Abstract
Nucleus is at the center stage of cellular drama orchestrated in the life of a cell and the nucleoplasm is surrounded by a double membranous compartment constituting the Nuclear membrane/envelope (NE) that separates it from the cytoplasm in nucleated cells. The initial understanding of the NE was that of a border security entity between the nucleus and the cytoplasm, separating gene regulation and transcription in the nucleus from translation in the cytoplasm. However, the discovery of a wide array of inherited diseases caused by mutations in genes encoding proteins that reside or interact with NE diverted the interest into deciphering the lipid-protein-rich environment of the NE. Today, the NE is considered a dynamic organelle which forms a functional linkage between the nucleus and the rest of the cell. The exposure of NE to constant mechanical constraints by its connectivity to the large polymer network of the lamina and chromatin on one side, and to the cytoskeleton on the other side results, in a variety of shape changes. We discuss two such deformation, the formation of nuclear blebs and nucleoplasmic reticulum (NER). Although the protein and the lipid composition of NE comprises a small fraction of the total lipid-protein load of the cell, the ability to define the lipid-protein composition of Inner nuclear membrane (INM) and Outer nuclear membrane (ONM) with precision is crucial for obtaining a deeper mechanistic understanding of their lipid-protein interaction and the various signaling pathways that are triggered by them. In addition, this allows us to further understand the direct and indirect roles of NE machinery in the chromosomal organization and gene regulation.
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nucleus is at the center stage of cellular drama orchestrated in the life of a cell and the universality of its presence in plants and animals was first proposed in the 1830s by Brown in plants; and Valentin and Henle in animals (Osorio and Gomes, 2013). The history of the initial observations of the nucleus is extremely fascinating and has been covered in detail elsewhere (Osorio and Gomes, 2013). We have provided a schematic of the timeline of the scientists who contributed to this initial journey of observing the nucleus (Fig. 1). The nucleoplasm is surrounded by a double membranous compartment constituting the Nuclear membrane/envelope (NE) that separates it from the cytoplasm in nucleated cells. The initial understanding of the NE since its discovery in 1913 (Kite 1913) has been that of a border security entity between the nucleus and the cytoplasm, separating gene regulation and transcription in the nucleus from translation in the cytoplasm. The switch from the chromatin-centric understanding of the nucleus to unravelling the lipid-protein-rich environment of the NE was spearheaded by the discovery that mutations in genes encoding its protein components cause a wide array of inherited diseases often referred to as laminopathies or nuclear envelopathies. These pathologies include movement disorders and myopathies (Dauer and Worman 2009; Meinke and Schirmer 2016), aging-related diseases causing reduced life span and progeria (Kubben and Misteli 2017; Fichtman et al 2019), lethal defects in the embryo (Turner and Schlieker, 2016), and lipodystrophies (Shackleton et al. 2000). Disruption of NE stability is also common in cancer cells causing DNA damage, cancer-relevant chromosomal rearrangements, and the initiation of pro-inflammatory pathways (Lim et al. 2016; Umbreit and Pellman 2017; Hatch 2018; Selezneva et al. 2022), underscoring the need to unravel its organization and dynamics.
Timeline of the early observations marking the discovery of the nucleus and the nuclear membrane. First observation of the nucleus was reported by Leeuwenhoek in fish erythrocytes, followed by the observations of Trembley, Müller, Ehrenberg, Hewson, Fontana and Bauer. Bown, Valentin and Henle recognized the quasi universality of nucleus in plants and animals. Later in twentieth century, Kite used the term nuclear membrane for the first time to describe a membranous structure around the nucleus. Information obtained from Osorio and Gomes, (2013) and Kite (1913)
Today, the NE is considered a dynamic organelle which forms a functional linkage between the nucleus and the rest of the cell (Fig. 2). Biological mass-spectrometry techniques and analysis algorithms have expanded our knowledge of the nuclear protein repertoire. However, understanding its lipid-protein composition and their interaction remains challenging owing to (i) purifying nuclear membrane without Endoplasmic reticulum (ER) contamination is very difficult and (ii) due to the proximity of the two membranes of NE. Immunogold-label Electron microscopy (EM) remains the method of choice for determining the location of NE residing proteins with precision. In recent years, techniques such as rapamycin trapping, using spit-GFP constructs, metal-induced energy transfer, ensemble Fluorescence recovery after photobleaching (FRAP) and super-resolution microscopy have been adapted to address the differential understanding of Inner nuclear membrane (INM) vs outer nuclear membrane (ONM) (Tingey et al. 2019). In this mini-review, we summarize the current understanding of the lipid-protein machinery of the NE and the dynamic deformations that occur in the NE in physiological and pathological states.
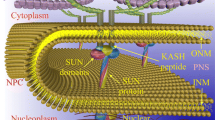
A schematic representation of NE showing the structure and dynamic deformations. NE is a dynamic double membrane organelle which forms a functional linkage between the nucleus and the rest of the cell. The two membranes are bridged by NPCs and the LINC complexes. Two types of deformations observed in the NE are the outward projecting blebs and inward projecting invaginations called NER
Protein Machinery of NE
Protein machinery of NE contributes to 1% (278 proteins) of all human proteins experimentally detected by the Human Protein Atlas with 238 of them having multiple locations outside of NE. Interestingly, 10% of eukaryotic transmembrane proteins are found to be residing in NE (Mudumbi et al. 2020). The ONM is continuous with the rough ER membrane, and they are generally similar in protein content with only a few proteins, namely nesprins and Itprip, preferentially concentrated in the ONM (Méndez-López et al. 2012; Cheng et al. 2019). In contrast, the protein machinery of the INM is still poorly understood. A comprehensive proteomic study identified 13 knowns and 67 putative proteins concentrated in the INM (Schirmer et al. 2003; Méndez-López et al. 2012; Cheng et al. 2019). The few signature proteins of INM include the integral proteins Lamin B receptor (LBR), Lamina-associated polypeptides (LAPs), emerin and MAN1 (also known as LEMD3), Ima1 (Ima1 present in fission yeast is a homologue of the human Samp1 and rat NET5 proteins) and Lem2 (Collas et al. 2000; Tange et al. 2016). Newly identified members of INM include Nurim, Mfsd10, Tmx4, and Arl6ip6 (Chen et al. 2012; Cheng et al. 2019). A recent review covered the current understanding of the diverse cellular functions of INM proteins in detail (Pawar and Kutay 2021).
The two membranes of the NE are separated by 30–50 nm, and NE bridges composed of Sad1 and UNC-84 (SUN) proteins in INM and Klarsicht, Anc-1, and Syne homology (KASH) proteins in ONM have been proposed to set and regulate nuclear envelope spacing (Sosa et al. 2012; Cain et al. 2014). These proteins also constitute the Linker of the nucleoskeleton and cytoskeleton (LINC) complex, which physically connects the nucleus and plasma membrane via the actin cytoskeleton to perform diverse functions including mechano-transduction from the extracellular environment to the nucleus (Ueda et al. 2022).
In addition to the LINC complex, large macromolecular assemblies (~ 100 mDa) constituting NPCs bridge the two membranes of NE. Nuclear pore complexes (NPCs) mediate molecular flux between the nucleus and the cytoplasm and are built by ~ 1000 protein subunits called Nucleoporins (NUP). The biogenesis of NPCs during the interphase of the cell cycle and their insertion in NE by fusion between the INM and ONM has been covered in detail elsewhere (Rothballer and Kutay 2013). NPCs have been traditionally studied as a selective pore allowing the trafficking of molecules through the double lipid bilayers, recent studies highlight its role in chromosomal organization and gene regulation, as it can interact with the genomic region enhancers and super-enhancers (Lin et al. 2019; Pascual-Garcia et al. 2019). Polar molecules, ions and macromolecules are allowed to pass between nucleoplasm and cytoplasm through the NPCs (Hampoelz et al. 2019).
Another important protein constituent of NE are the ion channels which have been discovered using genetic, immunological, pharmacological, and electrophysiological approaches (Matzke et al. 2010). The most well understood ion channels in NE are the Ca2+ channels in animals and plants (Bootman et al., 2009; Oliveira et al. 2014; Secondo et al. 2020; Pirayesh et al. 2021). The Ca2+ channels found in the INM include Inositol (1,4,5)-trisphosphate receptor (IP3R), Ryanodine receptor (RyR), and Nicotinic acid-adenine dinucleotide (NAADP) receptors. The ONM resident Ca2+ channels are IP3R, Ca2+-ATPases and inositol 1,3,4,5-tetrakisphosphate-operated Ca2+ channels (Becchetti 2011). In addition, chloride channels (Gururaja et al. 2020), potassium channels (Jang et al. 2015) Ca2+ -ATPase (Gerasimenko et al. 1995) and Na+/Ca2+ exchangers (Secondo et al. 2020) are also present in the NE.
The INM is lined by the Nuclear lamin (NL), belongs to type V intermediate filament proteins, and is divided into two subtypes, the A and B. The gene LMNA is spliced in two isoforms, the longer version encoding the protein Lamin A and the shorter isoform generating the Lamin C protein. Two different genes (LMNB1 and LMNB2) are responsible for encoding Lamin B1 and B2 proteins. Details of NL and their post-translational modifications and their active role in signaling have been covered elsewhere (Gauthier et al. 2021).
Grease of the NE: Lipids in Action
NE constitute a very small fraction (< 1%) of the total membrane content of the cell (Milo and Phillips 2015) and its composition has been analyzed by mass-spectrometry and multidimensional NMR (31P and 1H) (van Meer et al. 2008; Dazzoni et al. 2020a). NE lipid extract demonstrated a complex mixture of phospholipids with different fatty acyl chain lengths varying between 30 and 38 carbon atoms (two chains summed up) associated predominantly with Phosphatidylcholine (PC) head group (van Meer et al. 2008; Dazzoni et al. 2020a). Negatively charged lipids were also observed with an abundance of Phosphatidylinositol (PI), associated with chain length varying between 36 and 38 carbon (Dazzoni et al. 2020a). NE lipid extracts analyzed by 1H–,13C– and 31P-NMR showed the presence of cholesterol in addition to PE and PI (Dazzoni et al. 2020b). The presence of elevated levels of unsaturated fatty acid chains (with one to two double bonds per lipid species) accentuated the fluidity and elasticity of NE relative to plasma membrane (Dazzoni et al. 2020a, b). Despite the enhanced membrane fluidity, the INM and the ONM are only permeable to small non-polar molecules and sustain nucleoplasm membrane potential approximately − 15 mV with respect to the cytoplasm (Loewenstein and Kanno 1963; Mazzanti et al. 2001).
Recent studies have highlighted that although NE is not a major site of lipid synthesis compared to the ER, changes in NE lipid composition occur as an adaptive stress management response to the maintain the integrity of NE. Studies in yeast, fly and mammalian cells have shown that de novo PC synthesis can take place to relieve the curvature elastic stress and NE breakdown (Haider et al. 2018). Accumulation of very-long-chain fatty acids or phytoceramides by the action of NE resident very-long-chain fatty acid elongase Elo2 has been observed in yeast to prevent lethal defects associated with Lem2 and Bqt4 knockouts, which are conserved nuclear membrane proteins (Kinugasa et al. 2019). Interestingly, targeting ceramide synthesis suppresses nuclear abnormalities and improves the proliferation of aneuploid cells in yeasts and patients associated with Down syndrome (Hwang et al. 2019). In addition, a sphingolipid hydrolase (Smpd4) that releases ceramide spatially localizes to NPCs, suggesting a potential local role for sphingolipids and their precursors at the NE (Cheng et al. 2019).
How is Identity of the NE Maintained?
How is identity of the NE maintained despite the ONM being continuous with the ER? The code to understand the sorting of lipids and proteins within the ER/NE membranes and how NE maintains its identity is still not cracked. The ability to retain proteins in the INM has been linked to their affinity for nuclear components e.g., for LBR, SUN2, LAP2β and the phenomenon termed as “diffusion and retention” (Ungricht et al. 2017). Interestingly, many INM proteins contain NLS-like sequences (Lusk et al. 2007) and Kutay and group proposed that these could function as nuclear retention motifs, e.g., as part of DNA-binding domains (LaCasse and Lefebvre 1995; Cokol et al. 2000).
Recent evidence suggests that lipids produced in the ER are harnessed to remodel nuclear membranes (Barger et al. 2022).
Is there an asymmetry in the lipid composition of INM vs ONM? Specific lipids residing in INM have been shown to support viral proliferation (Marschall et al. 2011), NE dynamics (Hatch and Hetzer 2014), de novo lipid synthesis (Haider et al. 2018; Romanauska and Kohler 2018) and NPC biogenesis (Drin et al. 2007) underscoring the existence of mechanisms that differentially enrich and regulate specific lipid species at the INM. Currently limited knowledge exists to understand these processes and potential mechanisms that drive lipid asymmetry at the NE and lead to NE remodeling, despite the direct continuity of the lipid bilayers of the NE and ER. Two pathways have been proposed to attain this asymmetry: (1) the presence of a physical barrier that reduces the timescale of lateral diffusion for specific lipids from one area (peripheral ER/ONM) to the other (INM) and (2) differential spatial restriction of synthetic enzymes to generate a continuum of concentrations high in one area (e.g., peripheral ER) relative to the other (e.g., INM). These pathways have been discussed in detail in other recent reviews (Bahmanyar and Schlieker 2020; Barger et al. 2022).
NE Membrane Deformations
The exposure of NE to constant mechanical constraints by virtue of its connectivity to the large polymer network of the laimina and chromatin on one side, and to the cytoskeleton on the other side results, in a variety of shape changes. A recent review covered the mechanisms and functions of NE remodeling in exquisite detail (Ungricht and Kutay 2017). Here we discuss, two main types of NE deformations that have been observed: the outward projecting nuclear blebs and the inward projecting nuclear invaginations constituting the nucleoplasmic reticulum (NER).
Nuclear blebs are formed when the double NE separates from the lamina and chromatin, inflates and forms a round protrusion containing nucleoplasm which is not retracted like the plasma membrane blebs and possesses a high likelihood of bursting (Srivastava et al 2021). Nuclear blebs form as spherical protrusions filled with nucleoplasm and devoid of chromatin and are commonly associated with sites of local lamina weakness (Charras et al. 2008; Wiggan et al. 2017; Shah et al 2017). The membrane rupture occurs systematically in these swollen blebs, measuring ~ microns in diameter, as a consequence of internal pressure mounted by translocation of nucleoplasm inside the blebs (Srivastava et al 2021). However, these membrane rupture events are followed by rapid repair (Raab et al. 2016; Earle et al. 2020). Some of these blebs are associated with chromatin herniation, which results in protrusion of the chromatin through the local rupture of the lamina at the base of the bleb. A new lamina eventually reforms on the surface of the herniated chromatin as the herniation is not retracted, leading to a long-term nuclear shape alteration after the resealing of the envelope (Charras et al. 2008). The uncontrolled exchange between the nuclear interior and cytoplasm occurs in the nuclear blebs and these sites prime DNA damage (Shah et al. 2017). Nuclear blebs have been observed in several cancerous cells such as in monoblasts of acute monocytic leukaemia (McDuffie 1967), Burkitt lymphoma cells (Epsteln et al. 1965; Achong and Epstein 1966), and anaplastic giant cell carcinoma of the thyroid (Caryso et al 2011). Accumulation of these blebs has been observed in laminopathies such as premature Hutchinson–Gilford progeria syndrome and Emery–Dreifuss muscular dystrophy (Lattanzi et al. 2016). Interestingly, nuclear blebbing has also been observed in developing human and guineapig thymocytes (Törö and Oláh 1966; Sebuwufu 1966) suggesting NE plasticity during functional development and differentiation of the cells.
The continuity of NE is interrupted by invaginations that reach deep within the nucleoplasm and such a complex branched network of invaginations has been defined as NER (Malhas et al. 2011). Lipids extracts from NER were at least two orders of magnitude more elastic than the classical plasma membrane suggesting a physical explanation for the formation of NER (Dazzoni et al. 2020b). Morphological comparisons with the ER paved the nomenclature of these widespread intra-nuclear invaginations as NER (Echevarría et al. 2003; Fischer et al. 2003). NER structures are classified into 2 main classes: Type I invaginations where the INM alone invaginates into the nucleoplasm, whereas type II where both the INM and ONM enter the nucleoplasm allowing the presence of a cytoplasmic core (Drozdz, and Vaux 2017). The function of NER in physiology and pathology has been covered in detail in recent reviews (Drozdz, and Vaux 2017; Stiekema et al. 2022).
Dynamic Nature of NE
NE is a highly dynamic (temporally) organelle based on its ability and need to deform at small and large length scales. However, sparse studies exist which have looked at the mobility of lipids and protein in NE. Spectrometry analysis of NE of HEK 293 T cells showed that these membrane consist of PC (63%), PE (9%), SM (4%) and PI (12%) (Dazzoni et al. 2020a). Interestingly, cholesterol is also thought to be an important lipid in these membrane although its exact amount is still debatable. Measurements using solid-state NMR showed that the NE lipids derived from these cells are 100 times more elastic than plasma membranes (Dazzoni et al. 2020b). The abundance of unsaturated in the fatty acyl chains of PI coupled with its negative charge is thought to be balancing factor counteracting the rigidifying effect of cholesterol in NE (Dazzoni et al. 2020a).
FRAP based studies have shown that GFP-tagged emerin, MAN1 and LBR-are less mobile in the nuclear envelope than in the ER (Ellenberg et al. 1997; Östlund et al., 1999; Wu et al., 2002). Diffusional mobility (D) of emerin was decreased in INM (D = 0.10 ± 0.01 µm2/second) compared to the ER membrane (D = 0.32 ± 0.01 µm2/second). MAN1 also demonstrated a lower mobility in the INM (0.12 ± 0.02 µm2/second) relative to the ER pool (D ~ 0.28 ± 0.04 µm2/second). Early studies addressing the mobility of proteins in NE membranes using isolated nuclei chemically modified with citraconic acid showed that D for proteins in the INM bound by the fluorescently labeled lectin wheat germ agglutinin was 0.039 mm2/s (Schindler et al., 1984). Enhanced mobility of GFP-tagged emerin and MAN1, but not LBR, was observed in embryonic fibroblasts from lamin A knockout (Lmna−/−) mice implying that emerin and MAN1 are partly retained in the INM by binding to A-type lamins, while LBR depends on other binding partners for its retention.
Conclusion
This is a very exciting time in membrane biology, as the landscape of our understanding of membranes is beginning to expand from the classical plasma membrane to the intracellular organelles. NE is at the center stage of this revolution as we are understanding its many facets with the improvement of technology available to tease out its organization and dynamics. The challenges to overcome include the ability to define the lipid-protein composition of INM and ONM with precision to address the dynamic lipid-protein interaction in these complex membranes. Purifying and isolating nuclear membrane without ER contamination is still considered as the holy grail of the field. The traditional method for nuclei isolation involves the use of non-ionic detergent (Lee et al., 2010), which suffer from the tendency to cause unwanted nuclear aggregation and disrupt the NE resulting in stripping/loss of of some NE proteins in addition to leakage of nuclear matrix materials. Detergent free methods have been described that purify the nucleus without these side effects (Blobel and Potter 1966; Eski et al., 2020).
Solving the challenges mentioned above is quintessential for obtaining a deeper mechanistic understanding of the various signaling pathways that are triggered by NE residents or interacting protein-lipid machinery. In addition, this allows us to further understand its direct and indirect roles in the chromosomal organization and gene regulation.
Data Availability
All data generated or analyzed during this study are included in this published article which are listed in the references.
Abbreviations
- NE:
-
Nuclear membrane/envelope
- ER:
-
Endoplasmic reticulum
- EM:
-
Electron microscopy
- INM:
-
Inner nuclear membrane
- ONM:
-
Outer nuclear membrane
- LBR:
-
Lamin B receptor
- LAPs:
-
Lamina-associated polypeptides
- SUN:
-
Sad1 and UNC-84
- KASH:
-
Klarsicht, Anc-1, and Syne homology
- LINC:
-
Linker of the nucleoskeleton and cytoskeleton
- NPC:
-
Nuclear pore complexes
- NUP:
-
Nucleoporins
- NL:
-
Nuclear lamin
- PC:
-
Phosphatidylcholine
- PI:
-
Phosphatidylinositol
- NER:
-
Nucleoplasmic reticulum
- FRAP:
-
Fluorescence recovery after photobleaching
- D:
-
Diffusional mobility
- GFP:
-
Green fluorescent protein
- IP3R:
-
Inositol (1,4,5)-trisphosphate receptor
- RyR:
-
Ryanodine receptor
- NAADP:
-
Nicotinic acid-adenine dinucleotide
- RBC:
-
Red blood cells
References
Achong BG, Epstein MA (1966) Morphological and virological investigations on cultured Burkitt tumor lymphoblasts (strain Raji). J Nat Cancer Inst 36:877
Bahmanyar S, Schlieker C (2020) Lipid and protein dynamics that shape nuclear envelope identity. Mol Biol Cell 31:1315–1323
Barger SR, Penfield L, Bahmanyar S (2022) Coupling lipid synthesis with nuclear envelope remodeling. Trends Biochem Sci 47:52–65
Becchetti A (2011) Ion channels and transporters in cancer. 1. Ion channels and cell proliferation in cancer. Am J Physiol Cell Physiol 301:C255–C265
Blobel G, Potter VR (1966) Nuclei from rat liver: isolation method that combines purity with high yield. Science 154:1662–1665
Bootman MD, Fearnley C, Smyrnias I, MacDonald F, Roderick HL (2009) An update on nuclear calcium signalling. J Cell Sci 122:2337–2350
Cain NE, Tapley EC, McDonald KL, Cain BM, Starr DA (2014) The SUN protein UNC-84 is required only in force-bearing cells to maintain nuclear envelope architecture. J Cell Biol 206:163–172
Caryso RA, Fedele F, Crisafullj C, Paparo D, Parisi A, Luciano R, Cavallari V (2011) Abnormal nuclear structures (micronuclei, nuclear blebs, strings, and pockets) in a case of anaplastic giant cell carcinoma of the thyroid: an immunohistochemical and ultrastructural study. Ultrastruct Pathol 35:14–18
Charras GT, Coughlin M, Mitchison TJ, Mahadevan L (2008) Life and times of a cellular bleb. Biophys J 94:1836–1853
Chen H, Chen K, Chen J, Cheng H, Zhou R (2012) The integral nuclear membrane protein nurim plays a role in the suppression of apoptosis. Curr Mol Med 12:1372–1382
Cheng L-C, Baboo S, Lindsay C, Brusman L, Martinez-Bartolomé S, Tapia O, Zhang X, Yates JR, Gerace L (2019) Identification of new transmembrane proteins concentrated at the nuclear envelope using organellar proteomics of mesenchymal cells. Nucleus 10:126–143
Cokol M, Nair R, Rost B (2000) Finding nuclear localization signals. EMBO Rep 1:411–415
Collas P, Courvalin JC (2000) Sorting nuclear membrane proteins at mitosis. Trends Cell Biol 10:5–8
Dauer WT, Worman HJ (2009) The nuclear envelope as a signaling node in development and disease. Dev Cell 17:626–638
Dazzoni R, Buré C, Morvan E, Grélard A, Gounou C, Schmitter JM, Loquet A, Larijani B, Dufourc EJ (2020a) Tandem NMR and mass spectrometry analysis of human nuclear membrane lipids. Anal Chem 92:6858–6868
Dazzoni R, Grélard A, Morvan E, Bouter A, Applebee CJ, Loquet A, Larijani B, Dufourc EJ (2020b) The unprecedented membrane deformation of the human nuclear envelope, in a magnetic field, indicates formation of nuclear membrane invaginations. Sci Rep 10:5147
Drin G, Casella JF, Gautier R, Boehmer T, Schwartz TU, Antonny B (2007) A general amphipathic alpha-helical motif for sensing membrane curvature. Nat Struct Mol Biol 14:138–146
Drozdz MM, Vaux DJ (2017) Shared mechanisms in physiological and pathological nucleoplasmic reticulum formation. Nucleus 8:34–45
Earle AJ, Kirby TJ, Fedorchak GR, Isermann P, Patel J, Iruvanti S, Moore SA, Bonne G, Wallrath LL, Lammerding J (2020) Mutant lamins cause nuclear envelope rupture and DNA damage in skeletal muscle cells. Nat Mater 19:464–473
Echevarría W, Leite MF, Guerra MT, Zipfel WR, Nathanson MH (2003) Regulation of calcium signals in the nucleus by a nucleoplasmic reticulum. Nat Cell Biol 5:440–446
Ellenberg J, Siggia ED, Moreira JE, Smith CL, Presley JF, Worman HJ, Lippincott-Schwartz J (1997) Nuclear membrane dynamics and reassembly in living cells: targeting of an inner nuclear membrane protein in interphase and mitosis. J Cell Biol 138:1193–1206
Epsteln MA, Barr YM, Aehong BG (1965) In: Defendi V (ed) Methodological Approaches to the Study of Leukemias, 69. The Wistar Institute Press, Philadelphia
Eski SE, Dubois C, Singh SP (2020) Nuclei isolation from whole tissue using a detergent and enzyme-free method. J vis Exp. https://doi.org/10.3791/61471
Fichtman B, Zagairy F, Biran N, Barsheshet Y, Chervinsky E, Ben Neriah Z, Shaag A, Assa M, Elpeleg O, Harel A, Spiegel R (2019) Combined loss of LAP1B and LAP1C results in an early onset multisystemic nuclear envelopathy. Nat Commun 10:605
Fischer AH, Taysavang P, Jhiang SM (2003) Nuclear envelope irregularity is induced by RET/PTC during interphase. Am J Pathol 163:1091–1100
Gauthier BR, Comaills V (2021) Nuclear envelope integrity in health and disease: consequences on genome instability and inflammation. Int J Mol Sci. https://doi.org/10.3390/ijms22147281
Gerasimenko OV, Gerasimenko JV, Tepikin AV, Petersen OH (1995) ATP-dependent accumulation and inositol trisphosphate- or cyclic ADP-ribose-mediated release of Ca2+ from the nuclear envelope. Cell 80:439–444
Gururaja Rao S, Patel NJ, Singh H (2020) Intracellular chloride channels: novel biomarkers in diseases. Front Physiol 11:96
Haider A, Wei Y-C, Lim K, Barbosa AD, Liu C-H, Weber U, Mlodzik M, Oras K, Collier S, Hussain MM, Dong L, Patel S, Alvarez-Guaita A, Saudek V, Jenkins BJ, Koulman A, Dymond MK, Hardie RC, Siniossoglou S, Savage DB (2018) PCYT1A regulates phosphatidylcholine homeostasis from the inner nuclear membrane in response to membrane stored curvature elastic stress. Dev Cell 45:481–495
Hampoelz B, Andres-Pons A, Kastritis P, Beck M (2019) Structure and assembly of the nuclear pore complex. Annu Rev Biophys 48:515–536
Hatch EM (2018) Nuclear envelope rupture: little holes, big openings. Curr Opin Cell Biol 52:66–72
Hatch E, Hetzer M (2014) Breaching the nuclear envelope in development and disease. J Cell Biol 205:133–141
Hwang S, Williams JF, Kneissig M, Lioudyno M, Rivera I, Helguera P, Busciglio J, Storchova Z, King MC, Torres EM (2019) Suppressing aneuploidy-associated phenotypes improves the fitness of Trisomy 21 cells. Cell Rep 29:2473-2488.e5
Jang SH, Byun JK, Jeon WIl, Choi SY, Park J, Lee BH, Yang JE, Park JB, O’Grady SM, Kim DY, Ryu PD, Joo SW, Lee SY (2015) Nuclear localization and functional characteristics of voltage-gated potassium channel Kv1.3. J Biol Chem 290:12547–12557
Kinugasa Y, Hirano Y, Sawai M, Ohno Y, Shindo T, Asakawa H, Chikashige Y, Shibata S, Kihara A, Haraguchi T, Hiraoka Y (2019) The very-long-chain fatty acid elongase Elo2 rescues lethal defects associated with loss of the nuclear barrier function in fission yeast cells. J Cell Sci 132:jcs229021
Kinugasa Y, Shibata S, Kihara A, Haraguchi T, Hiraoka Y (2019) The very-long chain fatty acid elongase Elo2 rescues lethal defects associated with loss of the nuclear barrier function in fission yeast cells. J Cell Sci 132:jcs229021
Kite GL (1913) The relative permeability of the surface and interior portions of the cytoplasm of animal and plant cells. Biol Bull 25:1–7
Kubben N, Misteli T (2017) Shared molecular and cellular mechanisms of premature ageing and ageing-associated diseases. Nat Rev Mol Cell Biol 18:595–609
LaCasse EC, Lefebvre YA (1995) Nuclear localization signals overlap DNA- or RNA-binding domains in nucleic acid-binding proteins. Nucleic Acids Res 23:1647–1656
Lattanzi G, Benedetti S, D’Apice MR, Maggi L, Carboni N, Scarano E, Politano L (2016) Emerging perspectives on laminopathies. Cell Health Cytoskelet 8:25–35
Lee YH, Tan HT, Chung MC (2010) Subcellular fractionation methods and strategies for proteomics. Proteomics 10:3935–3956
Lim S, Quinton RJ, Ganem NJ (2016) Nuclear envelope rupture drives genome instability in cancer. Mol Biol Cell 27:3210–3213
Lin DH, Hoelz A (2019) The structure of the nuclear pore complex (An Update). Annu Rev Biochem 88:725–783
Loewenstein WR, Kanno Y (1963) Some electrical properties of a nuclear membrane examined with a microelectrode. J Gen Physiol 46:1123–1140
Lusk CP, Blobel G, King MC (2007) Highway to the inner nuclear membrane: rules for the road. Nat Rev Mol Cell Biol 8:414–420
Malhas A, Goulbourne C, Vaux DJ (2011) The nucleoplasmic reticulum: form and function. Trends Cell Biol 21:362–373
Marschall M, Feichtinger S, Milbradt J (2011) Regulatory roles of protein kinases in cytomegalovirus replication. Adv Virus Res 80:69–101
Matzke AJM, Weiger TM, Matzke M (2010) Ion channels at the nucleus: electrophysiology meets the genome. Mol Plant 3:642–652
Mazzanti M, Bustamante JO, Oberleithner H (2001) Electrical dimension of the nuclear envelope. Physiol Rev Am Physiol Soc 81:1–19
McDuffie NG (1967) Nuclear blebs in human leukaemic cells. Nature 214:1341–1342
Meinke P, Schirmer EC (2016) The increasing relevance of nuclear envelope myopathies. Curr Opin Neurol 29:651–661
Méndez-López I, Worman HJ (2012) Inner nuclear membrane proteins: impact on human disease. Chromosoma 121:153–167
Milo R, Phillips R (2015) Cell biology by the numbers. Garland Science, New York
Mudumbi KC, Czapiewski R, Ruba A, Junod SL, Li Y, Luo W, Ngo C, Ospina V, Schirmer EC, Yang W (2020) Nucleoplasmic signals promote directed transmembrane protein import simultaneously via multiple channels of nuclear pores. Nat Commun 11:2184
Oliveira AG, Guimarães ES, Andrade LM, Menezes GB, Fatima Leite M (2014) Decoding calcium signaling across the nucleus. Physiology (bethesda) 29(5):361–368
Osorio DS, Gomes ER (2013) The contemporary nucleus: a trip down memory lane. Biol Cell 105:430–441
Ostlund C, Ellenberg J, Hallberg E, Lippincott-Schwartz J, Worman HJ (1999) Intracellular trafficking of emerin, the Emery-Dreifuss muscular dystrophy protein. J Cell Sci 112(Pt 11):1709–1719
Pascual-Garcia P, Capelson M (2019) Nuclear pores in genome architecture and enhancer function. Curr Opin Cell Biol 58:126–133
Pawar S, Kutay U (2021) The diverse cellular functions of inner nuclear membrane proteins. Cold Spring Harb Perspect Biol 13:a040477
Pirayesh N, Giridhar M, Ben Khedher A, Vothknecht UC, Chigri F (2021) Organellar calcium signaling in plants: an update. Biochim Biophys Acta Mol Cell Res 1868:118948
Raab M, Gentili M, de Belly H, Thiam H-R, Vargas P, Jimenez AJ, Lautenschlaeger F, Voituriez R, Lennon-Dumenil A-M, Manel N, Piel M (2016) ESCRT III repairs nuclear envelope ruptures during cell migration to limit DNA damage and cell death. Science 352:359–362
Romanauska A, Kohler A (2018) The inner nuclear membrane is a metabolically active territory that generates nuclear lipid droplets. Cell 174:700-715.e718
Rothballer A, Kutay U (2013) Poring over pores: nuclear pore complex insertion into the nuclear envelope. Trends Biochem Sci 38:292–301
Schindler M (1984) Alterations in nuclear anatomy by chemical modification of proteins in isolated rat liver nuclei. Exp Cell Res 150:84–96
Schirmer EC, Florens L, Guan T, Yates JR, Gerace L (2003) Nuclear membrane proteins with potential disease links found by subtractive proteomics. Science 301:1380–1382
Sebuwufu PH (1966) Nuclear blebs in the human foetal thymus. Nature 212:1382–1383
Secondo A, Petrozziello T, Tedeschi V, Boscia F, Pannaccione A, Molinaro P, Annunziato L (2020) Nuclear localization of NCX: Role in Ca 2+ handling and pathophysiological implications. Cell Calcium 86:102143
Selezneva A, Gibb AJ, Willis D (2022) The Nuclear envelope as a regulator of immune cell function. Front Immunol 13:840069
Shackleton S, Lloyd DJ, Jackson SN, Evans R, Niermeijer MF, Singh BM, Schmidt H, Brabant G, Kumar S, Durrington PN et al (2000) LMNA, encoding lamin A/C, is mutated in partial lipodystrophy. Nat Genet 24:153–156
Shah P, Wolf K, Lammerding J (2017) Bursting the bubble—nuclear envelope rupture as a path to genomic instability? Trends Cell Biol 27:546–555
Sosa BA, Rothballer A, Kutay U, Schwartz TU (2012) LINC complexes form by binding of three KASH peptides to domain interfaces of trimeric SUN proteins. Cell 149:1035–1047
Srivastava N, Nader GPF, Williart A, Rollin R, Cuvelier D, Lomakin A, Piel M (2021) Nuclear fragility, blaming the blebs. Curr Opin Cell Biol 70:100–108
Stiekema M, Houben F, Verheyen F, Borgers M, Menzel J, Meschkat M, van Zandvoort MAMJ, Ramaekers FCS, Broers JLV (2022) The role of lamins in the nucleoplasmic reticulum, a pleiomorphic organelle that enhances nucleo-cytoplasmic interplay. Front Cell Dev Biol 10:914286
Tange Y, Chikashige Y, Takahata S, Kawakami K, Higashi M, Mori C, Kojidani T, Hirano Y, Asakawa H, Murakami Y, Haraguchi T, Hiraoka Y (2016) Inner nuclear membrane protein Lem2 augments heterochromatin formation in response to nutritional conditions. Genes Cells 21:812–832
Tingey M, Mudumbi KC, Schirmer EC, Yang W (2019) Casting a wider net: differentiating between inner nuclear envelope and outer nuclear envelope transmembrane proteins. Int J Mol Sci 20:5248
Törö I, Oláh I (1966) Nuclear blebs in the cells of the guinea-pig thymus. Nature 212:315–317
Turner EM, Schlieker C (2016) Pelger-Huët anomaly and Greenberg skeletal dysplasia: LBR-associated diseases of cholesterol metabolism. Rare Dis 4:e1241363
Ueda N, Maekawa M, Matsui TS, Deguchi S, Takata T, Katahira J, Higashiyama S, Hieda M (2022) Inner nuclear membrane protein, SUN1, is required for cytoskeletal force generation and focal adhesion maturation. Front Cell Dev Biol 10:885859
Umbreit NT, Pellman D (2017) Cancer biology: genome jail-break triggers lockdown. Nature 550:340–341
Ungricht R, Kutay U (2017) Mechanisms and functions of nuclear envelope remodelling. Nat Rev Mol Cell Biol 18:229–245
van Meer G, Voelker DR, Feigenson GW (2008) Membrane lipids: where they are and how they behave. Nat Rev Mol Cell Biol 9:112–124
Wiggan O, Schroder B, Krapf D, Bamburg JR, Deluca JG (2017) Cofilin regulates nuclear architecture through a myosin-ii dependent mechanotransduction module. Sci Rep 7:40953
Wu W, Lin F, Worman HJ (2002) Intracellular trafficking of MAN1, an integral protein of the nuclear envelope inner membrane. J Cell Sci 115(Pt 7):1361–1371
Acknowledgements
This work was supported by Royal Physiographic Society of Lund, Åke Wibergs Stiftelse of Stockholm and Carl Tryggers Stiftelse of Stockholm.
Funding
Open access funding provided by Lund University.
Author information
Authors and Affiliations
Contributions
DA and AC: wrote the manuscript and prepared the figures.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
The authors do not have any conflict of interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Anand, D., Chaudhuri, A. Grease in the Nucleus: Insights into the Dynamic Life of Nuclear Membranes. J Membrane Biol 256, 137–145 (2023). https://doi.org/10.1007/s00232-022-00272-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00232-022-00272-8