Abstract
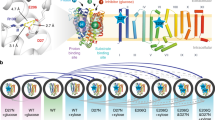
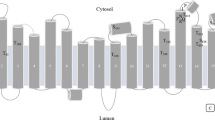
Vigna radiata H+–translocating pyrophosphatases (VrH+-PPases, EC 3.6.1.1) are present in various endomembranes of plants, bacteria, archaea, and certain protozoa. They transport H+ into the lumen by hydrolyzing pyrophosphate, which is a by-product of many essential anabolic reactions. Although the crystal structure of H+-PPases has been elucidated, the H+ translocation mechanism of H+-PPases in the solution state remains unclear. In this study, we used hydrogen–deuterium exchange (HDX) coupled with mass spectrometry (MS) to investigate the dynamics of H+-PPases between the previously proposed R state (resting state, Apo form), I state (intermediate state, bound to a substrate analog), and T state (transient state, bound to inorganic phosphate). When hydrogen was replaced by proteins in deuterium oxide solution, the backbone hydrogen atoms, which were exchanged with deuterium, were identified through MS. Accordingly, we used deuterium uptake to examine the structural dynamics and conformational changes of H+-PPases in solution. In the highly conserved substrate binding and proton exit regions, HDX-MS revealed the existence of a compact conformation with deuterium exchange when H+-PPases were bound with a substrate analog and product. Thus, a novel working model was developed to elucidate the in situ catalytic mechanism of pyrophosphate hydrolysis and proton transport. In this model, a proton is released in the I state, and the TM5 inner wall serves as a proton piston.
Graphical Abstract








Similar content being viewed by others
Abbreviations
- DDM:
-
n-Dodecyl-β-d-maltoside
- D2O:
-
Deuterium oxide
- IDP:
-
Imidodiphosphate
- MALDI:
-
Matrix-assisted laser desorption/ionization
- VrH+-PPase:
-
Vigna radiata H+–translocating pyrophosphatase
References
Anashkin VA, Baykov AA (2021) A lumenal loop associated with catalytic asymmetry in plant vacuolar H(+)-translocating pyrophosphatase. Int J Mol Sci. https://doi.org/10.3390/ijms222312902
Asaoka M, Segami S, Maeshima M (2014) Identification of the critical residues for the function of vacuolar H(+)-pyrophosphatase by mutational analysis based on the 3D structure. J Biochem 156:333–344. https://doi.org/10.1093/jb/mvu046
Bao A-K, Wang S-M, Wu G-Q, Xi J-J, Zhang J-L, Wang C-M (2009) Overexpression of the Arabidopsis H+-PPase enhanced resistance to salt and drought stress in transgenic alfalfa (Medicago sativa L.). Plant Sci 176:232–240. https://doi.org/10.1016/j.plantsci.2008.10.009
Belogurov GA, Lahti R (2002) A lysine substitute for K+. A460K mutation eliminates K+ dependence in H+-pyrophosphatase of Carboxydothermus hydrogenoformans. J Biol Chem 277:49651–49654. https://doi.org/10.1074/jbc.M210341200
Craig R, Cortens JP, Beavis RC (2004) Open source system for analyzing, validating, and storing protein identification data. J Proteome Res 3:1234–1242. https://doi.org/10.1021/pr049882h
Engen JR (2009) Analysis of protein conformation and dynamics by hydrogen/deuterium exchange MS. Anal Chem 81:7870–7875. https://doi.org/10.1021/ac901154s
Etxeberria E, Pozueta-Romero J, Gonzalez P (2012) In and out of the plant storage vacuole. Plant Sci 190:52–61. https://doi.org/10.1016/j.plantsci.2012.03.010
Gaxiola RA, Palmgren MG, Schumacher K (2007) Plant proton pumps. Febs Lett 581:2204–2214. https://doi.org/10.1016/j.febslet.2007.03.050
Hedrich R, Kurkdjian A, Guern J, Flugge UI (1989) Comparative studies on the electrical properties of the H+ translocating ATPase and pyrophosphatase of the vacuolar-lysosomal compartment. EMBO J 8:2835–2841
Hsiao YY, Van RC, Hung SH, Lin HH, Pan RL (2004) Roles of histidine residues in plant vacuolar H(+)-pyrophosphatase. Biochim Biophys Acta 1608:190–199. https://doi.org/10.1016/j.bbabio.2004.01.001
Hsiao YY, Pan YJ, Hsu SH, Huang YT, Liu TH, Lee CH et al (2007) Functional roles of arginine residues in mung bean vacuolar H+-pyrophosphatase. Biochim Biophys Acta 1767:965–973. https://doi.org/10.1016/j.bbabio.2007.04.007
Hsu SH, Lo YY, Liu TH, Pan YJ, Huang YT, Sun YJ et al (2015) Substrate-induced changes in domain interaction of vacuolar H(+)-pyrophosphatase. J Biol Chem 290:1197–1209. https://doi.org/10.1074/jbc.M114.568139
Huang YT, Liu TH, Chen YW, Lee CH, Chen HH, Huang TW et al (2010) Distance variations between active sites of H(+)-pyrophosphatase determined by fluorescence resonance energy transfer. J Biol Chem 285:23655–23664. https://doi.org/10.1074/jbc.M110.134916
Jiang L, Phillips TE, Hamm CA, Drozdowicz YM, Rea PA, Maeshima M et al (2001) The protein storage vacuole: a unique compound organelle. J Cell Biol 155:991–1002. https://doi.org/10.1083/jcb.200107012
Kan ZY, Ye X, Skinner JJ, Mayne L, Englander SW (2019) ExMS2: an integrated solution for hydrogen-deuterium exchange mass spectrometry data analysis. Anal Chem 91:7474–7481. https://doi.org/10.1021/acs.analchem.9b01682
Kellosalo J, Kajander T, Kogan K, Pokharel K, Goldman A (2012) The structure and catalytic cycle of a sodium-pumping pyrophosphatase. Science 337:473–476. https://doi.org/10.1126/science.1222505
Khadilkar AS, Yadav UP, Salazar C, Shulaev V, Paez-Valencia J, Pizzio GA et al (2016) Constitutive and companion cell-specific overexpression of AVP1, encoding a proton-pumping pyrophosphatase, enhances biomass accumulation, phloem loading, and long-distance transport. Plant Physiol 170:401–414. https://doi.org/10.1104/pp.15.01409
Lee CH, Pan YJ, Huang YT, Liu TH, Hsu SH, Lee CH et al (2011) Identification of essential lysines involved in substrate binding of vacuolar H+-pyrophosphatase. J Biol Chem 286:11970–11976. https://doi.org/10.1074/jbc.M110.190215
Li KM, Wilkinson C, Kellosalo J, Tsai JY, Kajander T, Jeuken LJ et al (2016) Membrane pyrophosphatases from Thermotoga maritima and Vigna radiata suggest a conserved coupling mechanism. Nat Commun 7:13596. https://doi.org/10.1038/ncomms13596
Lin SM, Tsai JY, Hsiao CD, Huang YT, Chiu CL, Liu MH et al (2012) Crystal structure of a membrane-embedded H+-translocating pyrophosphatase. Nature 484:399–403. https://doi.org/10.1038/nature10963
Liu S, Zheng L, Xue Y, Zhang Q, Wang L, Shou H (2010) Overexpression of OsVP1 and OsNHX1 increases tolerance to drought and salinity in Rice. J Plant Biol 53:444–452. https://doi.org/10.1007/s12374-010-9135-6
Luoto HH, Belogurov GA, Baykov AA, Lahti R, Malinen AM (2011) Na+-translocating membrane pyrophosphatases are widespread in the microbial world and evolutionarily precede H+-translocating pyrophosphatases. J Biol Chem 286:21633–21642. https://doi.org/10.1074/jbc.M111.244483
Maeshima M (2000) Vacuolar H(+)-pyrophosphatase. Biochim Biophys Acta 1465:37–51
Manabe F, Shoun H, Wakagi T (2011) Conserved residues in membrane-bound acid pyrophosphatase from Sulfolobus tokodaii, a thermoacidophilic archaeon. Extremophiles 15:359–364. https://doi.org/10.1007/s00792-011-0367-2
Martinoia E, Maeshima M, Neuhaus HE (2007) Vacuolar transporters and their essential role in plant metabolism. J Exp Bot 58:83–102. https://doi.org/10.1093/jxb/erl183
Mimura H, Nakanishi Y, Maeshima M (2005) Oligomerization of H(+)-pyrophosphatase and its structural and functional consequences. Biochim Biophys Acta 1708:393–403. https://doi.org/10.1016/j.bbabio.2005.05.004
Nakanishi Y, Saijo T, Wada Y, Maeshima M (2001) Mutagenic analysis of functional residues in putative substrate-binding site and acidic domains of vacuolar H+-pyrophosphatase. J Biol Chem 276:7654–7660. https://doi.org/10.1074/jbc.M009743200
Segami S, Nakanishi Y, Sato MH, Maeshima M (2010) Quantification, organ-specific accumulation and intracellular localization of type II H(+)-pyrophosphatase in Arabidopsis thaliana. Plant Cell Physiol 51:1350–1360. https://doi.org/10.1093/pcp/pcq096
Serrano A, Perez-Castineira JR, Baltscheffsky M, Baltscheffsky H (2007) H+-PPases: yesterday, today and tomorrow. IUBMB Life 59:76–83. https://doi.org/10.1080/15216540701258132
Tsai JY, Tang KZ, Li KM, Hsu BL, Chiang YW, Goldman A et al (2019) Roles of the hydrophobic gate and exit channel in vigna radiata pyrophosphatase ion translocation. J Mol Biol 431:1619–1632. https://doi.org/10.1016/j.jmb.2019.03.009
Vidilaseris K, Kiriazis A, Turku A, Khattab A, Johansson NG, Leino TO et al (2019) Asymmetry in catalysis by Thermotoga maritima membrane-bound pyrophosphatase demonstrated by a nonphosphorus allosteric inhibitor. Sci Adv 5:eaav7574. https://doi.org/10.1126/sciadv.aav7574
Yang SJ, Jiang SS, Hsiao YY, Van RC, Pan YJ, Pan RL (2004) Thermoinactivation analysis of vacuolar H(+)-pyrophosphatase. Biochim Biophys Acta 1656:88–95. https://doi.org/10.1016/j.bbabio.2004.02.001
Author information
Authors and Affiliations
Contributions
L-KH: Formal analysis, Writing - Original Draft Visualization, Writing - Review & Editing Y-CH: Validation Formal analysis, Investigation, Data Curation P-CC: ValidationFormal analysis, Investigation C-HL: Validation S-ML: Validation Y-HHH: Methodology Resources, Data Curation, Writing - Review & Editing, Supervision, Project administration R-LP: Conceptualization Ideas, Resources Writing - Review & Editing, Supervision, Project administration
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Huang, LK., Huang, YC., Chen, PC. et al. Exploration of the Catalytic Cycle Dynamics of Vigna Radiata H+–Translocating Pyrophosphatases Through Hydrogen–Deuterium Exchange Mass Spectrometry. J Membrane Biol 256, 443–458 (2023). https://doi.org/10.1007/s00232-023-00295-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-023-00295-9