Abstract
Purpose
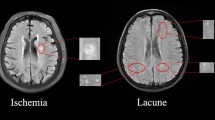
White matter hyperintensity (WMHI) lesions on MR images are an important indication of various types of brain diseases that involve inflammation and blood vessel abnormalities. Automated quantification of the WMHI can be valuable for the clinical management of patients, but existing automated software is often developed for a single type of disease and may not be applicable for clinical scans with thick slices and different scanning protocols. The purpose of the study is to develop and validate an algorithm for automatic quantification of white matter hyperintensity suitable for heterogeneous MRI data with different disease types.
Methods
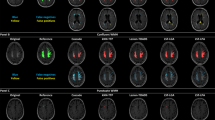
We developed and evaluated “DeepWML”, a deep learning method for fully automated white matter lesion (WML) segmentation of multicentre FLAIR images. We used MRI from 507 patients, including three distinct white matter diseases, obtained in 9 centres, with a wide range of scanners and acquisition protocols. The automated delineation tool was evaluated through quantitative parameters of Dice similarity, sensitivity and precision compared to manual delineation (gold standard).
Results
The overall median Dice similarity coefficient was 0.78 (range 0.64 ~ 0.86) across the three disease types and multiple centres. The median sensitivity and precision were 0.84 (range 0.67 ~ 0.94) and 0.81 (range 0.64 ~ 0.92), respectively. The tool’s performance increased with larger lesion volumes.
Conclusion
DeepWML was successfully applied to a wide spectrum of MRI data in the three white matter disease types, which has the potential to improve the practical workflow of white matter lesion delineation.




Similar content being viewed by others
Data availability
Data that support the findings in this study are available from the corresponding author upon reasonable request.
Code availability
The source code generated during the current study is available from the corresponding author on reasonable request.
References
Rovira A, León A (2008) MR in the diagnosis and monitoring of multiple sclerosis: an overview. Eur J Radiol 67(3):409–414
Wattjes MP, Rovira À, Miller D, Yousry TA, Sormani MP, de Stefano MP et al (2015) Evidence-based guidelines: MAGNIMS consensus guidelines on the use of MRI in multiple sclerosis–establishing disease prognosis and monitoring patients. Nat Rev Neurol 11(10):597–606
Asgari N, Skejoe HPB, Lillevang ST, Steenstrup T, Stenager E, Kyvik KO (2013) Modifications of longitudinally extensive transverse myelitis and brainstem lesions in the course of neuromyelitis optica (NMO): a population-based, descriptive study. BMC Neurol 13:33–33
Dutra BG, da Rocha AJ, Nunes RH, Maia ACMJ (2018) Neuromyelitis optica spectrum disorders: spectrum of MR imaging findings and their differential diagnosis. Radiographics : a review publication of the Radiological Society of North America, Inc. 38(1): p. 169–193
Zhang X, Tang Y, Xie Y, Ding C, Xiao J, Jiang X et al (2017) Total magnetic resonance imaging burden of cerebral small-vessel disease is associated with post-stroke depression in patients with acute lacunar stroke. Eur J Neurol 24(2):374–380
Li G, Zhu C, Li J, Wang X, Zhang Q, Zheng H et al (2018) Increased level of procalcitonin is associated with total MRI burden of cerebral small vessel disease in patients with ischemic stroke. Neurosci Lett 662:242–246
McKinley R, Wepfer R, Grunder L, Aschwanden F, Fischer T, Friedli C et al (2019) Automatic detection of lesion load change in multiple sclerosis using convolutional neural networks with segmentation confidence. NeuroImage Clinical 25:102104–102104
Salem M, Valverde S, Cabezas M, Pareto D, Oliver A, Salvi J et al (2019) A fully convolutional neural network for new T2-w lesion detection in multiple sclerosis. NeuroImage Clinical 25:102149–102149
Zijdenbos AP, Forghani R, Evans AC (2002) Automatic “pipeline” analysis of 3-D MRI data for clinical trials: application to multiple sclerosis. IEEE Trans Med Imaging 21(10):1280–1291
García-Lorenzo D, Francis S, Narayanan S, Arnold DL, Collins DL (2013) Review of automatic segmentation methods of multiple sclerosis white matter lesions on conventional magnetic resonance imaging. Med Image Anal 17(1):1–18
Dadar M, Maranzano J, Misquitta K, Anor CJ, Fonov VS, Tartaglia MC, et al. Performance comparison of 10 different classification techniques in segmenting white matter hyperintensities in aging. (1095–9572 (Electronic)).
Diniz PHB, Valente TLA, Diniz JOB, Silva AC, Gattass M, Ventura N et al (2018) Detection of white matter lesion regions in MRI using SLIC0 and convolutional neural network. Comput Methods Programs Biomed 167:49–63
Li H, Jiang G, Zhang J, Wang R, Wang Z, Zheng W-S et al (2018) Fully convolutional network ensembles for white matter hyperintensities segmentation in MR images. Neuroimage 183:650–665
Xu B, Chai Y, Galarza CM, Vu CQ, Tamrazi B, Gaonkar B et al (2018) Orchestral fully convolutional networks for small lesion segmentation in brain MRI. . Proceedings IEEE International Symposium on Biomedical Imaging 2018:889–892
Duong MT, Rudie JD, Wang J, Xie L, Mohan S, Gee JC et al (2019) Convolutional neural network for automated FLAIR lesion segmentation on clinical brain MR imaging. AJNR Am J Neuroradiol 40(8):1282–1290
Schmidt P, Gaser C, Arsic M, Buck D, Förschler A, Berthele A et al (2012) An automated tool for detection of FLAIR-hyperintense white-matter lesions in multiple sclerosis. Neuroimage 59(4):3774–3783
Rachmadi MF, Valdés-Hernández MdC, Agan MLF, Di Perri C, Komura T (2018) Segmentation of white matter hyperintensities using convolutional neural networks with global spatial information in routine clinical brain MRI with none or mild vascular pathology. Computerized Medical Imaging and Graphics. 66: p. 28-43
La Rosa F, Abdulkadir A, Fartaria MJ, Rahmanzadeh R, P.-J. Lu, R. Galbusera, et al. (2020). Multiple sclerosis cortical and WM lesion segmentation at 3T MRI: a deep learning method based on FLAIR and MP2RAGE. NeuroImage: Clinical. 27: p. 102335
Mongan JA-O, Moy LA-O, Kahn CEJA-O. Checklist for artificial intelligence in medical imaging (CLAIM): a guide for authors and reviewers. (2638–6100 (Electronic))
Heinen R, Steenwijk MD, Barkhof F, Biesbroek JM, van der Flier WM, Kuijf HJ et al (2019) Performance of five automated white matter hyperintensity segmentation methods in a multicenter dataset. Sci Rep 9(1):16742–16742
Le M, Tang LYW, Hernández-Torres E, Jarrett M, Brosch T, Metz L et al (2019) FLAIR(2) improves LesionTOADS automatic segmentation of multiple sclerosis lesions in non-homogenized, multi-center, 2D clinical magnetic resonance images. NeuroImage Clinical 23:101918–101918
Thompson AJ, Banwell BL, Barkhof F, Carroll WM, Coetzee T, Comi G et al (2018) Diagnosis of multiple sclerosis: 2017 revisions of the McDonald criteria. The Lancet Neurology 17(2):162–173
Wingerchuk DM, Banwell B, Bennett JL, Cabre P, Carroll W, Chitnis T et al (2015) International consensus diagnostic criteria for neuromyelitis optica spectrum disorders. Neurology 85(2):177–189
Wardlaw JM, Smith EE, Biessels GJ, Cordonnier C, Fazekas F, Frayne R et al (2013) Neuroimaging standards for research into small vessel disease and its contribution to ageing and neurodegeneration. The Lancet Neurology 12(8):822–838
Fedorov A, Beichel R, Kalpathy-Cramer J, Finet J, Fillion-Robin J-C, Pujol S et al (2012) 3D Slicer as an image computing platform for the Quantitative Imaging Network. Magn Reson Imaging 30(9):1323–1341
Ronneberger O, Fischer P, Brox T (2015) U-Net: Convolutional networks for biomedical image segmentation. Medical Image Computing and Computer-Assisted Intervention – MICCAI 2015. Cham: Springer International Publishing
Sudre CH, Li W, Vercauteren T, Ourselin S, Cardoso MJ (2017) Generalised Dice overlap as a deep learning loss function for highly unbalanced segmentations. Deep Learning in Medical Image Analysis and Multimodal Learning for Clinical Decision Support. Cham: Springer International Publishing
Kingma DP, Ba J (2014) Adam: a method for stochastic optimization. arXiv e-prints: p. arXiv:1412.6980
Paszke A, Gross S, Massa F, Lerer A, Bradbury J, Chanan G, et al. (2019). PyTorch: An Imperative Style, High-Performance Deep Learning Library. arXiv e-prints: p. arXiv:1912.01703
Ribaldi F, Altomare D, Jovicich J, Ferrari C, Picco A, Pizzini FB et al (2021) Accuracy and reproducibility of automated white matter hyperintensities segmentation with lesion segmentation tool: A European multi-site 3T study. Magn Reson Imaging 76:108–115
Steenwijk MD, Pouwels PJW, Daams M, van Dalen JW, Caan MWA, Richard E et al (2013) Accurate white matter lesion segmentation by k nearest neighbor classification with tissue type priors (kNN-TTPs). NeuroImage Clinical 3:462–469
Damangir S, Westman E, Simmons A, Vrenken H, Wahlund L-O, Spulber G (2017) Reproducible segmentation of white matter hyperintensities using a new statistical definition. Magma (New York, N.Y.). 30(3): p. 227–237
Acknowledgements
Frederik Barkhof is supported by the NIHR biomedical research center at UCLH. The authors are grateful to Dr. Susumu Mori for his helpful suggestions.
Funding
This work was supported by National Natural Science Foundation of China (grant numbers: 81571631, 81870958), Beijing Nova Program (grant number: xx2013045) and Natural Science Foundation of Beijing Municipality (grant number: 7133244).
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Data collection, methodology and data analysis were performed by Yajing Zhang, Yunyun Duan and Xiaoyang Wang. The first draft of the manuscript was written by Yajing Zhang and Yaou Liu, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethics approval
This study was approved by the institutional review boards of Beijing Tiantan Hospital, Capital Medical University, Beijing, China. The procedures used in this study adhere to the tenets of the Declaration of Helsinki.
Consent to participate
Written informed consent was obtained from each participant.
Consent for publication
Consent for publication was obtained from all participants.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Zhang, Y., Duan, Y., Wang, X. et al. A deep learning algorithm for white matter hyperintensity lesion detection and segmentation. Neuroradiology 64, 727–734 (2022). https://doi.org/10.1007/s00234-021-02820-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-021-02820-w