Abstract
Appendicularian tunicates are some of the most abundant mesozooplankton organisms with key roles in marine trophic webs and global carbon flux. Like most appendicularians with cosmopolitan distributions, Oikopleura dioica Fol, 1872 is considered a single species worldwide based on morphological features that distinguish them from other appendicularians. Despite their abundance, however, there are still only ~ 70 described appendicularian species, compared to over 2800 ascidian tunicates. Here we perform a molecular phylogenetic, morphological, and reproductive assessment of O. dioica specimens collected from the Ryukyu Archipelago, mainland Japan, and Europe. The specimens are morphologically very similar, with only detailed examination of the oikoplastic epithelium and quantitative measurements revealing minor distinguishing characteristics. Phylogenetic analyses of the ribosomal gene loci and mitochondrial cytochrome oxidase I (COI) gene strongly indicate that they form three separate genetic clades despite their morphological similarities. Finally, in vitro crosses between the Ryukyu and mainland Japanese specimens show total prezygotic reproductive isolation. Our results reveal that the current taxonomic O. dioica classification likely hides multiple cryptic species, highlighting the genetic diversity and complexity of their population structures. Cryptic organisms are often hidden under a single species name because their morphological similarities make them difficult to distinguish and their correct identification is fundamental to understanding Earth’s biodiversity. O. dioica is an attractive model to understand how morphological conservation can be maintained despite pronounced genetic divergence.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Appendicularians or larvaceans (Tunicata: Appendicularia) are abundant holoplanktonic tunicates with a global distribution. They are one of the most abundant taxonomic groups in the zooplankton community (Alldredge 1976; Clarke and Roff 1990; Hopcroft and Roff 1995). Under optimal conditions, appendicularians form rapid blooms (Essenberg 1922; Tokioka 1955) exceeding 53,000 individuals per cubic metre (Uye and Ichino 1995). The animals secrete an intricate structure known as a house, which they use to filter-feed suspended dissolved organic particles and microorganisms from the water (Sato et al. 2001); this allows them to feed on pico- and nanophytoplankton that are often too small to be captured by most zooplankton (Alldredge1976; Saito 2019). Therefore, they serve as an important intermediate link in the food chain connecting pico- and nanophytoplankton and larger zooplankton including fish larvae (Sakaguchi et al. 2017) copepods (Ohtsuka and Onbé 1989), ctenophores (Shiganova 2005; Purcell et al. 2010), jellyfish, and chaetognaths (Alldredge 1976). Most appendicularians measure from 2 to 8 mm in length, with houses ranging from 4 to 38 mm in diameter (Alldredge 1977). However, Bathochordaeus, known as giant larvaceans, have body lengths of ~ 10 cm and construct houses that exceed 1 m in diameter (Hamner 1992; Katija et al. 2017). Appendicularians maintain efficient feeding by continuously replacing houses: for instance, a single Oikopleura dioica can build approximately 50 houses during its life cycle (Sato et al. 2001). Discarded houses together with trapped materials form carbon-rich aggregates and are one of the main components of the marine snow that eventually sinks to the seabed. Therefore, appendicularians make a major contribution to vertical global carbon flux by transporting a substantial amount of surface productivity to the deep benthic fauna (Alldredge et al. 2005; Robison et al. 2005).
Despite their abundance and ecological importance in functioning ecosystems, the recorded appendicularian diversity remains low with only ~ 70 described species worldwide (Tokioka 1960; Hopcroft et al. 2005; Gorsky et al. 2017). This trend is especially apparent when compared with sessile tunicates, ascidians, which comprise over 2800 described species (Shenkar and Swalla 2011). The comparative lack of knowledge is partly due to the challenge of studying appendicularians. Soft-bodied appendicularians are extremely fragile; they are easily damaged during conventional plankton sampling, leaving little material for morphological identifications (Tokioka 1960). In addition, their distributions are often seasonal (Uye and Ichino 1995; Tomita et al. 2003); therefore, continuous and extensive sampling effort is needed to appreciate their true diversity. Furthermore, there might be potential cryptic speciation among currently recognized species (Hopcroft and Robison (Hopcroft 1999; Hopcroft et al. 2005): cryptic species are defined as “two or more distinct species that are erroneously classified (and hidden) under one species name” because they are superficially morphologically indistinguishable (Bickford et al. 2007). They are common in many marine taxa including foraminifera (Vargas et al. 1999), cnidarians (Dawson and Jacobs 2001), fishes (Borsa 2002), elasmobranchs (Quattro et al. 2006), and copepods, with numerous sibling species retaining strong morphological conservation (Chen and Hare 2011; Blanco-Bercial et al. 2014).
There have been three significant periods of appendicularian taxonomic discovery: exploratory studies led by Lohmann, Bückmann, and Hentschel in the late 19th to early twentieth centuries; Japanese exploration by Tokioka in the 1950s; and mesopelagic explorations led by Fenaux, Youngbluth, Hopcroft, Robison, and Flood in the late twentieth century. These efforts contributed to the current understanding of appendicularian diversity and distributions (Hopcroft et al. 2005). Most appendicularian species were previously delineated using classical morphology-based taxonomy. However, to assess accurately the true diversity of appendicularians, it is also important to examine molecular data that might uncover genetically distinct lineages hidden behind morphologically conserved populations.
Recent combined morphological and molecular taxonomic analyses have described new species of appendicularians and closely related taxa (Garić and Batistić 2016; Sherlock et al. 2017). The molecular approach provides additional evidence for species identification and enables a more accurate assessment of morphologically conserved taxa. Markers such as the cytochrome c oxidase subunit I (COI) and ribosomal DNA (rDNA) genes are often used to elucidate phylogenetic relationships (Zagoskin et al. 2014). In most eukaryotes including appendicularians, rDNA genes are organized in tandemly repeated clusters, each containing the 18S, 5.8S, and 28S ribosomal genes separated by internal transcribed spacers (ITS1 and ITS2) and an intergenic spacer (Singer and Berg 1991). Phylogenetic relationships at higher systematic levels (orders, families, and genera) are often resolved using the highly conserved 18S and 28S rDNA regions. The less conserved internal spacer sequences enable greater phylogenetic resolution of lower taxonomic groups, including species (Schlötterer et al. 1994; Hwang and Kim 1999; Dyomin et al. 2017).
Oikopleura dioica is an appendicularian model that contributes to a wide range of biological studies including developmental biology, evolution, genomics, and ecology (Nishida 2008; Lombard et al. 2009; Troedsson et al. 2013; Deng et al. 2018; Onuma and Nishida 2021; Ferrández-Roldán et al. 2021). Much of our knowledge of appendicularian biology comes from multigenerational culturing of O. dioica in the laboratory (Paffenhöfer 1973; Bouquet et al. 2009; Martí-Solans et al. 2015; Masunaga et al. 2020). The organism is mainly characterized by its ubiquitous distribution, dioecious nature, and presence of two subchordal cells on its tail. O. dioica is currently considered a single species worldwide; however, there are substantial nucleotide sequence variations between O. dioica collected from Norway, Osaka, and Okinawa (Denoeud et al. 2010; Wang et al. 2015a, 2020; Bliznina et al. 2021) despite the low level of phenotypic disparity.
Here we perform a molecular phylogenetic, morphological, and reproductive assessment of O. dioica collected from the Ryukyu Archipelago, mainland Japan, and Europe. Our results reveal three genetically distinct lineages despite near-indistinguishable morphologies. We provide compelling evidence that O. dioica collected from the Ryukyu Archipelago is a separate species from those found in mainland Japan and Europe.
Materials and methods:
Origins of O. dioica data
O. dioica specimens were either obtained from field collections (Kume, Amami, Nagasaki, Hyogo, Aomori) or laboratory cultures (Okinawa, Osaka, Barcelona). Sequence information for molecular phylogenetic analyses was obtained from specimens, or where publicly available, from GenBank or the supplementary material of the corresponding paper (Norway, Croatia). Images, size measurements, and fertilization rates were obtained using laboratory specimens. Sequencing data for all other appendicularians were obtained from GenBank. Collection sites are shown in Fig. 1, and detailed sample information is provided in Supplementary Table S1.
Field collections of O. dioica from the Ryukyu Archipelago
We surveyed O. dioica along the Ryukyu Archipelago from harbours and fishing ports using a hand-held plankton net (100-µm mesh) with a 500 mL volume cod-end (Masunaga et al. 2020). Animals were obtained in June and July 2019 from Kin bay on Okinawa island (26°25′39.6″N 127°49′56.5″E), Kume island (26°21′03.2″N 126°49′17.4″E), and Amami island (28°24′53.6″N 129°35′27.9″E; Fig. 1). O. dioica were identified by the presence of two subchordal cells at the distal half of their tails under a dissecting microscope (Fredriksson and Olsson 1991; Fig. 2B). Live animals from Okinawa island were transported to the Okinawa Institute of Science and Technology Graduate University (OIST) to establish laboratory cultures (Masunaga et al. 2020). Animals from Kume and Amami islands were isolated and maintained in beakers with portable paddles and motors until they reached maturity on-site. Once the animals matured, males were rinsed three times for 10 min with autoclaved filtered seawater, preserved in RNAlater (ThermoFisher AM7021), and transported back to OIST.
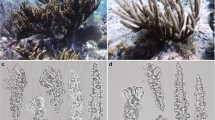
Okinawa and Osaka O. dioica laboratory strains. A Fully matured females (top) and males (bottom) of Osaka (left column) and Okinawan (right column) O. dioica maintained at the OIST culturing facility. B Okinawan O. dioica individual with two subchordal cells (white arrows) on its tail. (C) Dorsal and (D) lateral views of Okinawan O. dioica
Collections of O. dioica from mainland Japan and Europe
O. dioica specimens were generously donated by the Kujukushima Aquarium (UMI KIRARA; Nagasaki), the Nishida lab from Osaka University (Hyogo), the Nishino lab from Hirosaki University (Aomori), and the Cañestro lab from the University of Barcelona (Barcelona). One of the Osaka and the Barcelona samples are laboratory cultures of locally collected specimens.
Laboratory culturing of O. dioica specimens
Live O. dioica laboratory animals originally collected from Sakoshi bay in Hyogo prefecture (Osaka lab strain) were kindly provided by the Nishida lab and cultured at the OIST culturing facility. Both Okinawa and Osaka lab strains are cultured at 23 °C at OIST (Masunaga et al. 2020). The Barcelona laboratory strain is maintained by the Cañestro lab at the University of Barcelona (Martí-Solans et al. 2015). All animals are fed once or twice a day on a mixture of microalgae and cyanobacteria as previously described (Martí-Solans et al. 2015; Masunaga et al. 2020).
Genomic DNA isolation and sequencing
Fully mature males were rinsed and either processed immediately for DNA extraction or preserved in RNAlater for transport. Genomic DNA (gDNA) was isolated using a modified salting-out protocol. First, tissues were lysed with 10 µL/mg proteinase K for 30 min at 56 °C. Next, proteins were precipitated out with 50 µL 5 M NaCl and separated by centrifugation at 5000 × g for 15 min at 4 °C. The aqueous phase was slowly transferred with a wide-bore pipette to a tube containing 400 µL of cold 100% ethanol and 10 µg/mL glycogen. gDNA was precipitated at − 80 °C for 20 min and centrifuged at 6520 × g for 5 min at 4 °C. Finally, the pellet was washed with 200 µL of 70% ethanol, allowed to air dry for 5 min, and resuspended in molecular biology grade H2O. gDNA was quantified using a Qubit 3 Fluorometer (Thermo Fisher Scientific, Q32850), and its integrity was measured using Agilent 4200 TapeStation (Agilent, 5067-5365). Isolated gDNA were processed with the Ligation Sequencing Kit (Nanopore LSK109) according to the manufacturer’s protocol and sequenced on the Nanopore MinION platform.
18S and ITS phylogenetic analysis
We combined appendicularian 18S ribosomal DNA (rDNA) gene sequences from GenBank with 18S RNA gene sequences identified from our own genome assemblies. All sequences have been deposited in the GenBank database (accession numbers provided in Supplementary Table S1). To identify 18S rDNA genes, we searched genome sequences for the best-hit matches to the Rfam models for the eukaryotic small subunit (SSU; RF01960) and large subunit (LSU; RF02543) using cmsearch of the Infernal package (v.1.1.4). The SSU and LSU models from Rfam were downloaded in April 2021. The internal transcribed spacer (ITS) regions were obtained by extracting the region between SSU and LSU. Multiple sequence alignments for the 18S and ITS regions were created with MUSCLE (v5) within Seaview (v.3.2). Phylogenetic trees were estimated using maximum likelihood with IQ-TREE software (Trifinopoulos et al. 2016) and Bayesian inference with MrBayes (v.3.2.0) (Ronquist and Huelsenbeck 2003). The ML method was calculated using the best model estimated by ModelFinder (GTR + F + I + G4) (Kalyaanamoorthy et al. 2017) according to the Bayesian Information criterion with 1,000 ultrafast bootstrap replicates (Hoang et al. 2018). For Bayesian inference, a GTR substitution model with 6 gamma-distributed rate categories and an uninformative Dirichlet prior was used. Three independent MCMC chains were run with a maximum of 1,000,000 generations and 25% burn-in fraction, allowing the chain to terminate when the standard deviation of split frequencies was lower than 0.01. The 50% majority rule consensus trees with their posterior probabilities are presented.
COI phylogeny
The mitochondrial cytochrome oxidase I (COI) sequences representing the European lineage were obtained from (Pichon et al. 2019) (PMID: 32,148,763). The COI sequence representing the mainland Japanese lineage is contig comp28493_c0_seq1_38 from the transcriptome assembly of (Wang et al. 2015a), GEO ID: GSE64421, PMID: 26,032,664). The COI sequence representing the Ryukyu lineage is the transcript model TRINITY_DN9989_c0_g1_i1 from the transcriptome assembly from (Bliznina et al. 2021).
Imaging the oikoplastic epithelium
Okinawa and Osaka laboratory strains
To examine the oikoplastic epithelium (OE), immature adults were fixed with 4% paraformaldehyde in 0.1 M MOPS and 0.5 M NaCl buffer (PFA). Fixed animals were stored at 4 °C until use. The animals were washed three times with PBST (PBS with 0.1% Triton X), and nuclei were stained with DAPI. After 2 min incubation at room temperature, the animals were washed three times with PBST and mounted in 30% glycerol. The nuclei of the OE were imaged using a Nikon A1R confocal microscope with a 20 × objective. A volume of 40 planes (0.7 micron steps) was observed and two images were averaged for every plane. The volumes were 3D rendered using the NIS Elements imaging software (v.5.21.02) with a 3D LUT (Depth coded Alpha, Altitude). Maximum intensity projections (MIPs) were extracted as a single volume snapshot. Images were post-processed using Adobe Photoshop CC (22.4.1). The number of nuclei in the Fol and the Eisen domains of all specimens were manually identified and counted using the nomenclature described by (Kishi et al. 2017).
Barcelona laboratory strain
Barcelona animals were fixed with 4% paraformaldehyde in 0.1 M MOPS, 0.5 M NaCl, 2 mM MgSO4, 1 mM EGTA. The animals were washed with PBS with 0.1% Tween-20, and nuclei were stained with Hoechst. The animals were mounted in 80% glycerol. The nuclei were imaged using a Zeiss LSM880 microscope with a 25X objective.
Measuring trunk and tail ratios
Trunk and tail measurements were taken for approximately 50 animals of various developmental stages post-hatching. Each animal was transferred to a Petri dish containing 0.5% agarose. Excess seawater was removed until animals lay flat on the agarose bed and were immobilized. Each animal was photographed (Leica MC 190 HD or BFLY-U3-23S6C) and the trunk length (distal end of gonad to the mouth) and the tail length were measured using the Leica Las software (v.4.9) or FIJI (Schindelin et al. 2012). The trunk-to-tail ratio was calculated as trunk length (mm)/tail length (mm).
Measuring egg diameters
To compare egg diameters, approximately 20 females were collected from each laboratory strain. Females were rinsed three times with 5 mL of autoclaved filtered seawater (AFSW) to avoid the potential presence of sperm from the cultures. Each female was transferred to a Petri dish and allowed to spawn naturally. Photographs of eggs were taken, upon spawning, and five representative eggs from each clutch were selected for measurements. For each female, we report the mean diameter of five eggs. The data was not normally distributed; therefore, Kruskal–Wallis test was performed on both trunk and tail and egg diameter measurements using R (v.3.6.3, R Core Team 2020).
Crossing experiment
60 fully mature Okinawa and Osaka animals were randomly assigned to four mating groups during the experimental period of April to May 2018: 10 pairs of Okinawa males and Osaka females, 10 pairs of Okinawa females and Osaka males, and 5 pairs from each strain for positive controls (Okinawa males and females, Osaka males and females). Animals were rinsed three times with 5 mL of AFSW. Each male was transferred to an Eppendorf tube containing 50 µL of AFSW. Upon spawning, the sperm solution was diluted with 500 µL of AFSW and kept on ice. Each female was left to spawn naturally in a Petri dish with 5 mL of AFSW. Approximately 30–50 eggs were transferred to 500 µL of AFSW and fertilized with 10 µL of the sperm solution. Five minutes post-fertilization (pf), excess sperm was removed by moving the eggs to a new dish with 500 µL of AFSW. Eggs were monitored under the stereomicroscope (Leica M165 FC) and photographed at 15 min, 30 min, 1 h, and 3 h pf. The number of fertilized eggs was manually counted from the photographs taken at 30 min pf mark.
Results
Overall morphology
We collected specimens along the Ryukyu Archipelago using a hand-held plankton net. Specimens collected from Kin Bay in Okinawa Island (Okinawa lab strain) were cultured in the laboratory alongside the Osaka lab strain (Fig. 1; Material and Methods).
The overall morphologies of the Okinawa lab strain are congruent with previously described O. dioica characteristics (Fol 1872; Gorsky et al. 2017). It is the only known dioecious appendicularian species; the sexes of Okinawan individuals become visibly distinguishable towards the end of the lifecycle as the gonads at the rear of the body fill with sperm or eggs (Fig. 2A). Viewed dorsally, there are two spindle-shaped subchordal cells to the left of the notochord, generally located in the distal half of the tail (Fig. 2B), with the spacing of the two subchordal cells varying between individuals. The mouth is almost circular with upper and lower lips and there are two spherical buccal glands nearby. The oesophagus connects the mouth to the left lobe of the stomach, which then leads to the right lobe. There are round, ciliated spiracles on either side of the rectum that are connected to the pharynx. The U-shaped endostyle is situated ventrally to the pharynx. The brain is present in the anterodorsal region near the pharynx (Fig. 2C, D). According to these morphological criteria, we concluded that the specimens collected in Okinawa were O. dioica.
18S phylogeny
To compare the Ryukyu specimens to other appendicularian species, we estimated phylogenetic trees of 18S ribosomal rDNA sequences using maximum likelihood (ML) and Bayesian methods. We combined publicly available data from GenBank with 18S rDNA sequences retrieved from the genome sequences of our own and collaborators’ specimens. The resulting trees show that all O. dioica specimens form a distinct cluster from other appendicularian species (Fig. 3; ultrabootstrap support, 100%; posterior probability, 1.00).
Maximum likelihood phylogenetic tree estimated using ribosomal 18S DNA sequences. Each entry shows the species name, geographical origin, and sample ID. The support values of nodes with > 50% bootstrap support and > 0.5 posterior probability are shown. Branch support represents maximum likelihood ultrafast bootstrap value (1000 replicates)/posterior probability (Bayesian inference). Branch length is proportional to the size of the clade. Polytomies in the Bayesian tree are treated as supporting a clade
However, within the O. dioica cluster, it appears that the Ryukyu, mainland Japanese and European specimens each form distinct clades. Although the monophyly of the Ryukyu clade is supported by both ML and Bayesian trees, they present conflicting topologies with respect to its placement within O. dioica (Fig. 3; Supplementary Figure S1). We reasoned that since O. dioica 18S rDNA sequences are highly conserved, with pairwise sequence identities ranging from 99.8 to 100%–one to three nucleotide changes – it is difficult to resolve the close evolutionary relationships within the O. dioica monophyletic group (Table 1).
ITS phylogeny
To avoid relying on a single molecular marker and improve phylogenetic resolution, we constructed trees using the less conserved, non-coding, internal transcribed spacer (ITS) region of ribosomal loci (Fig. 4). This analysis also generated three highly supported clades within the O. dioica monophyletic group. ITS trees indicate that the mainland Japanese and European clades are most closely related, whereas the Ryukyu clade forms a more distant, highly supported lineage. The higher sequence divergence of ITS regions compared with the 18S rDNA genes enables phylogenetic reconstruction at a higher resolution (pairwise sequence identities vary from 84.0% for Amami (AMA-15) vs. Norway (OdB3) to 99.7% for Osaka (O9) vs. Nagasaki (Oidioi_NAG1), Osaka (O9) vs. Aomori (AOM-10). In comparison, the sequence identity between Ciona intestinalis and the congeneric Ciona robusta is 93.1% (Table 1). Although the Ryukyu clade is highly supported in the 18S and ITS trees, the identity of the basal group is ambiguous (Fig. 4, Supplementary Figure S1).
Maximum likelihood phylogenetic tree estimated for O. dioica internal transcribed spacer sequences. Support values for nodes with bootstrap support > 50% and posterior probability > 0.5 are shown. Branch support indicates maximum likelihood ultrafast bootstrap value (1000 replicates)/posterior probability (Bayesian inference). Branch length is proportional to the size of the clade. Polytomies in the Bayesian tree are treated as supporting a clade
COI phylogeny
To investigate further genetic differences between the three lineages, and to provide reference data for future studies, we searched the available transcriptomic data for mitochondrial cytochrome oxidase I (COI) sequences in the laboratory strain of each lineage. There are too few samples to construct phylogenetic trees, but alignments show pairwise nucleotide sequence identities between 83.4 and 85.5% (Table 1). Okinawa (OKI2018_I69_1.0) and Norway (OdB3) display the greatest pairwise divergence, which is consistent with the ITS comparisons. The translated protein sequences differ by only four or five amino acids in the C-terminal region and most changes occur among the hydrophobic amino acids Val-Leu-Met-Ile and Ala-Thr-Ser.
Morphological comparisons of the oikoplastic epithelium (OE)
Since our phylogenetic analyses revealed three distinct clades within the O. dioica monophyletic group, we decided to examine the morphologies of the OE in greater detail. The OE is a monolayer trunk epithelium responsible for synthesizing the cellulose house that surrounds the animals. The position, number, and shape of OE are considered to be specific to different appendicularian species (Spada et al. 2001; Flood et al. 2005). Cells are arranged in a bilaterally symmetric fashion (Ganot and Thompson 2002); therefore, we examined one side of the OE nuclei patterning in each specimen. We compared the OE nuclear pattern of Okinawa, Osaka, and Barcelona laboratory strains.
We found remarkable conservation in the number of nuclei and the spatial arrangement of the two most prominent domains of the OE (Fol and Eisen) between Okinawa, Osaka, and Barcelona lab strains (Fig. 5, Table 2). For all three lab strains, there are small specimen-to-specimen variations in the numbers of nuclei in the Chain of Pearls (CP) and Nasse (N) regions; these variabilities were also observed in O. dioica specimens from Osaka and Norway (Flood, et al. 2005; Kishi et al. 2017). Although the first row of the Posterior Fol (PF1) in Osaka was reported to have 10 cells (Kishi et al. 2017), our counts revealed 11 cells for Okinawa (n = 17), Osaka (n = 15), and Barcelona (n = 5) laboratory strains. Nuclear count details are shown in Supplementary Table S2.
Comparison of the nuclear pattern of the oikoplastic epithelium of Okinawa (left), Osaka (middle), and Barcelona (right) O. dioica lab strains. From the top row; the overview of the oikoplastic epithelium, close-up views of the Fol domain (orange), the Eisen domain (blue), and the mid-dorsal regions (yellow). The Fol domain is organized by: Anterior Crescent (AC), Anterior Elongate (AE), Giant Fol (GF), Nasse (N), and Posterior Fol (PF). The Eisen domain is organized by Eisen Giant cells (E), Eisen’s yard (EY), and Chain of Pearl (CP)
The number and the arrangement of the OE nuclei are highly conserved among the Okinawa, Osaka, and Barcelona individuals; however, closer examination revealed noticeable differences in the shape of nuclei in the mid-dorsal regions (Fig. 5). For Osaka animals, the nuclei are crescent-shaped compared with the roughly round shapes of the Okinawa and Barcelona strains. This observation was consistent throughout all the specimens examined during this study (Supplementary Figure S2).
Trunk/tail ratios and egg diameters
As DNA sequence analysis and visualization of the OE nuclei require instruments that are not yet readily available outside the laboratory, we sought additional morphological markers that are readily accessible in the field. We compared the trunk-to-tail ratios and the average egg diameters for the Okinawa, Osaka, and Barcelona laboratory strains (Fig. 6). There is a statistically significant difference in the trunk-to-tail ratios for Okinawa (mean ± 95% CI of 0.27 ± 0.007, n = 53), Barcelona (0.26 ± 0.006, n = 50), and Osaka strains (0.31 ± 0.004, n = 52; Kruskal–Wallis test, H (2) = 72.2, P < 2.2 e−16) with overlapping values (Fig. 6A). There is a statistically significant difference in the average egg diameter of Okinawa (mean ± 95% CI of 80 ± 0.0012 µm, n = 20), Osaka (96 ± 0.0015 µm, n = 21) and Barcelona females (96 ± 0.0039 µm, n = 21; Kruskal–Wallis test, H (2) = 38.7, P = 4e−09). Measurements on Okinawa and Osaka egg diameters show no overlapping values (Fig. 6B). Details on the trunk-to-tail ratios and the egg diameter measurements are shown in Supplementary Tables S3 and S4.
Gamete compatibility
Sixty Okinawa and Osaka O. dioica individuals were selected for in vitro fertilization (Fig. 7). 158 out of 179 (88.2%) eggs from Okinawa females and 194 out of 197 (98.5%) eggs from Osaka females were successfully fertilized in the positive control pairs. In contrast, none of the Okinawa-Osaka pairings led to fertilization: eggs inseminated with sperm from non-native males showed no signs of development 3 h pf, which is the approximate time to hatching under normal conditions at 20 °C. Individual measurements of the crossing experiment are shown in Supplementary Table S5. Therefore, the Okinawa and Osaka strains are reproductively incompatible.
Crossing experiments for Okinawa and Osaka laboratory strains. (A) Total number of eggs successfully fertilized for the Okinawa and Osaka mating pairs. (B) Photographs documenting development after crossing Okinawa and Osaka laboratory strains. Photographs were taken after 15 min, 30 min, 1 h, and 3 h post-fertilization attempt
Discussion
Here we presented evidence for cryptic speciation of O. dioica. We assessed three orthogonal pieces of evidence: phylogeny (Cracraft and Johnston 1983), morphology (Cronquist 1978), and reproductive compatibility (Mayr 1942). Although the Ryukyu, mainland Japanese, and European specimens exhibit remarkable conservation in morphology, our phylogenetic analyses suggest they comprise three genetically distinct lineages that are currently considered conspecific. It is possible, indeed likely, that the number of lineages will increase as samples from additional geographical locations are analysed in future studies.
Phylogeny
18S rDNA gene is a useful marker for distinguishing appendicularian species; however, it is too conserved to discriminate closely related O. dioica lineages (Fig. 3). The non-coding ITS region is useful for distinguishing recently diverged taxa within the O. dioica monophyletic group (Fig. 4). We therefore suggest ITS region as a candidate marker for O. dioica lineage identification and delimitation. The ITS region has been widely used in DNA barcoding analyses in many taxa including algae, protists, and animals and has been proposed as the standard DNA barcode for fungi and seed plants (Schlötterer et al. 1994; Yao et al. 2010; Wang et al. 2015b).
In addition to the three global lineages of O. dioica, there is local diversity within the Ryukyu clade (Fig. 4): there are three groups comprising (i) two individuals from Amami (AMA 15) and Kume islands (KUM-M3), (ii) another Amami individual (AMA 17) that clusters with specimens from Okinawa island (OKI2018_I69_1.0, Ishi1, and M2F1_6), and (iii) a Kume individual (KUM_M1) that clusters with two other Okinawa individuals (I1-5 and Ishi2). Despite this diversity, the placements suggest that specimens from Okinawa, Kume, and Amami islands form a single, continuous Ryukyu O. dioica population. Further examination of local samples will provide insights into the population structure and the distribution of the Ryukyu O. dioica clade and its boundaries with neighbouring populations.
Our results show that Oikopleura is not monophyletic, which disagrees with the current taxonomy available in resources such as the WORMS database. In future, a dedicated and detailed taxonomic study will be necessary to clarify the phylogeny of Oikopleura, Folia, Mesoikpleura and related genera.
Morphology
Closer examinations in the trunk-to-tail ratios and egg diameters reveal subtle quantitative differences. However, it is important to note that morphological measurements were taken from the specimens grown in laboratories. The size of animals including gonad length and egg diameters may vary depending on culture conditions, such as water quality and food availability.
Although not quantified, closer examinations in the trunk morphology reveal some differences in Barcelona specimens compared with Okinawan and Osaka. For example, the trunk of Barcelona specimens appears to be enlarged dorso-ventrally, whereas the Osaka and Okinawan specimens appear more elongated. In addition, the taper between the dorsal epithelium over the mouth and Fol domain of Barcelona specimens appear to be more pronounced (Fig. 5).
The oikoplastic epithelium (OE), which is responsible for creating the house structure in appendicularians species, is highly conserved between the three genetic O. dioica lineages. Only in the mid-dorsal region is it possible to observe distinct crescent-shaped nuclei in the Osaka specimens (Fig. 5). A similar observation was reported among Oikopleura longicauda, in which prominent differences in size and shape of the OE nuclei were found between specimens collected from different geographical regions (Flood, et al. 2005).
Many larger appendicularian species (trunk length > 1.4 mm), such as Oikopleura labradoriensis, O. vanhoeffeni, and O. villafrancae, exhibit more complex and arborized patterns of OE nuclei compared with smaller appendicularians such as O. dioica, O. gorskyi, O. parva, and O. longicauda (Flood, et al. 2005). These differences may reflect the changes in size and ploidy of OE cells. In O. dioica, OE cells stop mitotic division prior to metamorphosis and continue their growth by increasing the nuclear DNA content and size of nuclei. The nuclear morphologies change with the ploidy of OE cells as animals develop (Ganot and Thompson 2002). Therefore, despite our efforts to keep consistent staging throughout sample collection, it is possible that the observed differences in OE nuclei shape examined in this study could arise from differences in the developmental stages of individual specimens. It might be plausible that the numbers and the positions of the OE cells are invariable because they are responsible for the architecture of their house, which might directly affect their feeding ability and survival. On the other hand, the size of the OE cells and nuclear shapes are flexible because the nuclear DNA content of the OE cells increases as the animals grow. The crescent-shape nuclei we observed in the Osaka specimens may also be related to the size of the specimens.
The primary observation is that the number and arrangement of OE cells are highly conserved among genetically distinct lineages of O. dioica. Overall, the morphologies of the three O. dioica lineages are indistinguishable at a superficial level. Further morphometric comparisons are needed to evaluate whether there are morphological markers for reliable identification of O. dioica lineages in the field. The trunk shape, the size of Fol domain, and the taper between the dorsal epithelium over the mouth should be the first features to check in future morphometric work that will form an essential part of the process of describing a new species. It will be interesting to find out if such differences reflect normal variations between individuals, or if they correspond to lineage specific morphologies.
Reproductive isolation
Finally, we investigated the fertilization success of Okinawa and Osaka lab strains. Genetically distinct lineages of O. dioica from the Okinawa and Osaka lab strains do not interbreed under experimental conditions (Fig. 7). Reciprocal fertilization trials using individuals from these two strains always resulted in failure, suggesting strong prezygotic mechanisms against fertilization.
We observed the sperm of Okinawa and Osaka O. dioica gathered around the eggs of non-native strains, however, the eggs remained unfertilized. It is possible that the gamete recognition proteins in Okinawa and Osaka O. dioica are incompatible, preventing the entry of the sperm. Further studies will be needed to elucidate the underlying mechanism.
The intensity of this reproductive barrier between two relatively close geographical locations initially surprised us. It could be explained by the Kuroshio current—the subtropical western boundary current in the North Pacific that flows northwestwards along Okinawa Island and northeastwards along mainland Japan. The warm Kuroshio current transports many tropical and subtropical species to the north (Saito 2019). However, it can also act as a dispersal barrier, promoting lineage diversification in marine organisms (Kojima et al. 2000). Although appendicularians are present in the Kuroshio ecosystem (Saito 2019), O. dioica in particular is an inlet species and is most commonly found in bays and harbours (Tomita et al. 2003; Hidaka 2008). Therefore, even if a small population of O. dioica exists in the Kuroshio, the fast and strong meandering current at the Tokara gap (Fig. 1) may act as a potential dispersal barrier, limiting gene flow (Ogoh and Ohmiya 2005; Liu et al. 2008; Yamazaki et al. 2017) between Ryukyu and mainland Japanese O. dioica.
Implications and next steps
O. dioica is an important model organism for appendicularian and tunicate biology, partly owing to its global distribution, relative ease of sample collection, and ability to culture long-term in the laboratory. The original molecular comparisons between Norwegian and North Pacific specimens suggested a high level of DNA sequence conservation; however, more recent genomic and transcriptomic comparisons revealed more extensive divergence at the single nucleotide level. This study now confirms the impact of this divergence on the molecular phylogeny of different O. dioica specimens.
Studies have reported key developmental features that are not only common across European, Japanese and North American O. dioica, but also among chordates. In light of the high level of morphological conservation and similarities in developmental trajectories and pathways, many molecular, cellular and developmental biology discoveries in the different lineages will continue to apply across all O. dioica. Indeed, of great interest will be to understand the similarities and differences of distinct lineages, particularly in the context of their native marine environments.
Oikopleura is known to exhibit rapid protein evolution compared to other metazoans. The accelerated evolutionary rate may be coupled by the loss of some key components of DNA repair mechanisms, enhanced mutagenic environments of their habits (Denoeud et al. 2010), and their short life cycle. Considering these factors, it is perhaps not surprising to find a large degree of genetic diversity which could lead to more rapid reproductive isolation in O. dioica and related species.
Our work provides the first evidence that O. dioica might be hiding multiple cryptic species and establishes the basis for further investigations into the extent of O. dioica genetic diversity around the globe. The evidence suggests that at least the Ryukyu specimens should be considered a separate species to European and mainland Japanese O. dioica, which might require revision of current O. dioica taxonomy.
Our findings also raise interesting new questions to be tackled in future studies. What is the extent of geographical distributions of different lineages? Are different lineages sympatric or not? What is the level of genetic diversity within each lineage? Approaches targeting environmental DNA (eDNA) using ITS primers may provide molecular data that help us detect and distinguish O. dioica lineages. It is necessary to examine whether practical morphological characteristics could be discovered to aid field identification. We anticipate that the combined approach incorporating morphological and molecular markers will facilitate greater exploration into the diversity and biogeography of these ecologically important animals for ocean’s health.
Data availability
All sequence data have been deposited in the GenBank database (accession numbers provided in Supplementary Table S1). Raw data of our experiments are provided in the electronic supplementary files.
References
Alldredge A (1976) Appendicularians. Sci Am 235:94–105
Alldredge AL (1977) House morphology and mechanisms of feeding in the Oikopleuridae (Tunicata, Appendicularia). J Zool 181:175–188. https://doi.org/10.1111/j.1469-7998.1977.tb03236.x
Alldredge A (2005) The contribution of discarded appendicularian houses to the flux of particulate organic carbon from oceanic surface waters. In: Gorsky G, Youngbluth MJ, Deubel D (eds) Response of marine ecosystems to global change: ecological impact of appendicularians. Contemporary Publishing International, Paris, pp 309–326
Bickford D, Lohman DJ, Sodhi NS, Ng PKL, Meier R, Winker K, Ingram KK, Das I (2007) Cryptic species as a window on diversity and conservation. Trends Ecol Evol 22:148–155. https://doi.org/10.1016/j.tree.2006.11.004
Blanco-Bercial L, Cornils A, Copley N, Bucklin A (2014) DNA barcoding of marine copepods: assessment of analytical approaches to species identification. PLoS Curr 6:1–22. https://doi.org/10.1371/currents.tol.cdf8b74881f87e3b01d56b43791626d2
Bliznina A, Masunaga A, Mansfield MJ, Tan Y, Liu AW, West C, Rustagi T, Chien H-C, Kumar S, Pichon J, Plessy C, Luscombe NM (2021) Telomere-to-telomere assembly of the genome of an individual Oikopleura dioica from Okinawa using Nanopore-based sequencing. BMC Genomics 22:222. https://doi.org/10.1186/s12864-021-07512-6
Borsa P (2002) Allozyme, mitochondrial-DNA, and morphometric variability indicate cryptic species of anchovy (Engraulis encrasicolus). Biol J Linn Soc Lond 75:261–269. https://doi.org/10.1046/j.1095-8312.2002.00018.x
Bouquet J-M, Spriet E, Troedsson C, Otterå H, Chourrout D, Thompson EM (2009) Culture optimization for the emergent zooplanktonic model organism Oikopleura dioica. J Plankton Res 31:359–370. https://doi.org/10.1093/plankt/fbn132
Chen G, Hare MP (2011) Cryptic diversity and comparative phylogeography of the estuarine copepod Acartia tonsa on the US Atlantic coast. Mol Ecol 20:2425–2441. https://doi.org/10.1111/j.1365-294X.2011.05079.x
Clarke C, Roff JC (1990) Abundance and biomass of herbivorous zooplankton off Kingston, Jamaica, with estimates of their annual production. Estuar Coast Shelf Sci 31:423–437. https://doi.org/10.1016/0272-7714(90)90036-Q
Cracraft J (1983) Species Concepts and Speciation Analysis. In: Johnston RF (ed) Current Ornithology. Springer, New York, pp 159–187
Cronquist A (1978) Once again, what is a species? Biosystematics in agriculture. Beltsville Symposia in Agr Res 2:3–20
Dawson MN, Jacobs DK (2001) Molecular evidence for cryptic species of Aurelia aurita (Cnidaria, Scyphozoa). Biol Bull 200:92–96. https://doi.org/10.2307/1543089
de Vargas C, Norris R, Zaninetti L, Gibb SW, Pawlowski J (1999) Molecular evidence of cryptic speciation in planktonic foraminifers and their relation to oceanic provinces. Proc Natl Acad Sci USA 96:2864–2868. https://doi.org/10.1073/pnas.96.6.2864
Deng W, Henriet S, Chourrout D (2018) Prevalence of mutation-prone microhomology-mediated end joining in a chordate lacking the c-NHEJ DNA repair pathway. Curr Biol 28:3337-3341.e4. https://doi.org/10.1016/j.cub.2018.08.048
Denoeud F, Henriet S, Mungpakdee S, Aury J-M, Da Silva C, Brinkmann H, Mikhaleva J, Olsen LC, Jubin C, Cañestro C, Bouquet J-M, Danks G, Poulain J, Campsteijn C, Adamski M, Cross I, Yadetie F, Muffato M, Louis A, Butcher S, Tsagkogeorga G, Konrad A, Singh S, Jensen MF, Huynh Cong E, Eikeseth-Otteraa H, Noel B, Anthouard V, Porcel BM, Kachouri-Lafond R, Nishino A, Ugolini M, Chourrout P, Nishida H, Aasland R, Huzurbazar S, Westhof E, Delsuc F, Lehrach H, Reinhardt R, Weissenbach J, Roy SW, Artiguenave F, Postlethwait JH, Manak JR, Thompson EM, Jaillon O, Du Pasquier L, Boudinot P, Liberles DA, Volff J-N, Philippe H, Lenhard B, Roest Crollius H, Wincker P, Chourrout D (2010) Plasticity of animal genome architecture unmasked by rapid evolution of a pelagic tunicate. Science 330:1381–1385. https://doi.org/10.1126/science.1194167
Dyomin A, Volodkina V, Koshel E, Galkina S, Saifitdinova A, Gaginskaya E (2017) Evolution of ribosomal internal transcribed spacers in Deuterostomia. Mol Phylogenet Evol 116:87–96. https://doi.org/10.1016/j.ympev.2017.08.015
Essenberg CE (1922) The seasonal distribution of the appendicularia in the region of san Diego, California. Ecology 3:55–64. https://doi.org/10.2307/1929090
Ferrández-Roldán A, Fabregà-Torrus M, Sánchez-Serna G, Duran-Bello E, Joaquín-Lluís M, Bujosa P, Plana-Carmona M, Garcia-Fernàndez J, Albalat R, Cañestro C (2021) Cardiopharyngeal deconstruction and ancestral tunicate sessility. Nature 599:431–435. https://doi.org/10.1038/s41586-021-04041-w
Flood, (2005) Toward a photographic atlas on special taxonomic characters of oikopleurid appendicularia (Tunicata). In: Gorsky G, Youngbluth MJ, Deubel D (eds) Response of marine ecosystems to global change: ecological impact of appendicularians. Contemporary Publishing International, Paris, pp 59–85
Fol H (1872) Etudes sur les appendiculaires du détroit de Messine. Mem De La Soc De Phys Et D’histoire Nat De Genève 21:445–499
Fredriksson G, Olsson R (1991) The subchordal cells of Oikopleura dioica and O. albicans (Appendicularia, Chordata). Acta Zool 72:251–256. https://doi.org/10.1111/j.1463-6395.1991.tb01203.x
Ganot P, Thompson EM (2002) Patterning through differential endoreduplication in epithelial organogenesis of the chordate, Oikopleura dioica. Dev Biol 252:59–71. https://doi.org/10.1006/dbio.2002.0834
Garić R, Batistić M (2016) Description of Brooksia lacromae sp. nov. (Tunicata, Thaliacea) from the Adriatic Sea. EJT 196:1–13. https://doi.org/10.5852/ejt.2016.196
Gorsky G, Castellani C (2017) Chordata: Appendicularia. In: Castellani C, Edwards M (eds) Marine Plankton. A Practical Guide to Ecology, Methodology, and Taxonomy, 1st edn. Oxford University Press, Oxford, pp 599–606
Hamner R (1992) In situ observations of giant appendicularians in Monterey Bay. Deep-Sea Res i: Oceanogr Res Pap 39:1299–1313. https://doi.org/10.1016/0198-0149(92)90070-A
Hidaka, (2008) Species composition and horizontal distribution of the appendicularian community in waters adjacent to the Kuroshio in winter–early spring. Plankton Benthos Res 3:152–164. https://doi.org/10.3800/pbr.3.152
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Hopcroft R (1999) A new mesopelagic larvacean, Mesochordaeus erythrocephalus, sp. nov., from Monterey Bay, with a description of its filtering house. J Plankton Res 21:1923–1937
Hopcroft RR, Roff JC (1995) Zooplankton growth rates: extraordinary production by the larvacean Oikopleura dioica in tropical waters. J Plankton Res 17:205–220. https://doi.org/10.1093/plankt/17.2.205
Hopcroft, (2005) Diversity in larvaceans: How many species. In: Gorsky G, Youngbluth MJ, Deubel D (eds) Response of marine ecosystems to global change: ecological impact of appendicularians. Contemporary Publishing International, Paris, pp 45–57
Hwang UW, Kim W (1999) General properties and phylogenetic utilities of nuclear ribosomal DNA and mitochondrial DNA commonly used in molecular systematics. Korean J Parasitol 37:215–228. https://doi.org/10.3347/kjp.1999.37.4.215
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Katija K, Sherlock RE, Sherman AD, Robison BH (2017) New technology reveals the role of giant larvaceans in oceanic carbon cycling. Sci Adv 3:e1602374. https://doi.org/10.1126/sciadv.1602374
Kishi K, Hayashi M, Onuma TA, Nishida H (2017) Patterning and morphogenesis of the intricate but stereotyped oikoplastic epidermis of the appendicularian, Oikopleura dioica. Dev Biol 428:245–257. https://doi.org/10.1016/j.ydbio.2017.06.008
Kojima S, Segawa R, Hayashi I (2000) Stability of the courses of the warm coastal currents along the kyushu island suggested by the population structure of the Japanese Turban Shell, Turbo (Batillus) Cornutus. J Oceanogr 56:601–604. https://doi.org/10.1023/A:1011113430343
Liu S-YV, Kokita T, Dai C-F (2008) Population genetic structure of the neon damselfish (Pomacentrus coelestis) in the northwestern Pacific Ocean. Mar Biol 154:745–753. https://doi.org/10.1007/s00227-008-0967-2
Lombard F, Renaud F, Sainsbury C, Sciandra A, Gorsky G (2009) Appendicularian ecophysiology I: food concentration dependent clearance rate, assimilation efficiency, growth and reproduction of Oikopleura dioica. J Mar Syst 78:606–616. https://doi.org/10.1016/j.jmarsys.2009.01.004
Martí-Solans J, Ferrández-Roldán A, Godoy-Marín H, Badia-Ramentol J, Torres-Aguila NP, Rodríguez-Marí A, Bouquet JM, Chourrout D, Thompson EM, Albalat R, Cañestro C (2015) Oikopleura dioica culturing made easy: a low-cost facility for an emerging animal model in EvoDevo. Genesis 53:183–193. https://doi.org/10.1002/dvg.22800
Masunaga A, Liu AW, Tan Y, Scott A, Luscombe NM (2020) Streamlined sampling and cultivation of the pelagic cosmopolitan Larvacean. J Vis Exp, Oikopleura dioica. https://doi.org/10.3791/61279
Mayr, (1942) Systematics and theOrigin of Species. Cambridge University Press, New York, pp 1–334
Nishida H (2008) Development of the appendicularian Oikopleura dioica: culture, genome, and cell lineages. Dev Growth Differ 50:S239–S256. https://doi.org/10.1111/j.1440-169X.2008.01035.x
Ogoh K, Ohmiya Y (2005) Biogeography of luminous marine ostracod driven irreversibly by the Japan current. Mol Biol Evol 22:1543–1545. https://doi.org/10.1093/molbev/msi155
Ohtsuka S, Onbé T (1989) Evidence of selective feeding on larvaceans by the pelagic copepod Candacia bipinnata (Calanoida: Candaciidae). J Plankton Res 11:869–872. https://doi.org/10.1093/plankt/11.4.869
Onuma TA, Nishida H (2021) Developmental biology of the larvacean Oikopleura dioica: Genome resources, functional screening, and imaging. Dev Growth Differ 64:67–82. https://doi.org/10.1111/dgd.12769
Paffenhöfer G-A (1973) The cultivation of an appendicularian through numerous generations. Mar Biol 22:183–185. https://doi.org/10.1007/BF00391782
Pichon J, Luscombe NM, Plessy C (2019) Widespread use of the “ascidian” mitochondrial genetic code in tunicates. F1000Res 8:2072. https://doi.org/10.12688/f1000research.21551.2
Purcell JE, Hopcroft RR, Kosobokova KN, Whitledge TE (2010) Distribution, abundance, and predation effects of epipelagic ctenophores and jellyfish in the western Arctic Ocean. Deep Sea Res Part II Top Stud Oceanogr 57:127–135. https://doi.org/10.1016/j.dsr2.2009.08.011
Quattro JM, Stoner DS, Driggers WB, Anderson CA, Priede KA, Hoppmann EC, Campbell NH, Duncan KM, Grady JM (2006) Genetic evidence of cryptic speciation within hammerhead sharks (Genus Sphyrna). Mar Biol 148:1143–1155. https://doi.org/10.1007/s00227-005-0151-x
R Core Team (2020) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing. https://www.R-project.org/. Accessed 5 July 2022
Robison BH, Reisenbichler KR, Sherlock RE (2005) Giant larvacean houses: rapid carbon transport to the deep sea floor. Science 308:1609–1611. https://doi.org/10.1126/science.1109104
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Saito H (2019) The Kuroshio: its recognition, scientific activities and emerging issues. Kuroshio Current. John Wiley & Sons Inc, Hoboken, pp 3–11
Sakaguchi SO, Ikuta T, Ogawa G, Yamane K, Shiga N, Kitazato H, Fujikura K, Takishita K (2017) Morphological identity of a taxonomically unassigned cytochrome c oxidase subunit I sequence from stomach contents of juvenile chum salmon determined using polymerase chain reaction. Fish Sci 83:757–765. https://doi.org/10.1007/s12562-017-1106-0
Sato R, Tanaka Y, Ishimaru T (2001) House Production by Oikopleura dioica (Tunicata, Appendicularia) Under Laboratory Conditions. J Plankton Res 23:415–423. https://doi.org/10.1093/plankt/23.4.415
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez J-Y, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682. https://doi.org/10.1038/nmeth.2019
Schlötterer C, Hauser MT, von Haeseler A, Tautz D (1994) Comparative evolutionary analysis of rDNA ITS regions in Drosophila. Mol Biol Evol 11:513–522. https://doi.org/10.1093/oxfordjournals.molbev.a040131
Shenkar N, Swalla BJ (2011) Global diversity of Ascidiacea. PLoS ONE 6:e20657. https://doi.org/10.1371/journal.pone.0020657
Sherlock RE, Walz KR, Schlining KL, Robison BH (2017) Morphology, ecology, and molecular biology of a new species of giant larvacean in the eastern North Pacific: Bathochordaeus mcnutti sp. nov. Mar Biol 164:20. https://doi.org/10.1007/s00227-016-3046-0
Shiganova T (2005) Changes in appendicularian Oikopleura dioica abundance caused by invasion of alien ctenophores in the Black Sea. J Mar Biol Assoc UK 85:477–494. https://doi.org/10.1017/S0025315405011410
Singer M, Berg P (1991) Genes And Genomes. University Science Books, Mill Valley, CA
Spada F, Steen H, Troedsson C, Kallesoe T, Spriet E, Mann M, Thompson EM (2001) Molecular patterning of the oikoplastic epithelium of the larvacean tunicate Oikopleura dioica. J Biol Chem 276:20624–20632. https://doi.org/10.1074/jbc.M100438200
Tokioka T (1955) General consideration on Japanese appendicularian fauna. Publ Seto Mar Biol Lab 4:251–261. https://doi.org/10.5134/174523
Tokioka T (1960) Studies on the distribution of appendicularians and some thaliaceans of the north pacific, with some morphological notes. Publ Seto Mar Biol Lab 8:351–443. https://doi.org/10.5134/174644
Tomita M, Shiga N, Ikeda T (2003) Seasonal occurrence and vertical distribution of appendicularians in Toyama Bay, southern Japan Sea. J Plankton Res 25:579–589. https://doi.org/10.1093/plankt/25.6.579
Trifinopoulos J, Nguyen L-T, von Haeseler A, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44:W232–W235. https://doi.org/10.1093/nar/gkw256
Troedsson C, Bouquet J-M, Lobon CM, Novac A, Nejstgaard JC, Dupont S, Bosak S, Jakobsen HH, Romanova N, Pankoke LM, Isla A, Dutz J, Sazhin AF, Thompson EM (2013) Effects of ocean acidification, temperature and nutrient regimes on the appendicularian Oikopleura dioica: a mesocosm study. Mar Biol 160:2175–2187. https://doi.org/10.1007/s00227-012-2137-9
Uye S-I, Ichino S (1995) Seasonal variations in abundance, size composition, biomass and production rate of Oikopleura dioica (Fol) (Tunicata: Appendicularia) in a temperate eutrophic inlet. J Exp Mar Bio Ecol 189:1–11. https://doi.org/10.1016/0022-0981(95)00004-B
Wang K, Omotezako T, Kishi K, Nishida H, Onuma TA (2015a) Maternal and zygotic transcriptomes in the appendicularian, Oikopleura dioica: novel protein-encoding genes, intra-species sequence variations, and trans-spliced RNA leader. Dev Genes Evol 225:149–159. https://doi.org/10.1007/s00427-015-0502-7
Wang X-C, Liu C, Huang L, Bengtsson-Palme J, Chen H, Zhang J-H, Cai D, Li J-Q (2015b) ITS1: a DNA barcode better than ITS2 in eukaryotes? Mol Ecol Resour 15:573–586. https://doi.org/10.1111/1755-0998.12325
Wang K, Tomura R, Chen W, Kiyooka M, Ishizaki H, Aizu T, Minakuchi Y, Seki M, Suzuki Y, Omotezako T, Suyama R, Masunaga A, Plessy C, Luscombe NM, Dantec C, Lemaire P, Itoh T, Toyoda A, Nishida H, Onuma TA (2020) A genome database for a Japanese population of the larvacean Oikopleura dioica. Dev Growth Differ 62:450–461. https://doi.org/10.1111/dgd.12689
Yamazaki D, Miura O, Ikeda M, Kijima A, Van Tu D, Sasaki T, Chiba S (2017) Genetic diversification of intertidal gastropoda in an archipelago: the effects of islands, oceanic currents, and ecology. Mar Biol 164:184. https://doi.org/10.1007/s00227-017-3207-9
Yao H, Song J, Liu C, Luo K, Han J, Li Y, Pang X, Xu H, Zhu Y, Xiao P, Chen S (2010) Use of ITS2 region as the universal DNA barcode for plants and animals. PLoS ONE. https://doi.org/10.1371/journal.pone.0013102
Zagoskin MV, Lazareva VI, Grishanin AK, Mukha DV (2014) Phylogenetic information content of Copepoda ribosomal DNA repeat units: ITS1 and ITS2 impact. Biomed Res Int 2014:926342. https://doi.org/10.1155/2014/926342
Acknowledgements
We are grateful to Hiroki Nishida, Atsuo Nishino, Kouhei Hirose, Shouchi Nishino, for sending Oikopleura samples to us. We are thankful to Rade Garić for the advice and suggestion on taxonomy and morphological measurements. We thank Shinnosuke Teruya for allowing us to use the Deep Sea water research center on Kume Island. We are also thankful to the Kujukushima Aquarium Umi Kirara staff, especially Hisashi Akiyama, for their effort to identify and collect O. dioica for us. We thank Jai Denton and Jeffrey Jolly for their support and encouragement, especially during the early phase of the project. We thank Shinya Komoto from the OIST Imaging Section for assisting with Oikopleura epithelium imaging. Finally, we thank the Okinawa Institute of Science and Technology Graduate University (OIST) for funding, the DNA Sequencing Section and the Scientific Computing and Data Analysis Section of the Research Support Division at OIST, and the OIST Fieldwork Safety Committee for advice on safe sampling procedures. M.J.M. gratefully acknowledges funding from the Japan Society for the Promotion of Science as a JSPS International Research Fellow (Luscombe Unit, Okinawa Institute of Science and Technology Graduate University)
Funding
This work was supported by the Okinawa Institute of Science and Technology Graduate University. C.C. was supported by PID2019-110562GB-I00 grant from Ministerio de Ciencia y Innovación (Spain) and 2017-SGR-1665 from Generalitat de Catalunya; A.F-R. was supported by MS12 Margarita Salas program from Ministerio de Universidades (Spain).
Author information
Authors and Affiliations
Contributions
Aki Masunaga, Charles Plessy, Michael Mansfield, and Nicholas Luscombe contributed to the study conception and design. Material preparation was performed by Aki Masunaga, Yongkai Tan, Andrew Liu, Aleksandra Bliznina, and Takeshi Onuma. Data collection was performed by Aki Masunaga, Yongkai Tan, Paolo Barzaghi, Cristian Cañestro and Alfonso Ferrández-Roldán. Data analysis was performed by Aki Masunaga, Tamara Hodgetts, Michael Mansfield, and Charles Plessy. The first draft of the manuscript was written by Aki Masunaga and all authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
All authors included in this study declare that they have no conflict of interest.
Ethical approval
All applicable national and institutional guidelines for the care and use of animals were followed. This article does not contain any studies with human participants. All appendicularian specimen collections were approved by the OIST Fieldwork Safety Committee.
Additional information
Responsible Editor: Ulrich Sommer.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Masunaga, A., Mansfield, M.J., Tan, Y. et al. The cosmopolitan appendicularian Oikopleura dioica reveals hidden genetic diversity around the globe. Mar Biol 169, 157 (2022). https://doi.org/10.1007/s00227-022-04145-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00227-022-04145-5