Abstract
Denitrifying bacteria play a critical role in the estuarine nitrogen cycle. Through the transformation of nitrate into nitrogen gas, these organisms contribute to the loss of bioavailable (i.e., fixed) nitrogen from low-oxygen environments such as estuary sediments. Denitrifiers have been shown to vary in abundance and diversity across the spatial environmental gradients that characterize estuaries, such as salinity and nitrogen availability; however, little is known about how their communities change in response to temporal changes in those environmental properties. Here, we present a 1-year survey of sediment denitrifier communities along the estuarine salinity gradient of San Francisco Bay. We used quantitative PCR and sequencing of functional genes coding for a key denitrifying enzyme, dissimilatory nitrite reductase, to compare two groups of denitrifiers: those with nirK (encoding copper-dependent nitrite reductase) and those with nirS (encoding the cytochrome-cd 1-dependent variant). We found that nirS was consistently more abundant and more diverse than nirK in all parts of the estuary. The abundances of the two genes were tightly linked across space but differed temporally, with nirK peaking when temperature was low and nirS peaking when nitrate was high. Likewise, the diversity and composition of nirK- versus nirS-type communities differed in their responses to seasonal variations, though both were strongly determined by site. Furthermore, our sequence libraries detected deeply branching clades with no cultured isolates, evidence of enormous diversity within the denitrifiers that remains to be explored.






Similar content being viewed by others
References
Seitzinger S, Harrison JA, Böhlke JK et al (2006) Denitrification across landscapes and waterscapes: a synthesis. Ecol Appl Publ Ecol Soc Am 16:2064–2090
Jones CM, Stres B, Rosenquist M, Hallin S (2008) Phylogenetic analysis of nitrite, nitric oxide, and nitrous oxide respiratory enzymes reveal a complex evolutionary history for denitrification. Mol Biol Evol 25:1955–1966. doi:10.1093/molbev/msn146
Rinaldo S, Cutruzzola F (2007) Nitrite reductases in denitrification. In: Bothe H, Ferguson SJ, Newton WE (eds) Biology of the nitrogen cycle. Elsevier, Amsterdam, pp 37–55
Jones CM, Hallin S (2010) Ecological and evolutionary factors underlying global and local assembly of denitrifier communities. ISME J 4:633–641. doi:10.1038/ismej.2009.152
Abell GCJ, Revill AT, Smith C et al (2009) Archaeal ammonia oxidizers and nirS-type denitrifiers dominate sediment nitrifying and denitrifying populations in a subtropical macrotidal estuary. ISME J 4:286–300. doi:10.1038/ismej.2009.105
Braker G, Zhou J, Wu L et al (2000) Nitrite reductase genes (nirK and nirS) as functional markers to investigate diversity of denitrifying bacteria in Pacific Northwest marine sediment communities. Appl Environ Microbiol 66:2096–2104. doi:10.1128/AEM.66.5.2096-2104.2000
Francis CA, O’Mullan GD, Cornwell JC, Ward BB (2013) Transitions in nirS-type denitrifier diversity, community composition, and biogeochemical activity along the Chesapeake Bay Estuary. Front Aquat Microbiol 4:237. doi:10.3389/fmicb.2013.00237
Mosier AC, Francis CA (2010) Denitrifier abundance and activity across the San Francisco Bay estuary. Environ Microbiol Rep 2:667–676. doi:10.1111/j.1758-2229.2010.00156.x
Smith JM, Mosier AC, Francis CA (2015) Spatiotemporal relationships between the abundance, distribution, and potential activities of ammonia-oxidizing and denitrifying microorganisms in intertidal sediments. Microb Ecol 69:13–24. doi:10.1007/s00248-014-0450-1
Nogales B, Timmis KN, Nedwell DB, Osborn AM (2002) Detection and diversity of expressed denitrification genes in estuarine sediments after reverse transcription-PCR amplification from mRNA. Appl Environ Microbiol 68:5017–5025. doi:10.1128/AEM.68.10.5017-5025.2002
Braker G, Fesefeldt A, Witzel K-P (1998) Development of PCR primer systems for amplification of nitrite reductase genes (nirK and nirS) to detect denitrifying bacteria in environmental samples. Appl Environ Microbiol 64:3769–3775
Santoro AE, Boehm AB, Francis CA (2006) Denitrifier community composition along a nitrate and salinity gradient in a coastal aquifer. Appl Environ Microbiol 72:2102–2109. doi:10.1128/AEM.72.3.2102-2109.2006
Wei W, Isobe K, Nishizawa T et al (2015) Higher diversity and abundance of denitrifying microorganisms in environments than considered previously. ISME J. doi:10.1038/ismej.2015.9
Helen D, Kim H, Tytgat B, Anne W (2016) Highly diverse nirK genes comprise two major clades that harbour ammonium-producing denitrifiers. BMC Genomics 17:155. doi:10.1186/s12864-016-2465-0
Dang H, Wang C, Li J et al (2009) Diversity and distribution of sediment nirS-encoding bacterial assemblages in response to environmental gradients in the eutrophied Jiaozhou Bay, China. Microb Ecol 58:161–169. doi:10.1007/s00248-008-9469-5
Dong LF, Smith CJ, Papaspyrou S et al (2009) Changes in benthic denitrification, nitrate ammonification, and anammox process rates and nitrate and nitrite reductase gene abundances along an estuarine nutrient gradient (the Colne Estuary, United Kingdom). Appl Environ Microbiol 75:3171–3179. doi:10.1128/AEM.02511-08
Smith CJ, Nedwell DB, Dong LF, Osborn AM (2007) Diversity and abundance of nitrate reductase genes (narG and napA), nitrite reductase genes (nirS and nrfA), and their transcripts in estuarine sediments. Appl Environ Microbiol 73:3612–3622. doi:10.1128/AEM.02894-06
Law CS, Rees AP, Owens NJP (1991) Temporal variability of denitrification in estuarine sediments. Estuar Coast Shelf Sci 33:37–56. doi:10.1016/0272-7714(91)90069-N
Giblin AE, Weston NB, Banta GT et al (2010) The effects of salinity on nitrogen losses from an oligohaline estuarine sediment. Estuar Coasts 33:1054–1068. doi:10.1007/s12237-010-9280-7
Cornwell JC, Glibert PM, Owens MS (2014) Nutrient fluxes from sediments in the San Francisco Bay delta. Estuar Coasts 37:1120–1133. doi:10.1007/s12237-013-9755-4
Eyre BD, Maher DT, Squire P (2013) Quantity and quality of organic matter (detritus) drives N2 effluxes (net denitrification) across seasons, benthic habitats and estuaries. Glob Biogeochem Cycles 27:1083–1095. doi:10.1002/2013GB004631
Smith CJ, Dong LF, Wilson J et al (2015) Seasonal variation in denitrification and dissimilatory nitrate reduction to ammonia process rates and corresponding key functional genes along an estuarine nitrate gradient. Front Microbiol 6:542. doi:10.3389/fmicb.2015.00542
Conomos TJ, Smith RE, Gartner JW (1985) Environmental setting of San Francisco Bay. Hydrobiologia 129:1–12. doi:10.1007/BF00048684
Nichols FH, Cloern JE, Luoma SN, Peterson DH (1986) The modification of an estuary. Science 231:567–573. doi:10.1126/science.231.4738.567
Hager SW, Schemel LE (1992) Sources of nitrogen and phosphorus to Northern San Francisco Bay. Estuaries 15:40–52. doi:10.2307/1352708
Wankel SD, Kendall C, Francis CA, Paytan A (2006) Nitrogen sources and cycling in the San Francisco Bay estuary: a nitrate dual isotopic composition approach. Limnol Oceanogr 51:1654–1664
Caffrey JM (1995) Spatial and seasonal patterns in sediment nitrogen remineralization and ammonium concentrations in San Francisco Bay, California. Estuaries 18:219–233. doi:10.2307/1352632
Grenz C, Cloern JE, Hager SW, Cole BE (2000) Dynamics of nutrient cycling and related benthic nutrient and oxygen fluxes during a spring phytoplankton bloom in South San Francisco Bay (USA). Mar Ecol Prog Ser 197:67–80. doi:10.3354/meps197067
Bower CE, Holm-Hansen T (1980) A salicylate–hypochlorite method for determining ammonia in seawater. Can J Fish Aquat Sci 37:794–798. doi:10.1139/f80-106
Henry S, Baudoin E, López-Gutiérrez JC et al (2004) Quantification of denitrifying bacteria in soils by nirK gene targeted real-time PCR. J Microbiol Methods 59:327–335, doi:10.1016/j.mimet.2004.07.002
Suzuki MT, Taylor LT, DeLong EF (2000) Quantitative analysis of small-subunit rRNA genes in mixed microbial populations via 5′-nuclease assays. Appl Environ Microbiol 66:4605–4614. doi:10.1128/AEM.66.11.4605-4614.2000
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. doi:10.1093/bioinformatics/bts199
Edgar RC (2013) UPARSE: highly accurate OTU sequences from microbial amplicon reads. Nat Methods 10:996–998. doi:10.1038/nmeth.2604
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Stat 11:265–270
Hill MO (1973) Diversity and evenness: a unifying notation and its consequences. Ecology 54:427–432
Lozupone C, Lladser ME, Knights D et al (2011) UniFrac: an effective distance metric for microbial community comparison. ISME J 5:169–172. doi:10.1038/ismej.2010.133
Webb CO, Ackerly DD, McPeek MA, Donoghue MJ (2002) Phylogenies and community ecology. Annu Rev Ecol Syst 33:475–505. doi:10.1146/annurev.ecolsys.33.010802.150448
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217. doi:10.1371/journal.pone.0061217
Kembel SW, Cowan PD, Helmus MR et al (2010) Picante: R tools for integrating phylogenies and ecology. Bioinformatics 26:1463–1464. doi:10.1093/bioinformatics/btq166
Fish JA, Chai B, Wang Q et al (2013) FunGene: the functional gene pipeline and repository. Front Microbiol 4:291. doi:10.3389/fmicb.2013.00291
Letunic I, Bork P (2011) Interactive Tree Of Life v2: online annotation and display of phylogenetic trees made easy. Nucleic Acids Res 39:W475–W478. doi:10.1093/nar/gkr201
Oksanen J, Blanchet FG, Kindt R et al (2014) vegan: community ecology package
Abell G, Ross D, Keane J et al (2013) Nitrifying and denitrifying microbial communities and their relationship to nutrient fluxes and sediment geochemistry in the Derwent Estuary, Tasmania. Aquat Microb Ecol 70:63–75. doi:10.3354/ame01642
Anderson MJ (2001) A new method for non-parametric multivariate analysis of variance. Austral Ecol 26:32–46. doi:10.1111/j.1442-9993.2001.01070.pp.x
R Core Team (2014) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Graf DRH, Jones CM, Hallin S (2014) Intergenomic comparisons highlight modularity of the denitrification pathway and underpin the importance of community structure for N2O emissions. PLoS One 9:e114118. doi:10.1371/journal.pone.0114118
Kandeler E, Deiglmayr K, Tscherko D et al (2006) Abundance of narG, nirS, nirK, and nosZ genes of denitrifying bacteria during primary successions of a glacier foreland. Appl Environ Microbiol 72:5957–5962. doi:10.1128/AEM.00439-06
Procheş Ş, Wilson JRU, Cowling RM (2006) How much evolutionary history in a 10 × 10 m plot? Proc R Soc Lond B Biol Sci 273:1143–1148. doi:10.1098/rspb.2005.3427
Lozupone CA, Knight R (2007) Global patterns in bacterial diversity. Proc Natl Acad Sci 104:11436–11440. doi:10.1073/pnas.0611525104
Jayakumar A, O’Mullan GD, Naqvi SWA, Ward BB (2009) Denitrifying bacterial community composition changes associated with stages of denitrification in oxygen minimum zones. Microb Ecol 58:350–362. doi:10.1007/s00248-009-9487-y
Yoshida M, Ishii S, Otsuka S, Senoo K (2009) Temporal shifts in diversity and quantity of nirS and nirK in a rice paddy field soil. Soil Biol Biochem 41:2044–2051. doi:10.1016/j.soilbio.2009.07.012
Fortunato CS, Carlini DB, Ewers E, Bushaw-Newton C (2009) Nitrifier and denitrifier molecular operational taxonomic unit compositions from sites of a freshwater estuary of Chesapeake Bay. Can J Microbiol 55:333–346
Li M, Hong Y, Cao H et al (2013) Diversity, abundance, and distribution of NO-forming nitrite reductase–encoding genes in deep-sea subsurface sediments of the South China Sea. Geobiology 11:170–179. doi:10.1111/gbi.12020
Kim O-S, Imhoff JF, Witzel K-P, Junier P (2010) Distribution of denitrifying bacterial communities in the stratified water column and sediment–water interface in two freshwater lakes and the Baltic Sea. Aquat Ecol 45:99–112. doi:10.1007/s10452-010-9335-7
Li J, Wei G, Wang N, Gao Z (2014) Diversity and distribution of nirK-harboring denitrifying bacteria in the water column in the Yellow River estuary. Microbes Environ JSME 29:107–110
Fan H, Bolhuis H, Stal LJ (2015) Denitrification and the denitrifier community in coastal microbial mats. FEMS Microbiol Ecol 91:fiu033. doi:10.1093/femsec/fiu033
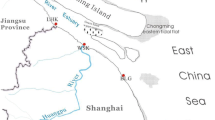
Dang H, Wang J (2007) Diversity and spatial distribution of sediment nirS-type denitrifying communities in response to environmental gradients in Changjiang Estuary and East China Sea. In: NCBI PopSet 16474642 Uncultured Bact. Clone S33-N-45 Putative Dissimilatory Reductase NirS Gene Partial Cds., http://www.ncbi.nlm.nih.gov/popset/164374642. Accessed 17 Jan 2015
Jayakumar DA, Francis CA, Naqvi SWA, Ward BB (2004) Diversity of nitrite reductase genes (nirS) in the denitrifying water column of the coastal Arabian Sea. Aquat Microb Ecol 34:69–78
Oakley BB, Francis CA, Roberts KJ et al (2007) Analysis of nitrite reductase (nirK and nirS) genes and cultivation reveal depauperate community of denitrifying bacteria in the Black Sea suboxic zone. Environ Microbiol 9:118–130. doi:10.1111/j.1462-2920.2006.01121.x
Bulow SE, Francis CA, Jackson GA, Ward BB (2008) Sediment denitrifier community composition and nirS gene expression investigated with functional gene microarrays. Environ Microbiol 10:3057–3069. doi:10.1111/j.1462-2920.2008.01765.x
Bowen JL, Byrnes JE, Weisman D, Colaneri C (2013) Functional gene pyrosequencing and network analysis: an approach to examine the response of denitrifying bacteria to increased nitrogen supply in salt marsh sediments. Front Terr Microbiol 4:342. doi:10.3389/fmicb.2013.00342
Hammond DE, Fuller C, Harmon D et al (1985) Benthic fluxes in San Francisco Bay. Hydrobiologia 129:69–90. doi:10.1007/BF00048688
Mosier AC, Francis CA (2008) Relative abundance and diversity of ammonia-oxidizing archaea and bacteria in the San Francisco Bay estuary. Environ Microbiol 10:3002–3016. doi:10.1111/j.1462-2920.2008.01764.x
Lee JA (2015) Biogeography of nitrogen-cycling microbial communities in San Francisco Bay, Doctoral Dissertation, Stanford University, Department of Environmental Earth System Science
Acknowledgments
This work was supported by NSF CAREER Grant OCE-0847266 (to C.A.F.) and by a Stanford Graduate Fellowship (from William R. and Sara Hart Kimball) and a Marshall-EPA Fellowship (to J.A.L.). We thank Julian Damashek for his extensive help with a great many aspects of this work, from assisting with sample collection to providing feedback on data presentation, and especially for carrying out the chemical analysis of the bottom-water samples. Arushi Atluri provided invaluable assistance with acquisition and processing of the nirK sequences. Finally, also owe extensive thanks to Jim Cloern, Jessica Dyke, Amy Kleckner, Jan Thompson, and the other USGS scientists and staff who made our work on the R/V Polaris possible.
Author information
Authors and Affiliations
Corresponding author
Additional information
The nucleotide sequences reported in this study have been deposited in GenBank under accession nos. KR060094—KR060621 (for nirK) and KR060622—KR061281 (for nirS).
Rights and permissions
About this article
Cite this article
Lee, J.A., Francis, C.A. Spatiotemporal Characterization of San Francisco Bay Denitrifying Communities: a Comparison of nirK and nirS Diversity and Abundance. Microb Ecol 73, 271–284 (2017). https://doi.org/10.1007/s00248-016-0865-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0865-y