Abstract
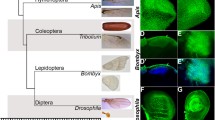
Hox genes encode Homeodomain-containing transcription factors, which specify segmental identities along the anterior–posterior axis. Functional changes in Hox genes have been directly implicated in the evolution of body plans across the metazoan lineage. The Hox protein Ultrabithorax (Ubx) is expressed and required in developing third thoracic (T3) segments in holometabolous insects studied so far, particularly, of the order Coleoptera, Lepidoptera and Diptera. Ubx function is key to specify differential development of the second (T2) and T3 thoracic segments in these insects. While Ubx is expressed in the third thoracic segment in developing larvae of Hymenopteran Apis mellifera, the morphological differences between T2 and T3 are subtle. To identify evolutionary changes that are behind the differential function of Ubx in Drosophila and Apis, which are diverged for more than 350 million years, we performed comparative analyses of genome wide Ubx-binding sites between these two insects. Our studies reveal that a motif with a TAAAT core is a preferred binding site for Ubx in Drosophila, but not in Apis. Biochemical and transgenic assays suggest that in Drosophila, the TAAAT core sequence in the Ubx binding sites is required for Ubx-mediated regulation of two of its target genes studied here; CG13222, a gene that is normally upregulated by Ubx and vestigial (vg), whose expression is repressed by Ubx in T3. Interestingly, changing the TAAT site to a TAAAT site was sufficient to bring an otherwise unresponsive enhancer of the vg gene from Apis under the control of Ubx in a Drosophila transgenic assay. Taken together, our results suggest an evolutionary mechanism by which critical wing patterning genes might have come under the regulation of Ubx in the Dipteran lineage.



Similar content being viewed by others
Data availability
Raw fastq files of ChIP sequencing for the Ubx protein in Apis mellifera hindwings can be accessed from NCBI (GSE71847). Raw fastq files of ChIP sequencing for the Ubx protein in Drosophila melanogaster halteres and RNA sequencing data for wing and haltere imaginal discs can be accessed from NCBI (GSE205177 and GSE205352).
References
Agrawal P, Habib F, Yelagandula R, Shashidhara LS (2011) Genome-level identification of targets of Hox protein Ultrabithorax in Drosophila: novel mechanisms for target selection. Sci Rep 1(i): 1–10. https://doi.org/10.1038/srep00205
Anders S, Pyl PT, Huber W (2015) HTSeq—a Python framework to work with high-throughput sequencing data. Bioinformatics 31(2):166–169. https://doi.org/10.1093/bioinformatics/btu638
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, Davis AP, Dolinski K, Dwight SS, Eppig JT et al (2000) Gene ontology: tool for the unification of biology. Nat Genet 25(1):25–29. https://doi.org/10.1038/75556
Bailey TL, Boden M, Buske FA, Frith M, Grant CE, Clementi L, Ren J, Li WW, Noble WS (2009) MEME suite: tools for motif discovery and searching. Nucleic Acids Res 37(SUPPL. 2):202–208. https://doi.org/10.1093/nar/gkp335
Cabrera CV, Botas J, Garcia-Bellido A (1985) Distribution of Ultrabithorax proteins in mutants of Drosophila bithorax complex and its transregulatory genes. Nature 318(6046):569–571. https://doi.org/10.1038/318569a0
Carbon S, Douglass E, Good BM, Unni DR, Harris NL, Mungall CJ, Basu S, Chisholm RL, Dodson RJ, Hartline E et al (2021) The gene ontology resource: enriching a GOld mine. Nucleic Acids Res 49(D1):D325–D334. https://doi.org/10.1093/nar/gkaa1113
Carroll SB (1995) Homeotic genes and the evolution of arthropods and chordates. Nature 376(6540):479–485. https://doi.org/10.1038/376479a0
Casares F, Mann RS (2000) A dual role for homothorax in inhibiting wing blade development and specifying proximal wing identities in Drosophila. Development 127(7):1499–1508. https://doi.org/10.1242/dev.127.7.1499
Castelli-Gair JE, Micol JL, Garcia-Bellido A (1990) Transvection in the Drosophila Ultrabithorax gene: a Cbx1 mutant allele induces ectopic expression of a normal allele in trans. Genetics 126(1):177–184. https://doi.org/10.1093/genetics/126.1.177
Choo SW, White R, Russell S (2011) Genome-wide analysis of the binding of the Hox protein Ultrabithoraxand the Hox cofactor Homothorax in Drosophila. PLoS One 6(4):e14778. https://doi.org/10.1371/journal.pone.0014778
Crocker J, Abe N, Rinaldi L, McGregor AP, Frankel N, Wang S, Alsawadi A, Valenti P, Plaza S, Payre F et al (2015) Low affinity binding site clusters confer Hox specificity and regulatory robustness. Cell 160(1–2):191–203. https://doi.org/10.1016/j.cell.2014.11.041
Galant R, Walsh CM, Carroll SB (2002) Hox repression of a target gene: extradenticle-independent, additive action through multiple monomer binding sites. Development 129(13):3115–3126. https://doi.org/10.1242/dev.129.13.3115
Heinz S, Benner C, Spann N, Bertolino E, Lin YC, Laslo P, Cheng JX, Murre C, Singh H, Glass CK (2010) Simple combinations of lineage-determining transcription factors prime cis-regulatory elements required for macrophage and B cell identities. Mol Cell 38(4):576–589. https://doi.org/10.1016/j.molcel.2010.05.004
Hersh BM, Carroll SB (2005) Direct regulation of knot gene expression by Ultrabithorax and the evolution of cis-regulatory elements in Drosophila. Development 132(7):1567–1577. https://doi.org/10.1242/dev.01737
Hersh BM, Nelson CE, Stoll SJ, Norton JE, Albert TJ, Carroll SB (2007) The UBX-regulated network in the haltere imaginal disc of D. melanogaster. Dev Biol 302: 717–727. https://doi.org/10.1016/j.ydbio.2006.11.011
Hrycaj SM, Wellik DM (2016) Hox genes and evolution. F1000Research 5: 859. https://doi.org/10.12688/f1000research.7663.1
Hudry B, Remacle S, Delfini MC, Rezsohazy R, Graba Y, Merabet S (2012) Hox proteins display a common and ancestral ability to diversify their interaction mode with the pbc class cofactors. PLoS Biol 10(6): e1001349. https://doi.org/10.1371/journal.pbio.1001351
Khan S, Dilsha C, Shashidhara LS (2020) Haltere development in D. melanogaster: implications for the evolution of appendage size, shape and function. Int J Dev Biol 64(1–3): 169–175. https://doi.org/10.1387/ijdb.190133LS
Kim J, Sebring A, Esch JJ, Kraus ME, Vorwerk K, Magee J, Carroll SB (1996) Integration of positional signals and regulation of wing formation and identity by Drosophila vestigial gene. Nature 382(6587):133–138. https://doi.org/10.1038/382133a0
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12(4):357–360. https://doi.org/10.1038/nmeth.3317
Klein T, Arias AM (1998) Different spatial and temporal interactions between Notch, wingless, and vestigial specify proximal and distal pattern elements of the wing in Drosophila. Dev Biol 194(2):196–212. https://doi.org/10.1006/dbio.1997.8829
Lewis EB (1978) A gene complex controlling segmentation in Drosophila. In: Lixpshitz HD (ed) Genes, development and cancer. The life and work of Edward B. Lewis, pp. 229–242. https://doi.org/10.1007/978-1-4020-6345-9_10
Lewis EB (1982) Control of body segment differentiation in Drosophila by the bithorax gene complex. Prog Clin Biol Res 85(Pt A): 269–288.
Loker R, Sanner JE, Mann RS (2021) Cell-type-specific Hox regulatory strategies orchestrate tissue identity. Curr Biol 31(19):4246-4255.e4. https://doi.org/10.1016/j.cub.2021.07.030
Madeira F, Park YM, Lee J, Buso N, Gur T, Madhusoodanan N, Basutkar P, Tivey ARN, Potter SC, Finn RD et al (2019) The EMBL-EBI search and sequence analysis tools APIs in 2019. Nucleic Acids Res 47(W1):W636–W641. https://doi.org/10.1093/nar/gkz268
Matsuoka Y, Monteiro A (2021) Hox genes are essential for the development of eyespots in Bicyclus anynana butterflies. Genetics 217: iyaa005. https://doi.org/10.1093/genetics/iyaa005
McGinnis W, Krumlauf R (1992) Homeobox genes and axial patterning. Cell 68(2):283–302. https://doi.org/10.1016/0092-8674(92)90471-N
Mohit P, Makhijani K, Madhavi MB, Bharathi V, Lal A, Sirdesai G, Reddy VR, Ramesh P, Kannan R, Dhawan J et al (2006) Modulation of AP and DV signaling pathways by the homeotic gene ultrabithorax during haltere development in Drosophila. Dev Biol 291:356–367. https://doi.org/10.1016/j.ydbio.2005.12.022
Neumann CJ, Cohen SM (1996) A hierarchy of cross-regulation involving Notch, wingless, vestigial and cut organizes the dorsal/ventral axis of the Drosophila wing. Development 122(11):3477–3485. https://doi.org/10.1242/dev.122.11.3477
Nguyen NTT, Contreras-Moreira B, Castro-Mondragon JA, Santana-Garcia W, Ossio R, Robles-Espinoza CD, Bahin M, Collombet S, Vincens P, Thieffry D et al (2018) RSAT 2018: regulatory sequence analysis tools 20th anniversary. Nucleic Acids Res 46(W1):W209–W214. https://doi.org/10.1093/nar/gky317
Pick L, Heffer A (2012) Hox gene evolution: multiple mechanisms contributing to evolutionary novelties: evolutionary changes in Hox function. Ann N Y Acad Sci 1256(1):15–32. https://doi.org/10.1111/j.1749-6632.2011.06385.x
Prasad N, Tarikere S, Khanale D, Habib F, Shashidhara LS (2016) A comparative genomic analysis of targets of Hox protein ultrabithorax amongst distant insect species. Sci Rep 6(1):27885. https://doi.org/10.1038/srep27885
Ramírez F, Ryan DP, Grüning B, Bhardwaj V, Kilpert F, Richter AS, Heyne S, Dündar F, Manke T (2016) deepTools2: a next generation web server for deep-sequencing data analysis. Nucleic Acids Res 44(W1):W160–W165. https://doi.org/10.1093/nar/gkw257
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140. https://doi.org/10.1093/bioinformatics/btp616
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26. https://doi.org/10.1038/nbt.1754
Sánchez-Higueras C, Rastogi C, Voutev R, Bussemaker HJ, Mann RS, Hombría JCG (2019) In vivo Hox binding specificity revealed by systematic changes to a single cis regulatory module. Nat Commun 10(1). https://doi.org/10.1038/s41467-019-11416-1
Shashidhara LS, Agrawal N, Bajpai R, Bharathi V, Sinha P (1999) Negative regulation of dorsoventral signaling by the homeotic gene ultrabithorax during haltere development in Drosophila. Dev Biol 212(2):491–502. https://doi.org/10.1006/dbio.1999.9341
Shlyueva D, Meireles-Filho ACA, Pagani M, Stark A (2016) Genome-wide ultrabithorax binding analysis reveals highly targeted genomic loci at developmental regulators and a potential connection to polycomb-mediated regulation. PLOS ONE 11(8): e0161997. https://doi.org/10.1371/journal.pone.0161997
Slattery M, Riley T, Liu P, Abe N, Gomez-Alcala P, Dror I, Zhou T, Rohs R, Honig B, Bussemaker HJ et al (2011) Cofactor binding evokes latent differences in DNA binding specificity between Hox proteins. Cell 147(6):1270–1282. https://doi.org/10.1016/j.cell.2011.10.053
Tomoyasu Y, Wheeler SR, Denell RE (2005) Ultrabithorax is required for membranous wing identity in the beetle Tribolium castaneum. Nature 433(7026):643–647. https://doi.org/10.1038/nature03272
Walsh CM, Carroll SB (2007) Collaboration between Smads and a Hox protein in target gene repression. Development 134(20):3585–3592. https://doi.org/10.1242/dev.009522
Waterhouse AM, Procter JB, Martin DMA, Clamp M, Barton GJ (2009) Jalview version 2-A multiple sequence alignment editor and analysis workbench. Bioinformatics 25(9):1189–1191. https://doi.org/10.1093/bioinformatics/btp033
Weatherbee SD, Halder G, Kim J, Hudson A, Carroll S (1998) Ultrabithorax regulates genes at several levels of the wing-patterning hierarchy to shape the development of the Drosophila haltere. Genes Dev 12(10):1474–1482. https://doi.org/10.1101/gad.12.10.1474
Weatherbee SD, Nijhout HF, Grunert LW, Halder G, Galant R, Selegue J, Carroll S (1999) Uitrabithorax function in butterfly wings and the evolution of insect wing patterns. Curr Biol 9(3):109–115. https://doi.org/10.1016/S0960-9822(99)80064-5
White RAH, Akam ME (1985) Contrabithorax mutations cause inappropriate expression of Ultrabithorax products in Drosophila. Nature 318(6046):567–569. https://doi.org/10.1038/318567a0
White RAH, Wilcox M (1985) Regulation of the distribution of Ultrabithorax proteins in Drosophila. Nature 318(6046):563–567. https://doi.org/10.1038/318563a0
Williams JA, Bell JB, Carroll SB (1991) Control of Drosophila wing and haltere development by the nuclear vestigial gene product. Genes Dev 5(12 PART B): 2481–2495. https://doi.org/10.1101/gad.5.12b.2481
Zhang Y, Liu T, Meyer CA, Eeckhoute J, Johnson DS, Bernstein BE, Nussbaum C, Myers RM, Brown M, Li W et al (2008) Model-based analysis of ChIP-Seq (MACS). Genome Biol 9(9). https://doi.org/10.1186/gb-2008-9-9-r137
Acknowledgements
We thank G. Deshpande and members of the LSS and SM laboratories for critical input. We thank S. Galande for providing reagents for Library preparation. This work was supported primarily by an Indo-French Research grant from CEFIPRA to SM; a JC Bose Fellowship and grant from the Department of Science and Technology, Government of India to LSS; and a Council of Scientific and Industrial Research (CSIR) Fellowship to SK.
Author information
Authors and Affiliations
Contributions
SK carried out all fly experiments, image analyses, and wrote the MS. SK and SP did ChIP-Seq and RNA-Seq and analyses. GG, FB and RP did EMSA experiments. LSS and SM conceived the project and wrote the MS.
Corresponding authors
Ethics declarations
Conflict of interest
We declare “no-conflict-of-interest”.
Additional information
Handling editor: Willie Swanson.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Khan, S., Pradhan, S.J., Giraud, G. et al. A Micro-evolutionary Change in Target Binding Sites as a Key Determinant of Ultrabithorax Function in Drosophila. J Mol Evol 91, 616–627 (2023). https://doi.org/10.1007/s00239-023-10123-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-023-10123-2