Abstract
Population structure and phylogeography of the loggerhead sea turtle (Caretta caretta) have so far been assessed mainly by mitochondrial DNA (mtDNA) single-gene sequencing studies. However, phylogenetic relationships amongst matrilines, genetic characterisation of rookeries and mixed-stock analyses have suffered from the limited resolution obtained by comparison of relatively short sequences such as from the mtDNA control region. Whole mitogenome sequencing can significantly improve population genetics, particularly in marine organisms showing female natal philopatry. Despite mitogenomics becoming increasingly common in biodiversity monitoring and conservation, only a few complete mitogenomes are available for C. caretta. In this study, we sequenced the complete mtDNA of 61 loggerhead turtles sampled between 2008 and 2021 along the Italian coastline and central Mediterranean Sea. We assigned complete mtDNA haplotypes to dead embryos and bycatch samples, and introduced a first nomenclature for loggerhead mitogenomes. Analysis of mtDNA diversity, Maximum Parsimony and Bayesian phylogenetic reconstruction allowed improved resolution of lineages with respect to studies reporting on partial mtDNA control region sequence comparisons, and we were able to further inform previous analyses on loggerhead ancestry based on control region haplogroups. Overall, whole mitogenome analysis has potential for considerable improvement of evolutionary history and phylogeographic investigations as well as mixed-stock surveys of loggerhead turtles.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Female natal philopatry is the tendency for individual females to breed at or near their place of birth (Shields 1982). This behaviour can shape population structure in many marine species including whales, sharks and chelonids (Miller 1997; Sheridan et al. 2010; Baker et al. 2013; Feldheim et al. 2014). Each breeding site generally hosts specific aggregates of maternal descent. However, exceptions episodically occur which lead a female to colonise new sites and significantly increase the species’ range (e.g. Carreras et al. 2018).
Population genetic approaches to study female philopatry have mostly relied upon mitochondrial DNA (mtDNA), which is maternally inherited and can therefore trace matrilines within populations (Moritz 1994). In particular, investigations on whole mtDNA genome sequences have significantly improved our understanding of intraspecific phylogeographic patterns in several marine taxa. For instance, mitogenome sequence analysis in killer whales helped clarifying phylogenetic relationships amongst sympatric ecotypes which were then classified as different species (Morin et al. 2010). In speartooth shark and sawfish, mitogenome sequences comparison allowed to better resolve population structure and inform effective conservation measures for critically endangered populations (Feutry et al. 2014, 2015).
In sea turtles, female natal philopatry and population genetic structure have long been investigated through the analysis of partial mtDNA control region sequence diversity (Jensen et al. 2013). More recently, whole mtDNA investigations helped characterising Chelonia mydas matrilines which could not be resolved by comparison of control region haplotypes alone and allowed definition of fine-scale population structure in the critically endangered Lepidochelys kempii (Frandsen et al. 2020). For the loggerhead sea turtle (Caretta caretta), a few complete mitogenomes have been characterised worldwide and used to reconstruct interspecific phylogenies (Drosopoulou et al. 2012; Duchene et al. 2012; Hernández-Fernández and Delgado Cano 2018; Otálora et al. 2018). In particular, Drosopoulou et al. (2012) provided the first thorough description of the mtDNA genome of C. caretta using a sample collected from a rookery in western Greece, however, more detailed patterns of intraspecific mitogenomic variation were not investigated.
The loggerhead sea turtle can be found in tropical and subtropical seas as well as temperate waters of the Mediterranean basin, which hosts one of the 10 Regional Management Units defined for C. caretta to prioritise conservation efforts worldwide (Wallace et al. 2010). The most recent assessment of the IUCN Red List classifies the loggerhead turtles from the Mediterranean as a “least concern” Regional Management Unit. However, populations are conservation-dependent, severely fragmented and characterised by a continuous decline of mature individuals mainly because of habitat encroachment, pollution, fisheries bycatch, climate change and intentional take (Casale et al. 2018, 2020). A variation in the distribution of the species induced by global warming is already detectable (Maffucci et al. 2016; Carreras et al. 2018; Hochscheid et al. 2022) and advocated by projection modelling (Mancino et al. 2022). Conservation efforts can make use of a comprehensive understanding of several life history traits. In particular, the definition of maternal descent of individuals from foraging stocks and the genetic profiling of nesting females and new nesting sites are fundamental to guide management programmes for C. caretta in the Mediterranean Sea.
Genetic characterisation of Mediterranean rookeries and foraging stocks for the loggerhead has been so far conducted by comparison of partial mtDNA control region sequences (Garofalo et al. 2009; Yilmaz et al. 2011; Saied et al. 2012; Carreras et al. 2014; Clusa et al. 2014). This information was used in conservation-oriented studies where, for instance, mixed-stock analysis described the contributions from known rookeries to aggregations of turtles at sea (Garofalo et al. 2013; Clusa et al. 2014; Splendiani et al. 2017; Tolve et al. 2018; Loisier et al. 2021). To this regard, comparison of control region haplotypes has reached a resolution threshold which cannot allow any further sensible improvement in fine-scale population structure analysis as the main Mediterranean rookeries appear to all share the same control region haplotypes (e.g. Tolve et al. 2018). The very same molecular marker was also used to study patterns of loggerhead Mediterranean colonisation and radiation (e.g. Clusa et al. 2013; Novelletto et al. 2016; Splendiani et al. 2017; Loisier et al. 2021). Nevertheless, limited information could be obtained on the evolution of different lineages because of the scarce resolution amongst matrilines defined by mtDNA control region sequences.
In this study, we produced a first whole mtDNA genome dataset for loggerhead turtles from the Central Mediterranean Sea in order to enhance resolution of matrilines and characterise rookeries which could be not distinguished by comparison of ubiquitous control region haplotypes. We obtained high-coverage mitogenome sequences from dead embryos collected at nesting sites and from bycaught and stranded turtles. Eight out of 27 nests were located in minor or occasional nesting areas, which were genetically described for the first time. We assessed intraspecific genetic diversity and performed phylogenetic analysis to reconstruct evolutionary relationships amongst matrilines and the time frame of radiation of haplogroups and mtDNA haplotypes within haplogroups. Using additional, published loggerhead mtDNA genome sequences, we provided an overview of mitogenomic diversity for C. caretta in the Central Mediterranean. We argue that our findings will help supporting detailed inference in mixed-stock analysis and foster thorough investigation of the phylogeographic history of loggerhead turtles.
Materials and methods
Sampling, DNA isolation and sequencing by synthesis
A total of 61 tissue biopsies were obtained from loggerhead turtles sampled along the coastline of the Italian peninsula and the island of Linosa and in the sea waters between Sicily and Tunisia from 2008 to 2021 (Fig. 1). Samples were collected from stranded turtles in Veneto and Emilia Romagna (N = 4), Tuscany (N = 1) and Latium (N = 2) and from bycaught turtles in the Strait of Sicily (N = 27). Samples were also collected from dead embryos from four nests in Tuscany, two nests in Latium, 19 nests in Calabria and two nests on the island of Linosa (Table S1). Numbered tags were applied to bycaught turtles before release.
Map of the study area. A total of 61 samples of Caretta caretta were collected between 2008 and 2021 along the coastline of the Italian peninsula and the Island of Linosa and in the sea waters of the Strait of Sicily. Nesting sites (circles) and stranded turtles (black dots) were sampled in Veneto and Emilia Romagna (ADR), Tuscany (TUS), Latium (LAT), Calabria (CAL) and the Island of Linosa (LIN). The horizontal shaded region shows location of the bycatch area between Sicily and Tunisia
Each sample was preserved in a cryovial with 95% ethanol and subsequently stored at − 80 °C. Whole DNA was extracted following a standard phenol:chloroform:isoamyl alcohol procedure (Green and Sambrook 2012). Integrity of DNA was assessed by 1% agarose gel electrophoresis and DNA concentration was measured using a Qubit 4 fluorometer Broad Range Assay (Invitrogen). Mitochondrial DNA whole genome sequencing was performed by first generating dual-indexed (i5 and i7) genomic libraries using a PCR-free Tagmentation Kit and IDT for Illumina indexes set A (Illumina) according to the manufacturer’s protocol. We envisaged a 1 × target nuclear genome coverage for all samples in order to obtain a high depth coverage (> 200 ×) for the mitochondrial DNA, which is represented by a much higher number of copies per cell than nuclear DNA (Michaels et al. 1982; Robin and Wong 1988; Wiesner et al. 1992). Considering a nuclear DNA genome size of 2.13 Gb (Chang et al. 2023) and a mtDNA genome size of 16,737 bp for C. caretta (Drosopoulou et al. 2012), we sequenced the libraries paired-end on an Illumina NovaSeq 6000 System using a 300-cycle NovaSeq 6000 SP Reagent Kit v 1.5 for an expected yield of approximately 250 Gb.
Mitogenomes reconstruction and alignment
Demultiplexing and conversion of sequencing data from bcl to fastq format were performed using bcl2fastq version 2.20 (Illumina). FastQC 11.9 (Andrews 2010) was used to check for reads quality. Adaptors and low-quality reads were removed using Trimmomatic 0.39 (Bolger et al. 2014). Reads were mapped to the C. caretta complete annotated mitochondrial genome (GenBank accession number: FR694649.1) published by Drosopoulou et al. (2012) using Bowtie2-2.5.0 (Langmead and Salzberg 2012). This mitogenome was chosen as reference for it was the only one published for C. caretta from the Mediterranean basin. Sequence Alignment Map (SAM) files were compressed in Binary Alignment Map (BAM) format, sorted and indexed using Samtools 1.16.1 (Danecek et al. 2021). Variant calling was performed with BCFtools 1.16 (Danecek et al. 2021) and Tablet alignment viewer (Milne et al. 2013) was used to visually check variants and relative coverage. Consensus sequences were extracted from Variant Calling Format (VCF) files using BCFtools by filtering out variants in the first 10 bp and those with a QUAL value < 30.
Mitogenomes alignment was performed by including five additional loggerhead mtDNA genome sequences available in GenBank (see also Table S2): FR694649.1 (Greece; used as reference mitogenome), JX454983.1 (Florida), JX454988.1 (Hawaii), MF554690.1 and MF579505.1 (Colombia) (Drosopoulou et al. 2012; Duchene et al. 2012; Hernández-Fernández and Delgado Cano 2018). The MUSCLE algorithm was used to produce a multiple sequence alignment, which was trimmed at position 16,419, ahead of the (TATAT)n microsatellite repeat and annotated using Geneious Prime 2022.2. We then partitioned the alignment in (i) 12S and 16S rRNAs, (ii) tRNAs, (iii) the control region and (iv) coding genes. The final nucleotide dataset had no ambiguous positions and was numbered with reference to the FR694649.1 mitogenome (Drosopoulou et al. 2012). Genbank accession numbers are reported in Table S1 for each complete mtDNA haplotype sequence described in this study.
Mitogenomic haplotype nomenclature
Mitochondrial haplotypes were named following the criteria set by Shamblin et al. (2012a) and the guidelines of the Archie Carr Center for Sea Turtles Research (https://accstr.ufl.edu/resources/mtdna-sequences/). Complete mtDNA haplotypes were defined by three serial numbers. The first two numbers refer to the 815 bp long C. caretta control region haplotype code, whilst the third number refers to variants recorded in other genes of the mitogenome outside of the control region, so that complete mtDNA haplotype names are basically variants of the current control region haplotype nomenclature. For example, CC-A2.1.1, CC-A2.1.2 and CC-A2.1.3 all refer to complete mtDNA haplotype variants of the control region haplotype CC-A2.1.
Nomenclature for groups of complete mtDNA haplotypes also followed that of C. caretta control region variants, which identify three main haplotype clusters named haplogroups IA, IB, and II (Shamblin et al. 2014). Haplogroup IA includes haplotypes characterising Pacific Ocean nests, haplogroup IB includes haplotypes which are found in Atlantic rookeries and one single haplotype recorded in Indian Ocean nests (Shamblin et al. 2014). Finally, haplogroup II comprises haplotypes from both Atlantic and Mediterranean rookeries.
Mitogenomic diversity and phylogenetic analysis
Number of haplotypes, haplotypic diversity, number of variants, nucleotide diversity and GC content were computed using DnaSP (Librado and Rozas 2009) and the R package pegas v1.1 (Paradis 2010). Analysis was performed on whole mitogenome sequences and then single mitochondrial genes using the entire dataset, first and then on haplogroup II only. McDonald–Kreitman test was implemented using DnaSP to check for neutrality of substitutions in the coding regions (McDonald and Kreitman 1991). The nonsynonymous (dN) to synonymous (dS) substitution rate ratio amongst haplogroup II sequences was then compared to the dN/dS ratio between haplogroup II sequences and mitogenomes of haplogroup IB (obtained by this study and from GenBank, accession number: MF579505.1). In order to rule out parallel patterns of nucleotide variations between haplogroup II and IB, the dN/dS ratio amongst haplogroup II variants was measured against the dN/dS between haplogroup II and the mitogenome of Lepidochelys olivacea as an outgroup (GenBank accession number: JX454987.1). Finally, differences in dN/dS amongst mitogenomes of the entire C. caretta dataset of our study were compared to dN/dS between our dataset and the mtDNA genome of L. olivacea.
Two haplotype networks were constructed using statistical parsimony (Templeton et al. 1992). The method, implemented in the TCS programme (Clement et al. 2000), links haplotypes with the smaller number of differences as defined by a 95% confidence criterion. The first network was based on 815 bp control region haplotypes only, whilst the second one was a network of complete mitogenome haplotypes.
A Maximum Parsimony (MP) graph based on the entire dataset was also constructed using MEGA11 (Kumar et al. 2018) with the “use all sites” option in order to visualise similarities between sequences and polarise mutational changes. We then applied a Bayesian divergence time analysis to provide a time frame for the process of radiation of haplogroup II with respect to the other two major C. caretta mitochondrial haplogroups IA and IB. Phylogeny and divergence time analyses provided poor resolution when using L. olivacea as outgroup. We therefore conducted MP and Bayesian inference of phylogeny using haplogroup IB as reference in order to better resolve intraspecific topology of haplogroup II, which included the vast majority of haplotypes defined in our study. We used BEAST suite v.1.10.4 (Suchard et al. 2018) on four mitogenome partitions (12S and 16S rRNAs, tRNAs, the control region and coding genes). We performed 20 independent test-runs in order to optimise priors for chain convergence and defined, for each partition, the “unlink” option flagged for substitution model and for molecular clock and the “link” option flagged for generation of the tree. The HKY (Hasegawa et al. 1985) substitution model and a strict molecular clock were selected for each partition. The “unlink parameters on codon positions” option was flagged for the fourth partition. The HKY model was selected because the tests conducted using more complex substitution models gave a poor Effective Sample Size (ESS), possibly due to over-parametrization. Likewise, we chose a strict molecular clock because multiple tests conducted using more complex uncorrelated relaxed clocks resulted in poor ESS and zero values for the parameters uced.mean and meanRate, which indicate a clock-like behaviour of the data. No prior was set on the time to the most recent common ancestor (TMRCA) and no divergence time estimate was assessed for the root of the tree. We converted the substitutions value from (site × million year)−1 to (site × generation)−1, where the generation time is calculated as age of maturity plus half of reproductive longevity (Pianka 1973). We used 25 years as age of maturity and 22 years as reproductive longevity which resulted in a 36-year generation time (Margaritoulis et al. 2020; Baldi et al. 2023). We set a normal distribution (mean = 1.2E-7, S.D. = 4.0E-8) as prior on the substitution rates for each mitogenomic partition. The distribution was truncated with a lower bound equal to 0.0 and an upper bound of 2.41E-7 (corresponding to 6.7E-3 million year−1), which is the highest value for mitochondrial substitution rates recorded in marine turtles (Dutton et al. 1999). Chain length was set to 10E8 steps with a 10% initial burn-in. We monitored the ESS for each output parameter after each run using Tracer v.1.7.1 (Rambaut et al. 2018). The overall BEAST output was summarised using TreeAnnotator (Drummond and Rambaut 2007) and the statistics related to the consensus tree nodes including node height, 95% Height Posterior Density (HPD) and posterior, were assessed with FigTree v1.4.4 (http://tree.bio.ed.ac.uk/software/figtree/).
Results
Mitogenomic diversity and complete mtDNA haplotype assignment
Whole mtDNA sequences were obtained for all 61 samples with an average genome coverage ( ×) of 272.4 ± 37.1SE (see Table S3). Mitogenome sequences were trimmed at position 16,425 so that the (TATAT)n microsatellite repeat positioned at the 3’ end of the control region was discarded from the analysis.
We recorded two mtDNA sequences in sample SIC04. We reported this pattern as a possible occurrence of heteroplasmy and assigned to SIC04 the mitogenomic haplotype name of the mtDNA variant (G > A at position 4671 of the genome) that was confirmed in 70% of the reads (Table S1).
All samples belonged to haplogroup II apart from the stranded adult TUS05, of unknown origin, which was assigned to haplogroup IB, control region haplotype CC-A1.1. We named this mitogenomic haplotype CC-A1.1.1. Downloaded mitogenome sequences FR694649.1 (our reference genome), JX454983.1 and MF554690.1 were assigned to haplogroup II, while MF579505.1 and JX454988.1 were assigned to haplogroups IB and IA, respectively (Table S2).
Haplogroup II sequence variants and their distribution across sampling sites are reported in Tables 1 and 2, respectively. The loggerhead mitogenomes characterised in this study showed a total of 27 polymorphic sites with respect to the C. caretta reference mtDNA FR694649.1. Of these, two segregating sites were recorded across all 61 samples of our dataset and in particular an indel in position 1631 in the 16S rRNA gene and a T > C substitution in position 9974 in the ND4L gene (Table1). Mitogenomic haplotype CC-A2.1.1 was the most abundant both in embryos and adult turtles (found in 26 out of 61 samples). The second most abundant haplotype was CC-A2.1.8, found in six samples from the Calabrian nests, one sample from the Island of Linosa and three samples collected from bycaught adults in the Strait of Sicily. Seven embryos from different nests in Calabria carried mitogenomic haplotype CC-A20.1.1. This mitogenome was not found elsewhere in the study area. Haplotype CC-A2.1.3 was found in three specimens in Tuscany and southern Italy. All the other mitogenomic haplotypes were represented by either one or two individuals. The nests from Tuscany, Latium and Linosa Island were characterised by haplotypes CC-A2.1.1 (TUS01, TUS02, LAT01, LAT02), CC-A2.1.3 (TUS03), CC-A2.1.9 (TUS04), CC-A2.1.8 (LIN01) and CC-A2.9.2 (LIN02) (Table S1).
Overall, we characterised 286 variable sites with respect to 55 polymorphisms recorded by comparing control region sequences alone and assigned samples to 23 different mitogenomic haplotypes. Ten haplotypes only were defined by control region sequence analysis (Table 3). Within haplogroup II, mitogenome sequences identified 27 variable sites and 20 haplotypes compared to six variable sites and seven haplotypes recorded by control region sequence comparison (Table 3; Fig. 2).
Genetic diversity analysis of whole mtDNA revealed a much higher number of haplotypes and higher haplotypic and nucleotide diversity than values recorded by control region sequences comparison (Table 3). Amongst the 16 gene partitions of the C. caretta mitogenome, the ND5 gene showed the highest number of segregating sites and haplotypic diversity (Tables S4 and S5). The same pattern was recovered by the analysis of haplogroup II mitogenomes for the high abundance (28%) of mitogenomic haplotypes CC-A2.1.8 and CC-A2.1.20 carrying the variant T at site 13,373 of the ND5 gene. Overall, nucleotide diversity (π) showed a threefold variation along the mtDNA molecule, with peak values at positions 3000–5000, 9000 to 10,000 and 13,000 to 14,000 (Fig. S1). When haplogroup II sequences only were considered, π values were the highest at positions 9000–10,000 and 13,000–14,000. The NADH dehydrogenase and ATP synthase subunit genes displayed the highest π values. Conversely, analysis restricted to haplogroup II mitogenomic haplotypes showed the highest π values for the ATP8 gene only (Tables S4 and S5).
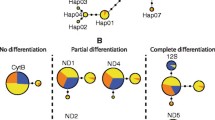
The TCS haplotype networks clustered both the partial mtDNA control region sequences and the mitogenomes in three unlinked haplogroups with 95% probability (Fig. 2). Links between haplogroups were obtained with a lower probability threshold by fixing connexion limits at 100 and 200 steps for control region and mitogenome haplotypes, respectively. Two internal ambiguities in the mitogenome cladogram were resolved by applying criteria derived from empirical validation of the coalescent theory (Crandall and Templeton 1993; Templeton and Sing 1993; Fetzner and Crandall 2003). Accordingly, a haplotype connexion was maintained to high-frequency haplotypes with an interior position in the network (Castelloe and Templeton 1994) rather than to low-frequency ones located in tip clades. Internal ambiguities could not be resolved for the mtDNA control region haplotypes network. We therefore maintained the three haplogroups unlinked. Haplogroup IA was represented by a single sequence in both reconstructions, while haplogroup IB included two haplotypes separated by 12 and 30 possible unobserved intermediate haplotypes in the partial mtDNA control region and mitogenomic network, respectively. The haplogroup II network based on control region sequences had a star-like configuration with a core CC-A2.1 haplotype and six equidistant derived haplotypes. A similar but more complex pattern was described by the mitogenomic analysis. Haplogroup II showed 10 equidistant haplotypes deriving from CC-A2.1.1, two haplotypes (CC-A2.1.10 and CC-A20.1.1) deriving from CC-A2.1.3 and CC-A2.1.8, respectively and five additional branches with haplotypes linked to CC-A2.1.1 by unobserved intermediate haplotype states.
Variation in the protein-coding regions
The McDonald-Kreitman neutrality test returned no significant values when comparing dN/dS of haplogroup II with dN/dS between haplogroup II and either haplogroup IB or the mitogenome of L. olivacea (Table S4). We found an excess of synonymous substitutions in all genes when comparing haplogroup II with haplogroup IB sequences and a similar result was obtained when comparing haplogroup II variants to the mitogenome of L. olivacea. Ten variants in the coding regions of haplogroup II sequences were synonymous and six variants were non-synonymous. In particular, ND2, ND3 and CYTB genes displayed no synonymous substitutions and a non-synonymous one. Comparison of dN/dS of our complete C. caretta dataset with dN/dS between C. caretta mitogenomes and L. olivacea mtDNA revealed an excess of non-synonymous substitutions for ND1, ND2, ATP6, ND3, ND5 and CYTB genes. Nevertheless, a significant value was shown for the ND3 gene only.
Phylogenetic analyses
The phylogenetic tree inferred by Maximum Parsimony resolved equally parsimonious topologies whereby haplogroup II (Atlantic and Mediterranean) and haplogroup IA (Pacific) shared the same cluster, while haplogroup IB (Atlantic) included the most phylogenetically divergent sequences (Fig. 3). All haplogroup II sequences coalesced to a long branch defined by 55 substitutions. Of these, 18 substitutions occurred in the control region, three were found in rRNA genes, one in a tRNA and 33 in coding genes with 28 synonymous and five non-synonymous substitutions (Table S6).
Unrooted maximum parsimony (MP) tree of Caretta caretta obtained using 61 mitogenomes from this study and five sequences from published literature (left). Haplogroup IA (Pacific Ocean) corresponds to Genbank accession number JX454988.1. Haplogroup IB (Atlantic Ocean) includes Genbank accession number MF579505.1 and mitogenomic haplotype TUS05 from this study (see Tables S1 and S2). Details of the haplogroup II subtree (right) show phylogenetic relationships amongst mtDNA sequences of 60 samples from this study (Mediterranean Sea) and three sequences from published literature (Mediterranean Sea and Atlantic Ocean). Numbers on tree branches represent number of nucleotide substitutions
Two main clades were defined within haplogroup II (Fig. 3). The clade characterised by the rare Mediterranean control region haplotype CC-A2.9 differed from the other haplogroup II sequences by the control region variant C at position 15,562, the derived state C > T at position 8398 in the ATP6 gene and the ancestral state A at position 9440 in the COIII gene. Mitogenome analysis further distinguished mitogenomic haplotypes CC-A2.9.1 and CC-A2.9.2 as a result of a A/G variant at position 5303 in the Cys tRNA gene (Fig. 3, Table S7). The second and most conspicuous clade was defined by a basal A > T transversion at position 9440 and included all CC-A2.1 mitogenomic variants as well as haplotypes corresponding to control region haplotypes CC-A3.1, CC-A10.4, CC-A20.1 and CC-A43.1.
Of the CC-A2.1 mitogenomic variants, six of them (CC-A2.1.2 to CC-A2.1.7 and CC-A2.1.9) derived each by one private variant from the basal CC-A2.1.1 mitogenomic haplotype. Similarly, haplotypes CC-A3.1.1 and CC-A10.4.1 each differed from CC-A2.1.1 by a single control region variant. Mitogenomic haplotype CC-A2.1.10 stemmed from CC-A2.1.3 by a synonymous substitution in the ND1 gene, while haplotype CC-A2.1.11 differed from the basal CC-A2.1.1 by two variants. Mitogenomic haplotype CC-A43.1.1 was the most divergent sequence characterised by two non-synonymous, two synonymous substitutions and the control region variant. Haplotype CC-A20.1.1 derived from CC-A2.1.8, which in turn stemmed from the basal clade by a C > T variant in position 13,373 in the ND5 gene (Table S7). All CC-A20.1 sequences were identical, denoting a recent common origin. Finally, the Atlantic mitogenomes JX454983.1 and MF554690.1 derived from CC-A2.1.1 by a single synonymous substitution and from the Mediterranean reference FR694649.1, respectively (Fig. 3, Tables S6 and S7).
The Bayesian tree confirmed the branching order of the haplogroups recovered by the MP tree (Fig. 4 and Table 4). The root of the tree (node I) separated haplogroup IB from haplogroups IA and II. The second node, positioned at 67% of the height of the total tree, divided haplogroup IA from haplogroup II mitogenomic haplotypes. According to Bayesian phylogeny, radiation of haplogroup IB (node III) occurred well before the branching of haplogroup II, which occupies only 4% of the total tree height. The Bayesian analysis also confirmed the basal position of the CC-A2.9 clade within haplogroup II. The only discrepancy between MP phylogeny and Bayesian reconstruction was the position of mitogenomic haplotype CC-A43.1.1, which resulted basal, instead of derived, with respect to CC-A2.1 variants. Other nodes that reached strong support (> 95% posterior probability) were those bifurcating CC-A20.1.1 and CC-A2.1.8, CC-A2.9.1 and CC-A2.9.2, CC-A2.1.3 and CC-A2.1.10, MF554690.1 and FR694649.1. All these nodes confirmed the phylogenetic clustering of the MP tree. The radiation of these mitogenomic haplotypes resulted to be a very recent event for they occupied just 1% of the total tree height. In particular, the most recently diverged mitogenomic sequences were FR694649.1 and MF554690.1, sampled in Greece and in the Atlantic Ocean, respectively (Table 4). Substitution rate estimates obtained for each mitogenome partition are reported in Table S8. The highest substitution rate (number of substitutions × (site × million year)−1) was recorded for the control region. The coding regions had a substitution rate three times lower than the control region while the rRNAs’ partition showed the lowest values.
Bayesian tree of Caretta caretta based on 61 mitogenomes from this study and five sequences from published literature. Nodes supporting > 95% posterior probability are shown in red. BEAST coerces multifurcations of identical sequences, so that nodes separating sequences assigned to the same mitogenomic haplotype should not be considered. Numbers on tree branches represent number of generations. Sample names and haplotypes assignment as for Fig. 1 and Table S1, respectively
Discussion
Improved resolution in loggerhead mitochondrial genetic diversity
We characterised the mitochondrial genomes of 61 loggerhead turtles sampled between 2008 and 2021 along the coastline of Italy, on the Island of Linosa and in sea waters of the Strait of Sicily and provided a first dataset on whole mtDNA sequences for Mediterranean C. caretta populations. Mitogenome analysis allowed an improved resolution of loggerhead matrilines with respect to previous studies based on partial mtDNA control region sequences comparison. The TCS haplotype networks based on statistical parsimony delineated similar star-like topologies in haplogroup II for both partial mtDNA control region and complete mtDNA analysis. However, statistical parsimony showed the increased resolution of the mitogenome haplotype network with respect to the 815 bp control region reconstruction, particularly for haplotype CC-A2.1.
The control region haplotype CC-A2.1 is the most abundant amongst the 13 main rookeries so far described in the Mediterranean Sea and it is also widespread in Atlantic nesting sites (Garofalo et al. 2009; Yilmaz et al. 2011; Saied et al. 2012; Carreras et al. 2014; Clusa et al. 2014). The ubiquity of haplotype CC-A2.1 has often hampered the characterisation of rookeries for turtles of unknown origin. In particular, mixed-stock analyses appeared to suffer from large confidence intervals associated to estimates of contribution by rookeries to specific mixed stocks. Confidence intervals with a lower value equal to zero most probably underestimated contribution of minor but perhaps genetically important rookeries (Garofalo et al. 2013; Clusa et al. 2014; Splendiani et al. 2017; Tolve et al. 2018; Loisier et al. 2021). In our study, 52 individual turtles all shared the same control region haplotype CC-A2.1 whilst whole mitogenome analysis recovered 11 distinct mitogenomic haplotypes (CC-A2.1.1-CC-A2.1.11), which significantly improved genetic characterisation of geographically distinct nesting sites. While the CC-A2.1.1 variant resulted the most abundant in our dataset with 28 records out of a total of 61 samples, characterisation of the other 10 mitogenomic haplotypes enabled distinction of nesting sites from approximately 30 km down to only 5 km apart along the coastline of Tuscany, Latium and Calabria. These nests could not be distinguished by mtDNA control region sequences analysis alone.
Similarly, haplotype CC-A1.1, which was carried by TUS05 and never found in Mediterranean rookeries, is very abundant and widespread in the Atlantic and therefore poorly informative of turtles’ provenance (Shamblin et al. 2012b). It is well known that CC-A1.1 turtles enter the western Mediterranean to forage and then migrate back to the Atlantic for reproduction (Bolten et al. 1998; Laurent et al. 1998; Carreras et al. 2006, 2011). It may therefore be difficult to envisage a Mediterranean origin for TUS05. Nevertheless, future mtDNA characterisation for Atlantic and Mediterranean loggerheads may unveil a number of mitogenomic variants of CC-A1.1 and allow description of differences in mtDNA genome frequencies across rookeries to improve resolution of turtles’ origin.
Our mitogenomic approach also improved the resolution of matrilines based on mtDNA control region haplotype CC-A2.9. This is a rare Mediterranean haplotype so far recorded only in Libya and in a small Israeli rookery. The two mitogenomic haplotypes CC-A2.9.1 and CC-A2.9.2 were found in a stranded adult in the northern Adriatic Sea and most important in one nest on the Island of Linosa, respectively. This is also the first evidence of a female carrying the rare haplotype nesting in a rookery other than Libya or Israel.
Mitogenomes comparison showed that in Calabrian nests there are at least five different mitogenomic haplotypes instead of only two variants (CC-A2.1 and CC-A20.1) detected by the analysis of mtDNA control region sequences. Of these, haplotype CC-A20.1.1 remains exclusive to the Calabrian rookery as for previous studies reporting on mtDNA control region haplotype CC-A20.1. However, we did not record mitogenomes characterised by control region haplotype CC-A31.1, also private to Calabrian nests (Garofalo et al. 2009; Yilmaz et al. 2011; Saied et al. 2012; Clusa et al. 2014).
Genetic characterisation of occasional and minor nesting sites
Major Mediterranean loggerhead rookeries are found in the eastern and southern sections of the basin from Greece through Turkey, Cyprus, Lebanon, Israel to Libya (Yilmaz et al. 2011; Saied et al. 2012; Carreras et al. 2014; Clusa et al. 2014). The western and central parts of the Mediterranean have been so far home of occasional nests and it was only in the last decade that nesting events were recorded on a more frequent and regular basis (Casale et al. 2018). According to Hochscheid et al. (2022), the number of turtle nests west of the main rookeries has increased from three nests per year in 2013 to 84 nests recorded in 2020, suggesting a 1300 km north-westward range expansion of loggerhead nesting distribution (Mancino et al. 2022). Samples collected from nests in Tuscany and Latium are evidence of such latitudinal dispersal, which seems now possible also as a result of a steady increase in water temperature (Casale et al. 2018; Hochscheid et al. 2022). All four nests recovered in Tuscany and the two nests from Latium had the same control region haplotype CC-A2.1, a highly shared sequence across both the Mediterranean basin and the Atlantic. On the other hand, analysis of mitogenome sequences recovered the presence of at least three distinct nesting females in this region of the northern Tyrrhenian Sea. Of these, a female that laid eggs in southern Tuscany (TUS03, see Table S1) carried mitogenomic haplotype CC-A2.1.3, which was also found in embryos from a nest on the Ionian coast of the Calabrian rookery (CAL1303). This pattern may suggest a southern Italian origin of the occasional nests recorded in Tuscany possibly as a result of the ongoing northward expansion of the distribution range of loggerheads in the Mediterranean (Mancino et al. 2022). As much as our evidence is based on a single observation, it nevertheless provides valuable information to further investigate dispersal of pioneer female loggerheads and the propagation of specific matrilines for the establishment of new rookeries.
In the Pelagian archipelago, Mingozzi et al. (2006) recorded nesting events on the Islands of Lampedusa and Linosa between 1985 and 1999 and regular, but not annual, nesting activities from 2000 to 2004. Nevertheless, genetic data was not produced during these surveys, so that ours represents the first study characterising the mitochondrial signature of the region. The two nests sampled in 2006 and 2008 were assigned to different matrilines, suggesting a relatively high degree of genetic diversity, given the limited size (540 ha) of Linosa Island. This is nevertheless coherent with the mitogenomic analysis of samples collected in the nearby Strait of Sicily where 11 mitogenomic haplotypes were recorded in 27 bycaught adult loggerheads. Additional surveys should assess the relevance of Linosa island as an additional nesting area for female loggerheads carrying a rare CC-A2.9.2 haplotype, considering that the CC-A2.9 matriline had so far been recorded in Libyan and Israeli nests only and in several individuals foraging in Mediterranean waters (e.g. Garofalo et al. 2013; Splendiani et al. 2017; Tolve et al. 2018). Moreover, the higher resolution offered by a mitogenomic approach could allow analysis of the genetic diversity and genomic differentiation between Linosa and the Island of Lampedusa, for which genetic characterisation of nesting sites is yet to be provided.
Heteroplasmy occurrence and patterns of substitutions in the coding regions
The deep sequencing coverage we obtained for C. caretta mitochondrial genomes revealed the presence of two mtDNA variants in one individual bycaught in the Strait of Sicily (SIC04). Heteroplasmy consisted of the coexistence of the CC-A2.1.1 mitogenomic haplotype with a mitogenome sequence carrying a truncating variant at mtDNA position 4671 on the ND2 gene, turning a tryptophan codon into a stop codon. The tryptophan codon located in the ND2 gene is a well-conserved sequence across vertebrates (Avise 1986) and the presence of a termination codon instead of a coding sequence during translation would suggest a possible detrimental effect on gene functioning (Yan et al. 2019). Although a mitogenome carrying such variant may not persist in homoplasy, we assigned a distinct mitogenomic haplotype name (CC-A2.1.2) to specimen SIC04. Tikochinski et al (2020) argued how intra-individual heteroplasmy may not be such a rare occurrence in sea turtles and could instead play a role in the evolution of population genetic diversity across generations. Unfortunately, the limitations of Sanger sequencing in the characterisation of low-frequency variants have impeded the development of studies on heteroplasmy in sea turtles. Nevertheless, regular use of high-throughput sequencing techniques should now lead to an increase in studies investigating this phenomenon and its evolutionary implications.
We found the highest nucleotide diversity in NADH dehydrogenase subunits genes. A similar pattern was previously recorded for genes encoding the ND1 and ND3 subunits (Novelletto et al. 2016). In particular, the ND5 subunit gene showed a relatively high proportion of polymorphism and the highest haplotype diversity. We also found a high nucleotide diversity in the genes encoding the subunits of ATP synthases across mitogenomes of haplogroup II, suggesting a pattern of sequence radiation within this clade. However, we did not record a significant excess of amino acid substitutions with respect to synonymous substitutions in haplogroup II mitogenomes. This would argue against adaptive selection acting on haplogroup II.
Mediterranean loggerhead phylogeography
The topologies of both Maximum Parsimony and Bayesian trees showed a marked star-like phylogeny of the main clade within haplogroup II, suggesting a fast and recent radiation of mitogenomic haplotypes from CC-A2.1.1 to CC-A2.1.10, CC-A3.1.1, CC-A10.4.1, CC-A20.1.1 and CC-A43.1.1. On the other hand, the sister branch defined by haplotypes CC-A2.9.1 and CC-A2.9.2 was basal in our phylogenetic reconstruction. A similar pattern was found for the CC-A2.9 mtDNA control region haplotype (Novelletto et al. 2016; Splendiani et al. 2017; Loisier et al. 2021), which was until now considered to be exclusive to Libyan and Israeli rookeries. This pattern supports the hypothesis of Clusa et al. (2013) by which the first loggerhead colonies of the Mediterranean settled in Libya during the Pleistocene, approximately 65 KYBP. The southern Mediterranean was probably a glacial refugium from where loggerhead turtles would have dispersed after the Last Glacial Maximum (LGM). Although our dataset only partially represents the Mediterranean mtDNA diversity, the time estimate we obtained for the divergence of the CC-A2.9 haplotype clade was consistent with a Pleistocenic colonisation of Libya. Our phylogeny did not show additional matrilines deriving from mtDNA carrying the control region haplotype CC-A2.9, which was previously recorded to have a close phylogenetic relationship with haplotypes CC-A26.1 and CC-A66.1 (Novelletto et al. 2016). A more comprehensive sampling including Ionian and eastern Mediterranean rookeries may reveal additional mitogenomic variants of the CC-A2.9 clade, which has so far been reported to have a limited radiation with respect to the sister branch CC-A2.1 (Garofalo et al. 2009; Yilmaz et al. 2011; Saied et al. 2012; Carreras et al. 2014; Clusa et al. 2014).
A more recent, second colonisation event of the Mediterranean from the Atlantic during the Holocene was suggested by Clusa et al. (2013) to explain the origin and expansion of the Calabrian rookery in southern Italy. This hypothesis was based on the presence of mtDNA control region haplotype CC-A20.1 in both Calabria and three rookeries in Florida (Shamblin et al. 2012b). The mitogenomic haplotype CC-A20.1.1 described in our study shared a recent ancestor with CC-A2.1.8 and both were recorded only in the southern part of our study area including Calabria, Linosa Island and the Strait of Sicily. This may suggest a local origin of these matrilines after the end of the LGM. Comparison of whole mitogenomic sequences from both Mediterranean and Atlantic stocks will be crucial to help understand origin and dispersal patterns of the CC-A20.1 clade.
On a wider, trans-oceanic scale, a number of biogeographical scenarios have been suggested advocating a first colonisation of the Mediterranean Sea by haplogroup II from Atlantic rookeries (Shamblin et al. 2014) or through a South African route (Bowen et al. 1994; Baltazar-Soares et al. 2020). Our topology recovered by both maximum parsimony and Bayesian analysis provided preliminary information on mitogenome shared ancestry. We envisage that further analysis based on e.g. an Approximate Bayesian Computation approach (see Baltazar-Soares et al. 2020) that makes use of whole mtDNA sequences from a wider sample set including individuals from the Atlantic, could help better elaborate current hypotheses and build a robust phylogeographic scenario of colonisation and dispersal of loggerheads in the Mediterranean Sea.
Conclusions
Analysis of complete mitogenome sequences can improve population genetic and phylogeographic studies, particularly for marine species that exhibit female natal philopatry. In this study, we investigated patterns of mtDNA diversity in loggerhead sea turtles and presented a high-coverage, complete mitogenome dataset for C. caretta in the central Mediterranean Sea along with a first genetic description of the Italian nesting sites of Tuscany, Latium and the Island of Linosa. We also proposed the first nomenclature for C. caretta mitogenomic haplotypes and described a larger number of variants than those so far recovered by mtDNA control region sequence comparisons. Mitogenome analysis provided a more comprehensive definition of matriline phylogeny and improved genetic characterisation of rookeries. Our study represents a first example of the potential of whole mitogenome analysis for the study of the evolutionary history of Mediterranean loggerheads and fine-scale characterisation of nesting sites to help mixed-stock analysis and the conservation of extant rookeries.
Data availability
Mitochondrial DNA haplotype sequences presented in this study were deposited in the GenBank database (accession numbers: OR166582 - OR166647).
References
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc
Avise JC (1986) Mitochondrial DNA and the evolutionary genetics of higher animals. Philos Trans R Soc Lond B 312:325–342. https://doi.org/10.1098/rstb.1986.0011
Baker CS, Steel D, Calambokidis J, Falcone E, González-Peral U, Barlow J, Burdin AM, Clapham PJ, Ford JK, Gabriele CM (2013) Strong maternal fidelity and natal philopatry shape genetic structure in North Pacific humpback whales. Mar Ecol Prog Ser 494:291–306. https://doi.org/10.3354/meps10508
Baldi G, Furii G, Del Vecchio M, Salvemini P, Vallini C, Angelini V, Pari S, Lombardi Moraes K, Margaritoulis D, Çurri A, Lazar B, Casale P (2023) Growth rates and age at maturity of Mediterranean loggerhead sea turtles estimated from a single-population foraging ground. Mar Biol 170:36. https://doi.org/10.1007/s00227-023-04189-1
Baltazar-Soares M, Klein JD, Correia SM, Reischig T, Taxonera A, Roque SM, Dos Passos L, Durão J, Lomba JP, Dinis H, Cameron SJK, Stiebens VA, Eizaguirre C (2020) Distribution of genetic diversity reveals colonization patterns and philopatry of the loggerhead sea turtles across geographic scales. Sci Rep 10:18001. https://doi.org/10.1038/s41598-020-74141-6
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120. https://doi.org/10.1093/bioinformatics/btu170
Bolten AB, Bjorndal KA, Martins HR, Dellinger T, Biscoito MJ, Encalada SE, Bowen BW (1998) Transatlantic developmental migrations of loggerhead sea turtles demonstrated by mtDNA sequence analysis. Ecol Appl 8:1–7. https://doi.org/10.1890/1051-0761(1998)008[0001:TDMOLS]2.0.CO;2
Bowen BW, Kamezaki N, Limpus CJ, Hughes GR, Meylan AB, Avise JC (1994) Global phylogeography of the loggerhead turtle (Caretta caretta) as indicated by mitochondrial DNA haplotypes. Evolution 48:1820–1828. https://doi.org/10.1111/j.1558-5646.1994.tb02217.x
Carreras C, Pont S, Maffucci F, Pascual M, Barceló A, Bentivegna F, Cardona L, Alegre F, SanFélix M, Fernández G, Aguilar A (2006) Genetic structuring of immature loggerhead sea turtles (Caretta caretta) in the Mediterranean Sea reflects water circulation patterns. Mar Biol 149:1269–1279. https://doi.org/10.1007/s00227-006-0282-8
Carreras C, Pascual M, Cardona L, Marco A, Bellido JJ, Castillo JJ, Tomás J, Raga JA, Sanfélix M, Fernández G, Aguilar A (2011) Living together but remaining apart: Atlantic and Mediterranean loggerhead sea turtles (Caretta caretta) in shared feeding grounds. J Hered 102:666–677. https://doi.org/10.1093/jhered/esr089
Carreras C, Pascual M, Tomas J (2018) Sporadic nesting reveals long distance colonisation in the philopatric loggerhead sea turtle (Caretta caretta). Sci Rep 8:1435. https://doi.org/10.1038/s41598-018-19887-w
Carreras C, Rees AF, Broderick AC, Godley BJ, Margaritoulis D (2014) Mitochondrial DNA markers of loggerhead marine turtles (Caretta caretta) (Testudines: Cheloniidae) nesting at Kyparissia Bay, Greece, confirm the western Greece unit and regional structuring. Sci Mar 78: 115–124. https://doi.org/10.3989/scimar.03865.27B
Casale P, Broderick AC, Camiñas JA, Cardona L, Carreras C, Demetropoulos A, Fuller WJ, Godley BJ, Hochscheid S, Kaska Y (2018) Mediterranean sea turtles: current knowledge and priorities for conservation and research. Endanger Species Res 36:229–267. https://doi.org/10.3354/esr00901
Casale P, Hochscheid S, Kaska Y, Panagopoulou A (2020) Sea Turtles in the Mediterranean Region: MTSG Annual Regional Report 2020. Report of the IUCN-SSC Marine Turtle Specialist Group
Castelloe J, Templeton AR (1994) Root probabilities for intraspecific gene trees under neutral coalescent theory. Mol Phylogenet Evol 3:102–113. https://doi.org/10.1006/mpev.1994.1013
Chang G, Jones S, Leelakumari S, Ashkani J, Culibrk L, O'Neill K, Tse K, Cheng D, Chuah E, McDonald H, Kirk H, Pandoh P, Pari S, Angelini V, Kyle C, Bertorelle G, Zhao Y, Mungall A, Moore R, Vilaça ST, Jones S (2023) The genome sequence of the Loggerhead sea turtle, Caretta caretta Linnaeus 1758 [version 2; peer review: 2 approved]. F1000Research 12. https://doi.org/10.12688/f1000research.131283.2
Clement M, Posada D, Crandall KA (2000) TCS: a computer program to estimate gene genealogies. Mol Ecol 9:1657–1659. https://doi.org/10.1046/j.1365-294x.2000.01020.x
Clusa M, Carreras C, Pascual M, Demetropoulos A, Margaritoulis D, Rees AF, Hamza AA, Khalil M, Aureggi M, Levy Y (2013) Mitochondrial DNA reveals Pleistocenic colonisation of the Mediterranean by loggerhead turtles (Caretta caretta). J Exp Mar Biol Ecol 439:15–24. https://doi.org/10.1016/j.jembe.2012.10.011
Clusa M, Carreras C, Pascual M, Gaughran SJ, Piovano S, Giacoma C, Fernández G, Levy Y, Tomás J, Raga JA, Maffucci F, Hochscheid S, Aguilar A, Cardona L (2014) Fine-scale distribution of juvenile Atlantic and Mediterranean loggerhead turtles (Caretta caretta) in the Mediterranean Sea. Mar Biol 161:509–519. https://doi.org/10.1007/s00227-013-2353-y
Crandall KA, Templeton AR (1993) Empirical tests of some predictions from coalescent theory with applications to intraspecific phylogeny reconstruction. Genetics 134:959–969. https://doi.org/10.1093/genetics/134.3.959
Danecek P, Bonfield JK, Liddle J, Marshall J, Ohan V, Pollard MO, Whitwham A, Keane T, McCarthy SA, Davies RM, Li H (2021) Twelve years of SAMtools and BCFtools. Gigascience 10: giab008. https://doi.org/10.1093/gigascience/giab008
Drosopoulou E, Tsiamis G, Mavropoulou M, Vittas S, Katselidis KA, Schofield G, Palaiologou D, Sartsidis T, Bourtzis K, Pantis J (2012) The complete mitochondrial genome of the loggerhead turtle Caretta caretta (Testudines: Cheloniidae): Genome description and phylogenetic considerations. Mitochondrial DNA 23:1–12. https://doi.org/10.3109/19401736.2011.637109
Drummond AJ, Rambaut A (2007) BEAST: Bayesian evolutionary analysis by sampling trees. BMC Evol Biol 7:214. https://doi.org/10.1186/1471-2148-7-214
Duchene S, Frey A, Alfaro-Núñez A, Dutton PH, Gilbert MTP, Morin PA (2012) Marine turtle mitogenome phylogenetics and evolution. Mol Phylogenet Evol 65:241–250. https://doi.org/10.1016/j.ympev.2012.06.010
Dutton PH, Bowen BW, Owens DW, Barragan A, Davis SK (1999) Global phylogeography of the leatherback turtle (Dermochelys coriacea). J Zool 248:397–409. https://doi.org/10.1111/j.1469-7998.1999.tb01038.x
Feldheim KA, Gruber SH, DiBattista JD, Babcock EA, Kessel ST, Hendry AP, Pikitch EK, Ashley MV, Chapman DD (2014) Two decades of genetic profiling yields first evidence of natal philopatry and long-term fidelity to parturition sites in sharks. Mol Ecol 23:110–117. https://doi.org/10.1111/mec.12583
Fetzner JW Jr, Crandall KA (2003) Linear habitats and the nested clade analysis: an empirical evaluation of geographic versus river distances using an Ozark crayfish (Decapoda: Cambaridae). Evolution 57:2101–2118. https://doi.org/10.1111/j.0014-3820.2003.tb00388.x
Feutry P, Kyne PM, Pillans RD, Chen X, Naylor GJ, Grewe PM (2014) Mitogenomics of the Speartooth Shark challenges ten years of control region sequencing. BMC Evol Biol 14:1–9. https://doi.org/10.1186/s12862-014-0232-x
Feutry P, Kyne PM, Pillans RD, Chen X, Marthick JR, Morgan DL, Grewe PM (2015) Whole mitogenome sequencing refines population structure of the Critically Endangered sawfish Pristis pristis. Mar Ecol Prog Ser 533:237–244. https://doi.org/10.3109/19401736.2015.1018208
Frandsen HR, Figueroa DF, George JA (2020) Mitochondrial genomes and genetic structure of the Kemp’s ridley sea turtle (Lepidochelys kempii). Ecol Evol 10:249–262. https://doi.org/10.1002/ece3.5891
Garofalo L, Mingozzi T, Mico A, Novelletto A (2009) Loggerhead turtle (Caretta caretta) matrilines in the Mediterranean: further evidence of genetic diversity and connectivity. Mar Biol 156:2085–2095. https://doi.org/10.1007/s00227-009-1239-5
Garofalo L, Mastrogiacomo A, Casale P, Carlini R, Eleni C, Freggi D, Gelli D, Knittweis L, Mifsud CR, Mingozzi T, Novarini N, Scaravelli D, Scillitani G, Oliverio M, Novelletto A (2013) Genetic characterization of central Mediterranean stocks of the loggerhead turtle (Caretta caretta) using mitochondrial and nuclear markers, and conservation implications. Aquat Conserv 23:868–884. https://doi.org/10.1002/aqc.2338
Green M, Sambrook J (2012) Molecular cloning: a laboratory manual. Cold Spring Harbor Laboratory Press, New York, NY
Hasegawa M, Kishino H, Yano T-a (1985) Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J Mol Evol 22:160–174. https://doi.org/10.1007/BF02101694
Hernández-Fernández J, Delgado Cano DA (2018) Complete mitochondrial genome of the three Colombian Caribbean loggerhead sea turtles, using next-generation sequencing. Mitochondrial DNA B 3:626–627. https://doi.org/10.1080/23802359.2018.1450664
Hochscheid S, Maffucci F, Abella E, Bradai MN, Camedda A, Carreras C, Claro F, de Lucia GA, Jribi I, Mancusi C, Marco A, Marrone N, Papetti L, Revuelta O, Urso S, Tomás J (2022) Nesting range expansion of loggerhead turtles in the Mediterranean: Phenology, spatial distribution, and conservation implications. Glob Ecol Conserv 38:e02194. https://doi.org/10.1016/j.gecco.2022.e02194
Jensen MP, FitzSimmons NN, Dutton PH, Michael P (2013) Molecular genetics of sea turtles. In: Wyneken J, Lohmann K, Musick JA (eds) The biology of sea turtles, vol III. CRC Press. Boca Raton, FL, pp 135–161
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Langmead B, Salzberg SL (2012) Fast gapped-read alignment with Bowtie 2. Nat Methods 9:357–359. https://doi.org/10.1038/nmeth.1923
Laurent L, Casale P, Bradai MN, Godley BJ, Broderick G, Gerosa AC, Schroth W, Schierwater B, Levy AM, Freggi D, Abd EM, Hadoud DA, Gomati HE, Domingo M, Hadjichristophprou M, Kornaraky L, Demirayak F, Gautier CH (1998) Molecular resolution of marine turtle stock composition in fishery bycatch: a case study in the Mediterranean. Mol Ecol 7:1529–1542. https://doi.org/10.1046/j.1365-294x.1998.00471.x
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Loisier A, Savelli M-P, Arnal V, Claro F, Gambaiani D, Sénégas JB, Cesarini C, Sacchi J, Miaud C, Montgelard C (2021) Genetic composition, origin and conservation of loggerhead sea turtles (Caretta caretta) frequenting the French Mediterranean coasts. Mar Biol 168:52. https://doi.org/10.1007/s00227-021-03855-6
Maffucci F, Corrado R, Palatella L, Borra M, Marullo S, Hochscheid S, Lacorata G, Iudicone D (2016) Seasonal heterogeneity of ocean warming: a mortality sink for ectotherm colonizers. Sci Rep 6:23983. https://doi.org/10.1038/srep23983
Mancino C, Canestrelli D, Maiorano L (2022) Going west: range expansion for loggerhead sea turtles in the Mediterranean Sea under climate change. Glob Ecol Conserv 38:e02264. https://doi.org/10.1016/j.gecco.2022.e02264
Margaritoulis D, Dean CJ, Lourenço G, Rees AF, Riggall TE (2020) Reproductive longevity of loggerhead sea turtles nesting in Greece. Chelonian Conserv Biol 19:133–136. https://doi.org/10.2744/CCB-1437.1
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654. https://doi.org/10.1038/351652a0
Michaels G, Hauswirth W, Laipis P (1982) Mitochondrial DNA copy number in bovine oocytes and somatic cells. Dev Biol 94:246–251. https://doi.org/10.1016/0012-1606(82)90088-4
Miller J (1997) Reproduction in sea turtles. In: Musick JA (ed) Lutz P. The biology of sea turtles. CRC Press, Boca Raton, FL, pp 51–81
Milne I, Stephen G, Bayer M, Cock P, Pritchard L, Cardle L, Shaw P, Marshall D (2013) Using Tablet for visual exploration of second-generation sequencing data. Brief Bioinform 14:193–202. https://doi.org/10.1093/bib/bbs012
Mingozzi T, Masciari G, Paolillo G, Pisani B, Russo M, Massolo A (2006) Discovery of a regular nesting area of loggerhead turtle Caretta caretta in southern Italy: a new perspective for national conservation. In: Hawksworth D, Bull A (eds) Biodiversity conservation in Europe Topics in biodiversity and conservation, vol 7. Springer, Dordrecht, pp 277–299
Morin PA, Archer FI, Foote AD, Vilstrup J, Allen EE, Wade P, Durban J, Parsons K, Pitman R, Li L, Bouffard P, Abel Nielsen SC, Rasmussen M, Willerslev E, Gilbert MT, Harkins T (2010) Complete mitochondrial genome phylogeographic analysis of killer whales (Orcinus orca) indicates multiple species. Genome Res 20:908–916. https://doi.org/10.1101/gr.102954.109
Moritz C (1994) Applications of mitochondrial DNA analysis in conservation: a critical review. Mol Ecol 3:401–411. https://doi.org/10.1111/j.1365-294X.1994.tb00080.x
Novelletto A, Testa L, Iacovelli F, Blasi P, Garofalo L, Mingozzi T, Falconi M (2016) Polymorphism in mitochondrial coding regions of Mediterranean loggerhead turtles: evolutionary relevance and structural effects. Physiol Biochem Zool 89:473–486. https://doi.org/10.1086/688679
Otálora K, Hernández-Fernández J, Carlos Salcedo-Reyes J (2018) Complete Colombian Caribbean loggerhead turtle mitochondrial genome: tRNA structure analysis and revisited marine turtle phylogeny. Univ Sci 23:355–381
Paradis E (2010) pegas: an R package for population genetics with an integrated–modular approach. Bioinformatics 26:419–420. https://doi.org/10.1093/bioinformatics/btp696
Pianka ER (1973) Evolutionary ecology. Harper & Row, New York, NY
Rambaut A, Drummond AJ, Xie D, Baele G, Suchard MA (2018) Posterior summarization in bayesian phylogenetics using Tracer 1.7. Syst Biol 67:901–904. https://doi.org/10.1093/sysbio/syy032
Robin E, Wong R (1988) Mitochondrial DNA molecules and virtual number of mitochondria per cell in mammalian cells. J Cell Physiol 136:507–513. https://doi.org/10.1002/jcp.1041360316
Saied A, Maffucci F, Hochscheid S, Dryag S, Swayeb B, Borra M, Ouerghi A, Procaccini G, Bentivegna F (2012) Loggerhead turtles nesting in Libya: an important management unit for the Mediterranean stock. Mar Ecol Prog Ser 450:207–218. https://doi.org/10.3354/meps09548
Shamblin BM, Bjorndal KA, Bolten AB, Hillis-Starr ZM, Lundgren I, Naro-Maciel E, Nairn CJ (2012a) Mitogenomic sequences better resolve stock structure of southern Greater Caribbean green turtle rookeries. Mol Ecol 21:2330–2340. https://doi.org/10.1111/j.1365-294X.2012.05530.x
Shamblin BM, Bolten AB, Bjorndal KA, Dutton PH, Neilen JT, Abreu-Grobois FA, Reich KJ, Witherington BE, Bagley DA, Ehrhart LM, Tucker AD, Addison DS, Arenas A, Johnson C, Carthy RR, Lamont MM, Dodd MG, Gaines MS, LaCasella E, Nairn CJ (2012b) Expanded mitochondrial control region sequences increase resolution of stock structure among North Atlantic loggerhead turtle rookeries. Mar Ecol Prog Ser 469:145–160. https://doi.org/10.3354/meps09980
Shamblin BM, Bolten AB, Abreu-Grobois FA, Bjorndal KA, Cardona L, Carreras C, Clusa M, Monzón-Argüello C, Nairn CJ, Nielsen JT (2014) Geographic patterns of genetic variation in a broadly distributed marine vertebrate: new insights into loggerhead turtle stock structure from expanded mitochondrial DNA sequences. PLoS ONE 9:e85956. https://doi.org/10.1371/journal.pone.0085956
Sheridan CM, Spotila JR, Bien WF, Avery HW (2010) Sex-biased dispersal and natal philopatry in the diamondback terrapin, Malaclemys terrapin. Mol Ecol 19:5497–5510. https://doi.org/10.1111/j.1365-294X.2010.04876.x
Shields WM (1982) Philopatry, inbreeding, and the evolution of sex. State University of New York Press, New York, NY
Splendiani A, Fioravanti T, Giovannotti M (2017) Mitochondrial DNA reveals the natal origin of Caretta caretta (Testudines: Cheloniidae) stranded or bycaught along the Southwestern Adriatic coasts. Eur Zool J 84:566–574. https://doi.org/10.1080/24750263.2017.1400597
Suchard MA, Lemey P, Baele G, Ayres DL, Drummond AJ, Rambaut A (2018) Bayesian phylogenetic and phylodynamic data integration using BEAST 1.10. Virus Evol 4. https://doi.org/10.1093/ve/vey016
Templeton AR, Crandall KA, Sing CF (1992) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping and DNA sequence data. III. Cladogram estimation. Genetics 132: 619–633. https://doi.org/10.1093/genetics/132.2.619
Templeton AR, Sing CF (1993) A cladistic analysis of phenotypic associations with haplotypes inferred from restriction endonuclease mapping. IV. Nested analyses with cladogram uncertainty and recombination. Genetics 134:659–669. https://doi.org/10.1093/genetics/134.2.659
Tikochinski Y, Carreras C, Tikochinski G, Vilaça ST (2020) Population-specific signatures of intra-individual mitochondrial DNA heteroplasmy and their potential evolutionary advantages. Sci Rep 10:211. https://doi.org/10.1038/s41598-019-56918-6
Tolve L, Casale P, Formia A, Garofalo L, Lazar B, Natali C, Novelletto A, Vallini C, Bužan E, Chelazzi G, Gaspari S, Fortuna C, Kocijan I, Marchiori E, Novarini N, Poppi L, Salvemini P, Ciofi C (2018) A comprehensive mitochondrial DNA mixed-stock analysis clarifies the composition of loggerhead turtle aggregates in the Adriatic Sea. Mar Biol 165:68. https://doi.org/10.1007/s00227-018-3325-z
Wallace BP, DiMatteo AD, Hurley BJ, Finkbeiner EM, Bolten AB, Chaloupka MY, Hutchinson BJ, Abreu-Grobois FA, Amorocho D, Bjorndal KA, Bourjea J, Bowen BW, Dueñas RB, Casale P, Choudhury BC, Costa A, Dutton PH, Fallabrino A, Girard A, Girondot M, Godfrey MH, Hamann M, López-Mendilaharsu M, Marcovaldi MA, Mortimer JA, Musick JA, Nel R, Pilcher NJ, Seminoff JA, Troëng S, Witherington B, Mast RB (2010) Regional Management Units for marine turtles: a novel framework for prioritizing conservation and research across multiple scales. PLoS ONE 5:e15465. https://doi.org/10.1371/journal.pone.0015465
Wiesner R, Rüegg J, Morano I (1992) Counting target molecules by exponential polymerase chain reaction: copy number of mitochondrial DNA in rat tissues. Biochem Biophys Res Commun 183:553–559. https://doi.org/10.1016/0006-291x(92)90517-o
Yan K, An T, Zhai M, Huang Y, Wang Q, Wang Y, Zhang R, Wang T, Liu J, Zhang Y, Zhang J, Wang K (2019) Mitochondrial miR-762 regulates apoptosis and myocardial infarction by impairing ND2. Cell Death Dis 10:500. https://doi.org/10.1038/s41419-019-1734-7
Yilmaz C, Turkozan O, Bardakci F (2011) Genetic structure of loggerhead turtle (Caretta caretta) populations in Turkey. Biochem Syst Ecol 39:266–276. https://doi.org/10.1016/j.bse.2011.08.018
Acknowledgements
We are grateful to the following collaborators involved in sample collection: Nicola Novarini of the Natural History Museum of Venice, Lisa Poppi, Caterina Raso, Luca Marini and collaborators of the NGO TartaLazio and all voluntary workers and former students involved in the TartaCare Project of Calabria University including Sandra Berlingeri, Giulia Cambiè, Francesca Crispino, Concetta Denaro, Simona Fabiano, Elena Giardinazzo, Teresa Malito, Carmela Mancuso, Giampiero Masciari, Alessandro Massolo, Nunzia Micò, Gianni Parise, Brunella Pisani, Manuela Policastrese, Patrizia Rima, Manuela Russo, Salvatore Salice and Salvatore Urso. We also thank Luana Papetti and collaborators of the NGO tartAmare and Isa Tonso and collaborators of Legambiente Arcipelago Toscano for support during collection of pipped eggs from nesting sites in Tuscany. A special thanks to Enrico Negrisolo for valuable suggestions during the preparation of the manuscript.
Funding
Open access funding provided by Università degli Studi di Firenze within the CRUI-CARE Agreement. This study was funded by the Tuscany Regional Authority NATura Network Toscana agreement in compliance to the Habitats Directive 92/43/EEC. Additional support was provided by the Italian Ministry of University and Research through the National Biodiversity Future Center, part of the National Recovery and Resilience Plan, Mission 4, Component 2, Investment 1.4, Project CN00000033. LT was granted a graduate research scholarship supported by the European Structural and Investment Funds through the National Operational Programme (PON) of the Italian Ministry of University and Research.
Author information
Authors and Affiliations
Contributions
LT, AI, ANo, LG and CCi designed and conceived the study. LG, ACD, ADL, CCo, CM, EM, GT, IC, LM, MALZ, SN and TM were involved in sample collection. CCi, AF, LM and CM secured funding for laboratory work and consumables. LT, AI and CN performed genetic analyses. LT, AI, ANo, ANi, FI, LG and MF contributed to data analysis. LT, AI, ANo, LG and CCi wrote the paper. All authors revised and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
Authors declare no conflict of interest.
Ethical standards
Sampling was conducted in compliance to the exemption provided by Article VII, paragraph 6 of CITES to the University of Pisa and the University of Siena (register of scientific institutions codes IT026 and IT007, respectively) and in accordance to permit 27676 of 16 March 2021 granted to MALZ by the Italian Ministry for the Ecological Transition. The study complies with institutional, national, and international ethics guidelines concerning the use of animals in research and/or the sampling of endangered species. None of the procedures used in the study met the criteria to define them “experiments” as defined in article 2 of the Council of European Communities Directive 86/609/EEC regarding the protection of animals used for experimental and other scientific purposes.
Additional information
Responsible Editor: C. Eizaguirre.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Andrea Novelletto Co-senior author.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Tolve, L., Iannucci, A., Garofalo, L. et al. Whole mitochondrial genome sequencing provides new insights into the phylogeography of loggerhead turtles (Caretta caretta) in the Mediterranean Sea. Mar Biol 171, 19 (2024). https://doi.org/10.1007/s00227-023-04325-x
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-023-04325-x