-year ice
- MYI :
-
multiyear ice
- DOM :
-
dissolved organic matter
- CTD :
-
conductivity, temperature, and depth
- PAR :
-
photosynthetically active radiation
- SOP :
-
Standard Operating Procedure
- OTU :
-
operational taxonomic unit
- ML :
-
maximum likelihood
- PR 2 :
-
Protist Ribosomal Reference database
- NB :
-
Nansen Basin
- TR :
-
Transition Region
- YP :
-
Yermak Plateau
- Chl a :
-
chlorophyll alpha
- PSW :
-
polar surface water
- AW :
-
Atlantic water
- MAW :
-
modified Atlantic water
- DOC :
-
dissolved organic carbon
References
Lindsay R, Schweiger A (2015) Arctic sea ice thickness loss determined using subsurface, aircraft, and satellite observations. Cryosphere 9:269–283
Meier WN, Hovelsrud GK, van Oort BEH, Key JR, Kovacs KM, Michel C, Haas C, Granskog MA, Gerland S, Perovich DK, Makshtas A, Reist JD (2014) Arctic sea ice in transformation: a review of recent observed changes and impacts on biology and human activity. Rev Geophys 52:185–217
Maslanik J, Stroeve J, Fowler C, Emery W (2011) Distribution and trends in Arctic sea ice age through spring 2011. Geophys Res Lett 38:L13502. https://doi.org/10.1029/2011GL047735
Parkinson CL, Comiso JC (2013) On the 2012 record low Arctic sea ice cover: combined impact of preconditioning and an August storm. Geophys Res Lett 40:1356–1361
Polyakov IV, Walsh JE, Kwok R (2012) Recent changes of Arctic multiyear sea ice coverage and the likely causes. Bull Am Meteorol Soc 93:145–151
Assmy P, Fernández-Méndez M, Duarte P, Meyer A, Randelhoff A, Mundy CJ, Olsen LM, Kauko HM, Bailey A, Chierici M, Cohen L, Doulgeris AP, Ehn JK, Fransson A, Gerland S, Hop H, Hudson SR, Hughes N, Itkin P, Johnsen G, King JA, Koch BP, Koenig Z, Kwasniewski S, Laney SR, Nicolaus M, Pavlov AK, Polashenski CM, Provost C, Rösel A, Sandbu M, Spreen G, Smedsrud LH, Sundfjord A, Taskjelle T, Tatarek A, Wiktor J, Wagner PM, Wold A, Steen H, Granskog MA (2017) Leads in Arctic pack ice enable early phytoplankton blooms below snow-covered sea ice. Sci Rep 7:40850. https://doi.org/10.1038/srep40850
Arrigo KR, Perovich DK, Pickart RS, Brown ZW, van Dijken GL, Lowry KE, Mills MM, Palmer MA, Balch WM, Bahr F, Bates NR, Benitez-Nelson C, Bowler B, Brownlee E, Ehn JK, Frey KE, Garley R, Laney SR, Lubelczyk L, Mathis J, Matsuoka A, Mitchell BG, Moore GWK, Ortega-Retuerta E, Pal S, Polashenski CM, Reynolds RA, Schieber B, Sosik HM, Stephens M, Swift JH (2012) Massive phytoplankton blooms under arctic sea ice. Science 336:1408
Comeau AM, Li WKW, Tremblay JÉ, Carmack EC, Lovejoy C (2011) Arctic Ocean microbial community structure before and after the 2007 record sea ice minimum. PLoS One 6(11):e27492. https://doi.org/10.1371/journal.pone.0027492
Harrison WG, Cota GF (1991) Primary production in polar waters: relation to nutrient availability. Polar Res 10:87–104
Rey F (2012) Declining silicate concentrations in the Norwegian and Barents Seas. ICES J Mar Sci 69:208–212
Bowman JS, Rasmussen S, Blom N, Deming JW, Rysgaard S, Sicheritz-Ponten T (2012) Microbial community structure of Arctic multiyear sea ice and surface seawater by 454 sequencing of the 16S RNA gene. ISME J 6:11–20
Hatam I, Charchuk R, Lange B, Beckers J, Haas C, Lanoil B (2014) Distinct bacterial assemblages reside at different depths in Arctic multiyear sea ice. FEMS Microbiol Ecol 90:115–125
Hatam I, Lange B, Beckers J, Haas C, Lanoil B (2016) Bacterial communities from Arctic seasonal sea ice are more compositionally variable than those from multi-year sea ice. ISME J 10:2543–2552
Brown MV, Bowman JP (2001) A molecular phylogenetic survey of sea-ice microbial communities (SIMCO). FEMS Microbiol Ecol 35:267–275
Wilson B, Müller O, Nordmann EL, Seuthe L, Bratbak G, Øvreås L (2017) Changes in marine prokaryote composition with season and depth over an Arctic polar year. Front Mar Sci 4:95. https://doi.org/10.3389/fmars.2017.00095
Müller O, Wilson B, Paulsen ML, Rumińska A, Armo HR, Bratbak G, Øvreås L (2018) Spatiotemporal dynamics of ammonia-oxidizing Thaumarchaeota in distinct Arctic water masses. Front Microbiol 9:24. https://doi.org/10.3389/fmicb.2018.00024
Metfies K, Von Appen WJ, Kilias E, Nicolaus A, Nöthig EM (2016) Biogeography and photosynthetic biomass of arctic marine pico-eukaroytes during summer of the record sea ice minimum 2012. PLoS One 11:e0148512. https://doi.org/10.1371/journal.pone.0148512
Meshram AR, Vader A, Kristiansen S, Gabrielsen TM (2017) Microbial eukaryotes in an arctic under-ice spring bloom north of Svalbard. Front Microbiol 8:1099. https://doi.org/10.3389/fmicb.2017.01099
Li WKW, McLaughlin FA, Lovejoy C, Carmack EC (2009) Smallest algae thrive as the Arctic Ocean freshens. Science 326:539
Lasternas S, Agustí S (2010) Phytoplankton community structure during the record Arctic ice-melting of summer 2007. Polar Biol 33:1709–1717
Lalande C, Bauerfeind E, Nöthig EM, Beszczynska-Möller A (2013) Impact of a warm anomaly on export fluxes of biogenic matter in the eastern Fram Strait. Prog Oceanogr 109:70–77
Nöthig EM, Bracher A, Engel A, Metfies K, Niehoff B, Peeken I, Bauerfeind E, Cherkasheva A, Gäbler-Schwarz S, Hardge K, Kilias E, Kraft A, Mebrahtom Kidane Y, Lalande C, Piontek J, Thomisch K, Wurst M (2015) Summertime plankton ecology in fram strait - a compilation of long-and short-term observations. Polar Res 34:23349. https://doi.org/10.3402/polar.v34.23349
Bowman JS (2015) The relationship between sea ice bacterial community structure and biogeochemistry: a synthesis of current knowledge and known unknowns. Elem Sci Anthr 3:000072. https://doi.org/10.12952/journal.elementa.000072
Fernández-Méndez M, Turk-Kubo KA, Buttigieg PL, Rapp JZ, Krumpen T, Zehr JP, Boetius A (2016) Diazotroph diversity in the sea ice, melt ponds, and surface waters of the Eurasian basin of the Central Arctic Ocean. Front Microbiol 7:1884. https://doi.org/10.3389/fmicb.2016.01884
Pedrós-Alió C, Potvin M, Lovejoy C (2015) Diversity of planktonic microorganisms in the Arctic Ocean. Prog Oceanogr 139:233–243
Granskog MA et al (2016) Arctic research on thin ice: consequences of Arctic Sea Ice Loss. EOS Trans AGU 97(5):22–26
Peterson AK et al N-ICE2015 (2016) Ocean turbulent fluxes from under-ice turbulence cluster (TIC) [v1.0]. https://doi.org/10.21334/npolar.2016.ab29f1e2
Assmy P et al (2016) N-ICE2015 water column biogeochemistry [v1.0]. https://doi.org/10.21334/npolar.2016.3ebb7f64
Meyer A, Sundfjord A, Fer I, Provost C, Villacieros Robineau N, Koenig Z, Onarheim IH, Smedsrud LH, Duarte P, Dodd PA, Graham RM, Schmidtko S, Kauko HM (2017) Winter to summer oceanographic observations in the Arctic Ocean north of Svalbard. J Geophys Res Ocean 122:6218–6237
Bauer RK (2017) Oceanmap: a plotting toolbox for 2D oceanographic data
Polyakov IV, Pnyushkov AV, Alkire MB, Ashik IM, Baumann TM, Carmack EC, Goszczko I, Guthrie J, Ivanov VV, Kanzow T, Krishfield R, Kwok R, Sundfjord A, Morison J, Rember R, Yulin A (2017) Greater role for Atlantic inflows on sea-ice loss in the Eurasian Basin of the Arctic Ocean. Science 356:285–291
Meyer et al (2016) N-ICE2015 ocean microstructure profiles (MSS90L). [Data set]. Norwegian Polar Institute. https://doi.org/10.21334/npolar.2016.774bf6ab
Kopf A, Bicak M, Kottmann R, Schnetzer J, Kostadinov I, Lehmann K, Fernandez-Guerra A, Jeanthon C, Rahav E, Ullrich M, Wichels A, Gerdts G, Polymenakou P, Kotoulas G, Siam R, Abdallah RZ, Sonnenschein EC, Cariou T, O’Gara F, Jackson S, Orlic S, Steinke M, Busch J, Duarte B, Caçador I, Canning-Clode J, Bobrova O, Marteinsson V, Reynisson E, Loureiro CM, Luna GM, Quero GM, Löscher CR, Kremp A, DeLorenzo ME, Øvreås L, Tolman J, LaRoche J, Penna A, Frischer M, Davis T, Katherine B, Meyer CP, Ramos S, Magalhães C, Jude-Lemeilleur F, Aguirre-Macedo ML, Wang S, Poulton N, Jones S, Collin R, Fuhrman JA, Conan P, Alonso C, Stambler N, Goodwin K, Yakimov MM, Baltar F, Bodrossy L, van de Kamp J, Frampton DMF, Ostrowski M, van Ruth P, Malthouse P, Claus S, Deneudt K, Mortelmans J, Pitois S, Wallom D, Salter I, Costa R, Schroeder DC, Kandil MM, Amaral V, Biancalana F, Santana R, Pedrotti ML, Yoshida T, Ogata H, Ingleton T, Munnik K, Rodriguez-Ezpeleta N, Berteaux-Lecellier V, Wecker P, Cancio I, Vaulot D, Bienhold C, Ghazal H, Chaouni B, Essayeh S, Ettamimi S, Zaid EH, Boukhatem N, Bouali A, Chahboune R, Barrijal S, Timinouni M, el Otmani F, Bennani M, Mea M, Todorova N, Karamfilov V, ten Hoopen P, Cochrane G, L’Haridon S, Bizsel KC, Vezzi A, Lauro FM, Martin P, Jensen RM, Hinks J, Gebbels S, Rosselli R, de Pascale F, Schiavon R, dos Santos A, Villar E, Pesant S, Cataletto B, Malfatti F, Edirisinghe R, Silveira JAH, Barbier M, Turk V, Tinta T, Fuller WJ, Salihoglu I, Serakinci N, Ergoren MC, Bresnan E, Iriberri J, Nyhus PAF, Bente E, Karlsen HE, Golyshin PN, Gasol JM, Moncheva S, Dzhembekova N, Johnson Z, Sinigalliano CD, Gidley ML, Zingone A, Danovaro R, Tsiamis G, Clark MS, Costa AC, el Bour M, Martins AM, Collins RE, Ducluzeau AL, Martinez J, Costello MJ, Amaral-Zettler LA, Gilbert JA, Davies N, Field D, Glöckner FO (2015) The ocean sampling day consortium. Gigascience 4:27. https://doi.org/10.1186/s13742-015-0066-5
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci 108:4516–4522
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Huntley J, Fierer N, Owens SM, Betley J, Fraser L, Bauer M, Gormley N, Gilbert JA, Smith G, Knight R (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Apprill A, Mcnally S, Parsons R, Weber L (2015) Minor revision to V4 region SSU rRNA 806R gene primer greatly increases detection of SAR11 bacterioplankton. Aquat Microb Ecol 75:129–137
Parada AE, Needham DM, Fuhrman JA (2016) Every base matters: assessing small subunit rRNA primers for marine microbiomes with mock communities, time series and global field samples. Environ Microbiol 18:1403–1414
Stoeck T et al (2010) Multiple marker parallel tag environmental DNA sequencing reveals a highly complex eukaryotic community in marine anoxic water. Mol Ecol 19:21–31
Piredda R et al (2017) Diversity and temporal patterns of planktonic protist assemblages at a Mediterranean Long Term Ecological Research site. FEMS Microbiol Ecol 93:1–14
Ribeiro H, de Sousa T, Santos JP, Sousa AGG, Teixeira C, Monteiro MR, Salgado P, Mucha AP, Almeida CMR, Torgo L, Magalhães C (2018) Potential of dissimilatory nitrate reduction pathways in polycyclic aromatic hydrocarbon degradation. Chemosphere 199:54–67
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the miseq illumina sequencing platform. Appl Environ Microbiol 79:5112–5120
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541
Quast C et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41:590–596
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Westcott SL, Schloss PD (2017) OptiClust, an improved method for assigning amplicon-based sequence data to operational taxonomic units. mSphere 2:e00073–e00017. https://doi.org/10.1128/mSphereDirect.00073-17
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high- throughput community sequencing data. Nat Methods 7:335–336
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
Shannon CE (1948) A mathematical theory of communication. Bell Syst Tech J 27:379–423
Faith DP (1992) Conservation evaluation and phylogentic diversity. Biol Conserv 61:1–10
Faith DP, Baker AM (2006) Phylogenetic diversity (PD) and biodiversity conservation: some bioinformatics challenges. Evol Bioinforma 2:121–128
Pielou EC (1966) The measurement of diversity in different types of biological collections. J Theor Biol 13:131–144
Lozupone C, Knight R (2005) UniFrac : a new phylogenetic method for comparing microbial communities. Appl Environ Microbiol 71:8228–8235
Lozupone CA, Hamady M, Kelley ST, Knight R (2007) Quantitative and qualitative β diversity measures lead to different insights into factors that structure microbial communities. Appl Environ Microbiol 73:1576–1585
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8(4):e61217. https://doi.org/10.1371/journal.pone.0061217
Harrell JFE (2018) Hmisc: Harrell Miscellaneous. https://CRAN.R-project.org/package=Hmisc. Accessed March 2017
Wei T & Simko V (2017) R package ‘corrplot’: visualization of a correlation matrix (version 0.84). https://github.com/taiyun/corrplot. Accessed March 2017
R Core Team (2018) R: a language and environment for statistical computing. R Found Stat Comput, Vienna, Austria
Wickham H. ggplot2: elegant graphics for data analysis. (Springer-Verlag New York, 2009)
Kurtz ZD, Müller CL, Miraldi ER, Littman DR, Blaser MJ, Bonneau RA (2015) Sparse and compositionally robust inference of microbial ecological networks. PLoS Comput Biol 11:1–25. https://doi.org/10.1371/journal.pcbi.1004226
Guillou L et al (2013) The Protist Ribosomal Reference database (PR2): a catalog of unicellular eukaryote Small Sub-Unit rRNA sequences with curated taxonomy. Nucleic Acids Res 41:597–604
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. Peer J 4:e2584. https://doi.org/10.7717/peerj.2584
Mitchell A, Bucchini F, Cochrane G, Denise H, Hoopen P, Fraser M, Pesseat S, Potter S, Scheremetjew M, Sterk P, Finn RD (2016) EBI metagenomics in 2016 - an expanding and evolving resource for the analysis and archiving of metagenomic data. Nucleic Acids Res 44:D595–D603. https://doi.org/10.1093/nar/gkv1195
DeSantis TZ et al (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410
Chen IMA et al (2017) IMG/M: integrated genome and metagenome comparative data analysis system. Nucleic Acids Res 45:D507–D516. https://doi.org/10.1093/nar/gkr975
Becker RA, Wilks AR, Brownrigg R, Minka TP & Deckmyn A (2017) Maps: draw geographical maps
Fuhrman JA, Steele JA, Hewson I, Schwalbach MS, Brown MV, Green JL, Brown JH (2008) A latitudinal diversity gradient in planktonic marine bacteria. Proc Natl Acad Sci 105:7774–7778
Ghiglione J-F, Galand PE, Pommier T, Pedros-Alio C, Maas EW, Bakker K, Bertilson S, Kirchman DL, Lovejoy C, Yager PL, Murray AE (2012) Pole-to-pole biogeography of surface and deep marine bacterial communities. Proc Natl Acad Sci 109:17633–17638
Countway PD, Gast RJ, Dennett MR, Savai P, Rose JM, Caron DA (2007) Distinct protistan assemblages characterize the euphotic zone and deep sea (2500 m) of the western North Atlantic (Sargasso Sea and Gulf Stream). Environ Microbiol 9:1219–1232
Xu D, Jiao N, Ren R, Warren A (2017) Distribution and diversity of micriobial eukaryotes in bathypelagic waters of the south China Sea. J Eukaryot Microbiol 64:370–382
Xu D, Li R, Hu C, Sun P, Jiao N, Warren A (2017) Microbial eukaryote diversity and activity in the water column of the South China sea based on DNA and RNA high throughput sequencing. Front Microbiol 8:1121. https://doi.org/10.3389/fmicb.2017.01121
Galand PE, Potvin M, Casamayor EO, Lovejoy C (2010) Hydrography shapes bacterial biogeography of the deep Arctic Ocean. ISME J 4:564–576
Morris RM, Rappé MS, Connon SA, Vergin KL, Siebold WA, Carlson CA, Giovannoni SJ (2002) SAR11 clade dominates ocean surface bacterioplankton communities. Nature 420:806–810
Kirchman DL, Cottrell MT, Lovejoy C (2010) The structure of bacterial communities in the western Arctic Ocean as revealed by pyrosequencing of 16S rRNA genes. Environ Microbiol 12:1132–1143
Giovannoni SJ et al (2005) Genome streamlining in a cosmopolitan oceanic bacterium. Science 309:1242–1245
Steindler L, Schwalbach MS, Smith DP, Chan F, Giovannoni SJ (2011) Energy starved candidatus pelagibacter ubique substitutes light-mediated ATP production for endogenous carbon respiration. PLoS One 6:1–10. https://doi.org/10.1371/journal.pone.0019725
Tripp HJ, Kitner JB, Schwalbach MS, Dacey JWH, Wilhelm LJ, Giovannoni SJ (2008) SAR11 marine bacteria require exogenous reduced sulphur for growth. Nature 452:741–744
Malmstrom RR, Kiene RP, Cottrell MT, Kirchman DL (2004) Contribution of SAR11 bacteria to dissolved dimethylsulfoniopropionate and amino acid uptake in the North Atlantic Ocean. Appl Environ Microbiol 70:4129–4135
Liss P, Malin G, Turner S, Holligan P (1994) Dimethyl sulphide and Phaeocystis: a review. J Mar Syst 5:41–53. https://doi.org/10.1016/0924-7963(94)90015-9
Matrai PA, Vernet M (1997) Dynamics of the vernal bloom in the marginal ice zone of the Barents Sea: dimethyl sulfide and dimethylsulfoniopropionate budgets. J Geophys Res 102:22965–22979
Levasseur M (2013) Impact of Arctic meltdown on the microbial cycling of sulphur. Nat Geosci 6:691–700
Dacey JWD, Wakeham SG (1986) Oceanic dimethylsulfide: production during zooplankton grazing on phytoplankton. Science 233:1314–1316
Stingl U, Desiderio RA, Cho JC, Vergin KL, Giovannoni SJ (2007) The SAR92 clade: an abundant coastal clade of culturable marine bacteria possessing proteorhodopsin. Appl Environ Microbiol 73:2290–2296
Klindworth A, Mann AJ, Huang S, Wichels A, Quast C, Waldmann J, Teeling H, Glöckner FO (2014) Diversity and activity of marine bacterioplankton during a diatom bloom in the North Sea assessed by total RNA and pyrotag sequencing. Mar Genomics 18:185–192
Teeling H et al (2016) Recurring patterns in bacterioplankton dynamics during coastal spring algae blooms. Elife 5:1–31. https://doi.org/10.7554/eLife.11888.001
Wemheuer B, Güllert S, Billerbeck S, Giebel HA, Voget S, Simon M, Daniel R (2014) Impact of a phytoplankton bloom on the diversity of the active bacterial community in the southern North Sea as revealed by metatranscriptomic approaches. FEMS Microbiol Ecol 87:378–389
Eronen-Rasimus E, Piiparinen J, Karkman A, Lyra C, Gerland S, Kaartokallio H (2016) Bacterial communities in Arctic first-year drift ice during the winter/spring transition. Environ Microbiol Rep 8:527–535
Han D, Kang I, Ha HK, Kim HC, Kim OS, Lee BY, Cho JC, Hur HG, Lee YK (2014) Bacterial communities of surface mixed layer in the pacific sector of the western Arctic Ocean during sea-ice melting. PLoS One 9:e86887. https://doi.org/10.1371/journal.pone.0086887
Bano N, Hollibaugh JT (2002) Phylogenetic composition of bacterioplankton assemblages from the Arctic Ocean. Appl Environ Microbiol 68:505–518
Yakimov MM, Timmis KN, Golyshin PN (2007) Obligate oil-degrading marine bacteria. Curr Opin Biotechnol 18:257–266
Giudice AL, Bruni V, de Domenico M & Michaud L (2010) Psychrophiles-cold-adapted hydrocarbon-degrading microorganisms. Springer Berlin Heidelberg.
Green DH, Llewellyn LE, Negri AP, Blackburn SI, Bolch CJS (2004) Phylogenetic and functional diversity of the cultivable bacterial community associated with the paralytic shellfish poisoning dinoflagellate Gymnodinium catenatum. FEMS Microbiol Ecol 47:345–357
Lea-Smith DJ, Biller SJ, Davey MP, Cotton CAR, Perez Sepulveda BM, Turchyn AV, Scanlan DJ, Smith AG, Chisholm SW, Howe CJ (2015) Contribution of cyanobacterial alkane production to the ocean hydrocarbon cycle. Proc Natl Acad Sci 112:13591–13596
Taylor JD, Cottingham SD, Billinge J, Cunliffe M (2014) Seasonal microbial community dynamics correlate with phytoplankton-derived polysaccharides in surface coastal waters. ISME J 8:245–248
Tully BJ, Tully BJ, Sachdeva R, Heidelberg KB, Heidelberg JF (2014) Comparative genomics of planktonic Flavobacteriaceae from the Gulf of Maine using metagenomic data. Microbiome 2:1–14. https://doi.org/10.1186/2049-2618-2-34
Nikrad MP, Cottrell MT, Kirchman DL (2012) Abundance and single-cell activity of heterotrophic bacterial groups in the western Arctic Ocean in summer and winter. Appl Environ Microbiol 78:2402–2409
Malmstrom R, Straza T, Cottrell M, Kirchman D (2007) Diversity, abundance, and biomass production of bacterial groups in the western Arctic Ocean. Aquat Microb Ecol 47:45–55
Abell GCJ, Bowman JP (2005) Ecological and biogeographic relationships of class Flavobacteria in the Southern Ocean. FEMS Microbiol Ecol 51:265–277
Grzymski JJ, Riesenfeld CS, Williams TJ, Dussaq AM, Ducklow H, Erickson M, Cavicchioli R, Murray AE (2012) A metagenomic assessment of winter and summer bacterioplankton from Antarctica Peninsula coastal surface waters. ISME J 6:1901–1915
Ladau J, Sharpton TJ, Finucane MM, Jospin G, Kembel SW, O'Dwyer J, Koeppel AF, Green JL, Pollard KS (2013) Global marine bacterial diversity peaks at high latitudes in winter. ISME J 7:1669–1677
Gosink JJ, Woese CR, Staley JT (1998) Polaribacter gen. nov., with three new species, P. irgensii sp. nov., P. franzmannii sp. nov. and P. filamentus sp. nov., gas vacuolate polar marine bacteria of the Cytophaga-Flavobacterium-Bacteroides group and reclassification of ‘Flectobacillus glomera’. Int J Syst Bacteriol 48:223–235
Staley JT, Gosink JJ (1999) Poles apart: biodiversity and biogeography of sea ice bacteria. Annu Rev Microbiol 53:189–215
Gonzalez JM, Fernandez-Gomez B, Fernandez-Guerra A, Gomez-Consarnau L, Sanchez O, Coll-Llado M, del Campo J, Escudero L, Rodriguez-Martinez R, Alonso-Saez L, Latasa M, Paulsen I, Nedashkovskaya O, Lekunberri I, Pinhassi J, Pedros-Alio C (2008) Genome analysis of the proteorhodopsin-containing marine bacterium Polaribacter sp. MED152 (Flavobacteria). Proc Natl Acad Sci 105:8724–8729
Delmont TO, Hammar KM, Ducklow HW, Yager PL, Post AF (2014) Phaeocystis antarctica blooms strongly influence bacterial community structures in the Amundsen Sea polynya. Front Microbiol 5:1–13. https://doi.org/10.3389/fmicb.2014.00646
Wollenburg JE, Katlein C, Nehrke G, Nöthig EM, Matthiessen J, Wolf- Gladrow DA, Nikolopoulos A, Gázquez-Sanchez F, Rossmann L, Assmy P, Babin M, Bruyant F, Beaulieu M, Dybwad C, Peeken I (2018) Ballasting by cryogenic gypsum enhances carbon export in a Phaeocystis under-ice bloom. Sci Rep 8:1–9. https://doi.org/10.1038/s41598-018-26016-0
Alonso-Sáez L, Sánchez O, Gasol JM, Balagué V, Pedrós-Alio C (2008) Winter-to-summer changes in the composition and single-cell activity of near-surface Arctic prokaryotes. Environ Microbiol 10:2444–2454
Alonso-Saez L, Waller AS, Mende DR, Bakker K, Farnelid H, Yager PL, Lovejoy C, Tremblay JE, Potvin M, Heinrich F, Estrada M, Riemann L, Bork P, Pedros-Alio C, Bertilsson S (2012) Role for urea in nitrification by polar marine Archaea. Proc Natl Acad Sci 109:17989–17994
DeLong EF, Wu KY, Prézelin BB, Jovine RVM (1994) High abundance of Archaea in Antarctic marine picoplankton. Nature 371:695–697
Kirchman DL, Elifantz H, Dittel AI, Malmstrom RR, Cottrell MT (2007) Standing stocks and activity of Archaea and Bacteria in the western Arctic Ocean. Limnol Oceanogr 52:495–507
Kim J-G, Park SJ, Sinninghe Damsté JS, Schouten S, Rijpstra WIC, Jung MY, Kim SJ, Gwak JH, Hong H, Si OJ, Lee SH, Madsen EL, Rhee SK (2016) Hydrogen peroxide detoxification is a key mechanism for growth of ammonia-oxidizing archaea. Proc Natl Acad Sci 113:7888–7893
Tolar BB, Powers LC, Miller WL, Wallsgrove NJ, Popp BN, Hollibaugh JT (2016) Ammonia oxidation in the ocean can be inhibited by nanomolar concentrations of hydrogen peroxide. Front Mar Sci 3:1–16. https://doi.org/10.3389/fmars.2016.00237
Christman GD, Cottrell MT, Popp BN, Gier E, Kirchman DL (2011) Abundance, diversity, and activity of ammonia-oxidizing prokaryotes in the coastal arctic ocean in summer and winter. Appl Environ Microbiol 77:2026–2034
Pedneault E, Galand PE, Potvin M, Tremblay JÉ, Lovejoy C (2014) Archaeal amoA and ureC genes and their transcriptional activity in the Arctic Ocean. Sci Rep 4:4661. https://doi.org/10.1038/srep04661
Berg IA, Kockelkorn D, Buckel W, Fuchs G (2007) A 3-hydroxypropionate/4-hydroxybutyrate autotrophic carbon dioxide assimilation pathway in archaea. Science 318:1782–1786
Hatzenpichler R (2012) Diversity, physiology, and niche differentiation of ammonia-oxidizing archaea. Appl Environ Microbiol 78:7501–7510
Connelly TL, Baer SE, Cooper JT, Bronk DA (2014) Urea uptake and carbon fixation by marine pelagic bacteria and archaea during the Arctic summer and winter seasons. Appl Environ Microbiol 80:6013–6022
Mußmann M et al (2011) Thaumarchaeotes abundant in refinery nitrifying sludges express amoA but are not obligate autotrophic ammonia oxidizers. Proc Natl Acad Sci 108:16771–16776
Hawley AK, Brewer HM, Norbeck AD, Pa a-Toli L, Hallam SJ (2014) Metaproteomics reveals differential modes of metabolic coupling among ubiquitous oxygen minimum zone microbes. Proc Natl Acad Sci 111:11395–11400
Needham DM, Sachdeva R, Fuhrman JA (2017) Ecological dynamics and co-occurrence among marine phytoplankton, bacteria and myoviruses shows microdiversity matters. ISME J 11:1614–1629
Tolar BB, Wallsgrove NJ, Popp BN, Hollibaugh JT (2017) Oxidation of urea-derived nitrogen by thaumarchaeota-dominated marine nitrifying communities. Environ Microbiol 19:4838–4850
Grossmann L, Bock C, Schweikert M, Boenigk J (2016) Small but Manifold-Hidden Diversity in ‘Spumella-like Flagellates’. J Eukaryot Microbiol 63:419–439
Nolte V et al (2010) Contrasting seasonal niche separation between rare and abundant taxa conceals the extent of protist diversity. Mol Ecol 19:2908–2915
Rottberger J, Gruber A, Boenigk J, Kroth PG (2013) Influence of nutrients and light on autotrophic, mixotrophic and heterotrophic freshwater chrysophytes. Aquat Microb Ecol 71:179–191
McKie-Krisberg ZM, Sanders RW (2014) Phagotrophy by the picoeukaryotic green alga Micromonas: implications for Arctic Oceans. ISME J 8:1953–1961
Lovejoy C, Vincent WF, Bonilla S, Roy S, Martineau MJ, Terrado R, Potvin M, Massana R, Pedrós-Alió C (2007) Distribution, phylogeny, and growth of cold-adapted picoprasinophytes in arctic seas. J Phycol 43:78–89
Foulon E, Not F, Jalabert F, Cariou T, Massana R, Simon N (2008) Ecological niche partitioning in the picoplanktonic green alga Micromonas pusilla: evidence from environmental surveys using phylogenetic probes. Environ Microbiol 10:2433–2443
Kilias E, Kattner G, Wolf C, Frickenhaus S, Metfies K (2014) A molecular survey of protist diversity through the central Arctic Ocean. Polar Biol 37:1271–1287
Acknowledgments
The author AGGS would like to address a special acknowledgement to the Master in Cell and Molecular Biology (M:BCM) held during 2015-17 at Faculty of Sciences, University of Porto (FCUP). We also would like to thank Dr. Anna Silyakova (Centre for Arctic Gas Hydrate, Environment and Climate, Tromsø) for critical reviewing the section “High frequencies of hydrocarbon-degrading bacteria in the Arctic.”
Funding
This work was financially supported by the Centre for Ice, Climate and Ecosystems at the Norwegian Polar Institute and the Research Council of Norway (project Boom or Bust no. 244646) and Structured Program of R&D&I MarInfo-NORTE-01-0145-FEDER-000031, funded by the NORTE2020 through the European Regional Development Fund (ERDF). MF-M and PA were supported by Norwegian Ministries of Foreign Affairs and Climate and Environment through the program Arktis 2030 (project ID Arctic). This project was also funded by PROPOLAR through a grant to CM and a scholarship to AGGS (NITRONICE project) and by Portuguese Science and Technology Foundation (FCT) through a grant to CM (NITROLIMIT-PTDC 2017-PTDC/CTA-AMB/30997/2017).
Author information
Authors and Affiliations
Contributions
AGGS carried out the bioinformatics analysis of SSU rRNA amplicon datasets, analyzed all the data, and wrote the manuscript. MPT and CM supervised all the work. CM and PD designed the sampling campaign. PD and MFM collected and filtrated the samples aboard the R.V. Lance. AGGS and CM extracted the DNA to sequence. LT developed R scripts used in this study. JS, RBL, and JBPL performed phylogenetic analysis. MPT and HR critically reviewed the bioinformatics pipelines, particularly, the 18S rRNA amplicon analysis, and the marine hydrocarbon-degrading bacteria part, respectively. PA and CM funded the work. All authors improved, reviewed, and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
The authors declare that they have no competing interests.
Electronic supplementary material
Supplementary Figure S1
- Correspondence between samples and water mass based on conservative temperature (°C) and absolute salinity (g·kg-1). The values of the physical variables were retrieved from Table 1. Only seven dots are displayed in the scatter plot since the conservative temperature and absolute salinity are the same for the samples NB_5, NB_50, TR_5, and TR_50. AW stands for Atlantic Water; PSW, Polar Surface Water; MAW, Modified Atlantic Water; PSWw, warm Polar Surface Water.
Supplementary Figure S2
- Microbial ecological networks for the top 100 prokaryotic and eukaryotic OTUs. a Network of the top 100 prokaryotic and eukaryotic OTUs. b Network highlighting the SAR11-Dinophyceae associations. c Network highlighting the Alcanivorax-Marinobacter-Dinophyceae associations. d Network highlighting the Thaumarchaeota-Nitrospinae-Chlorohyta associations. e Network highlighting the Bacteroidetes-Diatomea-Phaeocystis-SAR92 associations. Positive associations are represented by green edges and negative associations by red edges. Node size is proportional to the geometric mean of the relative abundance of OTUs. The “Otu” prefix “p” means prokaryotic and the “e” means eukaryotic.
Supplementary Figure S3
– Comparison of taxonomic profiles of 16S rRNA genes obtained with amplicon and metagenomic datasets. Percentage of 16S rRNA reads retrieved from the surface (5 m), subsurface (20 or 50 m) and mesopelagic (250 m) seawater at Nansen Basin (NB, 09.03.2015), Transition Region (TR, 27.04.2015) and Yermak Plateau (YP, 16.06.2015; see Supplementary Table S1). a Percentage of the most abundant taxa (≥ 20% across samples) from metagenomic 16S rRNA reads. b Absolute number of metagenomic 16S rRNA reads. c Percentage of Alcanivorax and Marinobacter genera from 16S rRNA amplicon reads. d Percentage of Alcanivorax and Marinobacter genera from metagenomic 16S rRNA reads.
Supplementary Figure S4
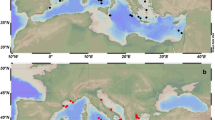
– Biogeographical distribution of environmental Alcanivorax 16S rRNA gene sequences. The maximum likelihood tree under the GTRGAMMA model with 1000 bootstrap replicates includes 38 nucleotide sequences spanning 357 bp positions. Two Alcanivorax-OTUs from the 16S rRNA amplicon dataset (“pOtu00002” and “pOtu00028”), six Alcanivorax metagenomic 16S rRNA reads recovered from the metagenomic 16S rRNA reads dataset (from sample NB_250, named “MG-1”-“MG-6”), and twenty nine Alcanivorax metagenomic sequences from the 16S rRNA Public Assembled Metagenomes database (identified through the accession number of the project in IMG/M). It was assumed a mesopelagic depth (200–1000 m) for sequences retrieved from “Ga0031697” and “Ga0031693” since both reported the collection depth as “deep ocean”. The 16S rRNA gene sequence of Rhodospirillum sulfurexigens JA143T (NCBI accession number AM710622.1) was included as outgroup. The scale bar represents nucleotide substitutions per site. Bootstrap values >50% are displayed.
Supplementary Table S1
– Features of sampling conditions of microbial N-ICE2015 collection.
Supplementary Table S2
- Description of 16S rRNA libraries from N-ICE2015 project.
Supplementary Table S3
- Description of 18S rRNA libraries from N-ICE2015 project.
Supplementary Table S4
- Distribution of prokaryotic taxa across N-ICE2015 collection at phylum, class, order, family, genus and OTU levels. Percentage of taxa at given taxonomic level, inclusive the absolute number of reads assigned to each OTU (97% sequence similarity threshold), within the samples and across the prokaryotic N-ICE2015 collection. The OTU table includes just prokaryotic taxa (see the lineages removed in the “Methods” section) classified against SILVA reference database (v. 1.2.8) after excluding the rare clusters (<5 observations across samples) and rarefying at even sampling depth (38,232 sequences).
Supplementary Table S5
- Raw prokaryotic OTU table from the 16S rRNA libraries of N-ICE2015 collection. The OTU table includes all the taxa (inclusive those lineages that were removed from the main OTU table used in this manuscript, see the “Methods” section) classified against SILVA reference database (v. 1.2.8) without excluding the rare clusters neither rarefying (the no. of sequences goes from 59,627, at TR_5, to 212,529, at YP_5).
Supplementary Table S6
- Distribution of eukaryotic taxa across N-ICE2015 collection at phylum, class, order, family, genus and OTU levels. Percentage of taxa at given taxonomic level, inclusive the absolute number of reads assigned to each OTU (98% sequence similarity threshold), within the samples and across the eukaryotic N-ICE2015 collection. The OTU table includes just interesting protist taxa (see the lineages removed in the “Methods” section) classified against SILVA reference database (v. 1.2.8) after excluding the rare clusters (<5 observations across samples) and rarefying at even sampling depth (43,289 sequences).
Supplementary Table S7
- Eukaryotic OTU table from the 18S rRNA libraries of N-ICE2015 collection assigned against the Protist Ribosomal Reference database (PR2). Percentage and absolute number of reads assigned to each OTU (98% sequence similarity threshold), within the samples and across the eukaryotic N-ICE2015 collection. The OTU table includes protist taxa (see the lineages removed in the “Methods” section) classified against PR2 (v. 4.5) after excluding the rare clusters (<5 observations across samples) and rarefying at even sampling depth (43,647 sequences).
Supplementary Table S8
– Prokaryotic taxonomic table from the metagenomic 16S rRNA reads dataset.
Rights and permissions
About this article
Cite this article
de Sousa, A.G.G., Tomasino, M.P., Duarte, P. et al. Diversity and Composition of Pelagic Prokaryotic and Protist Communities in a Thin Arctic Sea-Ice Regime. Microb Ecol 78, 388–408 (2019). https://doi.org/10.1007/s00248-018-01314-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-01314-2