Abstract
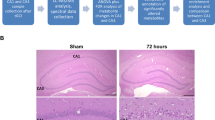
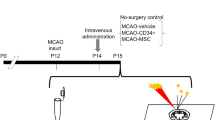
The N-methyl-D-aspartate (NMDA) receptor is a crucial mediator of pathological glutamate-driven excitotoxicity and subsequent neuronal death in acute ischemic stroke. Although the roles of the NMDAR’s composite GluN2A-C subunits have been investigated in this phenomenon, the relative importance of the GluN2D subunit has yet to be evaluated. Herein, GluN2D−/− mice were studied in a model of ischemic stroke using MALDI FT-ICR mass spectrometry imaging to investigate the role of the GluN2D subunit of the NMDA receptor in brain ischemia. GluN2D−/− mice underwent middle cerebral artery occlusion (MCAO) and brain tissue was subsequently harvested, frozen, and cryosectioned. Tissue sections were analyzed via MALDI FT-ICR mass spectrometry imaging. MALDI analyses revealed increases in several calcium-related species, namely vitamin D metabolites, LysoPC, and several PS species, in wild-type mouse brain tissue when compared to wild type. In addition, GluN2D−/− mice also displayed an increase in PC, as well as a decrease in DG, suggesting reduced free fatty acid release from brain ischemia. These trends indicate that GluN2D−/− mice show enhanced rates of neurorecovery and neuroprotection from ischemic strokes compared to wild-type mice. The cause of neuroprotection may be the result of an increase in PGP in knockout mice, contributing to greater cardiolipin synthesis and decreased sensitivity to apoptotic signals.

Graphical abstract







Similar content being viewed by others
References
Paoletti P, Bellone C, Zhou Q. NMDA receptor subunit diversity: impact on receptor properties, synaptic plasticity and disease. Nat Rev Neurosci. 2013;14:383–400.
Nakazawa K, McHugh TJ, Wilson MA, Tonegawa S. NMDA receptors, place cells and hippocampal spatial memory. Nat Rev Neurosci. 2004;5:361–72.
Dingledine R, Borges K, Bowie D, Traynelis SF. The glutamate receptor ion channels. Pharmacol Rev. 1999;51:7–61.
Traynelis SF, Wollmuth LP, McBain CJ, Menniti FS, Vance KM, Ogden KK, et al. Glutamate receptor ion channels: structure, regulation, and function. Pharmacol Rev. 2010;62:405–96.
Cheriyan J, Balsara RD, Hansen KB, Casetellino FJ. Pharmacology of triheteromeric N-methyl-d-aspartate receptors. Neurosci Lett. 2016;617:240–6.
Monyer H, Burnashev N, Laurie DJ, Sakmann B, Seeburg PH. Developmental and regional expression in the rat brain and functional properties of four NMDA receptors. Neuron. 1994;12:529–40.
Cull-Candy SG, Leszkiewicz DN. Role of distinct NMDA receptor subtypes at central synapses. Science STKE. 2004;re16.
Sheng M, Cummings J, Roldan LA, Jan YN, Jan LY. Changing subunit composition of heteromeric NMDA receptors during development of rat cortex. Nature. 1994;368:144–7.
Gray JA, Shi Y, Usui H, During MJ, Sakimura K, Nicoll RA. Distinct modes of AMPA receptor suppression at developing synapses by GluN2A and GluN2B: single-cell NMDA receptor subunit deletion in vivo. Neuron. 2011;71:1085–101.
Holmes A, Zhou N, Donahue DL, Balsara R, Castellino FJ. A deficiency of the GluN2C subunit of the N-methyl-D-aspartate receptor is neuroprotective in a mouse model of ischemic stroke. Biochem Biophys Res Commun. 2018;495:136–44.
Dudley E. MALDI profiling and applications in medicine. Adv Exp Med Biol. 2019;1140:27–43.
Ahmen M, Broeck G, Baggerman G, Schildermans K, Pauwels P, Van Craenenbroeck AH, et al. Next-generation protein analysis in the pathology department. J Clin Pathol. 2019. https://doi.org/10.1136/jclinpath-2019-205864.
Berghmans E, Van Raemdonck G, Schildermans K, Willems H, Boonen K, Maes E, et al. MALDI mass spectrometry imaging linked with top-down proteomics as a tool to study the non-small-cell lung cancer tumor microenvironment. Methods and Protocols. 2019;2:44.
Andrews WT, Skube SB, Hummon AB. Magnetic bead-based peptide extraction methodology for tissue imaging. Analyst. 2017;143:133–40.
Mallah K, Quanico J, Raffo-Romero A, Cardon T, Aboulouard S, Devos D, et al. Matrix-assisted laser desorption/ionization-mass spectrometry imaging of lipids in experimental model of traumatic brain injury detecting acylcaritines as injury related markers. Anal Chem. 2019. https://doi.org/10.1021/acs.analchem.9b02633.
Tobias F, Olson MT, Cologna SM. Mass spectrometry imaging of lipids: untargeted consensus spectra reveal spatial distributions in Niemann-Pick disease type C1. J Lipid Res. 2018;12:2446–55.
Liu X, Flinders C, Mumenthaler SM, Hummon AB. MALDI mass spectrometry imaging for evaluation of therapeutics in colorectal tumor organoids. J Am Soc Mass Spectrom. 2018;29:516–26.
Liu X, Hummon AB. Chemical imaging of platinum-based drugs and their metabolites. Sci Rep. 2016. https://doi.org/10.1038/srep38507.
Eveque-Mourroux MR, Emans PJ, Zautsen RRM, Boonen A, Heeren RMA, Cilero-Pastor B. Spatially resolved endogenous improved metabolite detection in human osteoarthritis cartilage by matrix assisted laser desorption ionization mass spectrometry imaging. Analyst. 2019. https://doi.org/10.1039/c9an00944b.
Chughtai K, Heeren RMA. Mass spectrometric imaging for biomedical tissue analysis. Chem Rev. 2011;110:3237–77.
Stoeckli M, Chaurand P, Hallahan DE, Caprioli RM. Imaging mass spectrometry: a new technology for the analysis of protein expression in mammalian tissues. Nat Med. 2001;7:493–6.
Palmer A, Phapale P, Chernyavsky I, Lavigne R, Fay D, Tarasov A, et al. FDR-controlled metabolite annotation for high-resolution imaging mass spectrometry. Nat Methods. 2016;14:57–60.
Alexandrov T, Ovchnnikova K, Palmer AJ, Kovalev V, Tarasov A, Stuart L, Nigmetzianov R, Fay D. METASPACE: A community-populated knowledge base of spatial metabolomes in health and disease. BioRxiv. 2019. https://doi.org/10.1101/539478.
Ellis SR, Paine MRL, Eijkel GB, Pauling JK, Husen P, Jervelund MW, et al. Automated, parallel mass spectrometry imaging and structural identification of lipids. Nat Methods. 2018;5:515–8.
Vaysse PM, Heeren RMA, Porta T, Balluff B. Mass spectrometry imaging for clinical research-latest developments, applications, and current limitations. Analyst. 2017;142:2690–712.
Gemperline E, Horn HA, DeLaney K, Currie CR, Li L. Imaging with mass spectrometry of bacteria on the exoskeleton of fungus-growing ants. ACS Chem Biol. 2017;12(8):1980–5.
Duenas ME, Essner JJ, Lee YJ. 3D MALDI mass spectrometry imaging of a single cell: spatial mapping of lipids in the embryonic development of zebrafish. Sci Rep. 2017;7:14946.
Summer LW, Amberg A, Barrett D, Beale MH, Beger R, Daykin CA, et al. Proposed minimum reporting standards for chemical analysis Chemical Analysis Working Group (CAWG) Metabolomics Standards Initiative (MSI). Metabolomics. 2007;3(3):211–21.
Falkenburger BH, Jensen JB, Dickson EJ, Suh BC, Hille B. Phosphoinositides: lipid regulators of membrane proteins. J Physiol. 2010;588:3179–85.
Heuser D, Guggenberger H. Ionic changes in brain ischemia and alterations produced by drugs. Br J Anesthesia. 1985;57(1):22–33.
White BC, Wiegenstein JG, Winegar CD. Brain ischemia and anoxia: mechanisms of injury. JAMA. 1984;251:158–690.
Farber JL, Chien KR, Mittnacht S Jr. Myocardial ischemia: the pathogenesis of irreversible cell injury in ischemia. Am J Pathol. 1981;102(2):271–81.
Blunt JW, DeLuca HF. The synthesis of 25-hydroxycholecalciferol. A biologically active metabolite of vitamin D3. Biochemistry. 1969;8(2):671–5.
Kopic S, Geibel JP. Gastric acid, calcium absorption, and their impact of bone health. Physiol Rev. 2013;93(1):189–268.
Sturkie PD. Hormonal Regulation of Calcium Metabolism. Basic Physiology. New York, NY: Springer-Verlag New York Incorporated; 1981. p. 414–8.
Borah M, Dhar S, Gogoi DM, Ruram AA. Association of serum calcium levels with infarct size in acute ischemic stroke: observations from northeast India. J Neurosci Rural Pract. 2016;7(S1):S41–5.
Canning P, Kenny BA, Prise V, Glenn J, Sarker MH, Hudson N, et al. Lipoprotein-associated phospholipase A2 (Lp-PLA2) as a therapeutic target to prevent retinal vasopermeability during diabetes. Proc Natl Acad Sci U S A. 2016;113(26):7213–8.
Law SH, Chan ML, Marathe GK, Parveen F, Chen CH, Ke LY. An updated review of lysophosphatidylcholine metabolism in human diseases. Int J Mol. 2019;20(5):1149.
Zhou F, Liu Y, Huang Q, Zhou J. Relation between lipoprotein-associated phospholipase A2 mass and incident ischemic stroke severity. Neurol Sci. 2018;39(9):1591–6.
Wang Y, Hu S, Ren L, Lan T, Cai J, Li C. Lp-PLA2 as a risk factor of early neurological deterioration in acute ischemic stroke with TOAST type of large arterial atherosclerosis. Neurol Res. 2019;41(1):1–8.
Ding CY, Cai HP, Ge HL, Yu LH, Lin YX, Kang DZ. Assessment of lipoprotein-associated phospholipase A2 level and its changes in the early stages as predictors of delayed cerebral ischemia in patients with aneurysmal subarachnoid hemorrhage. J Neurosurg. 2019:1–7.
Suzuki J, Fujii T, Imao T, Ishihara K, Kuba H, Nagata S. Calcium-dependent phospholipid scramblase activity of TMEM16 protein family members. J Biol Chem. 2013;288(19):13305–16.
Bull RK, Jevons S, Barton PG. Complexes of prothrombin with calcium ions and phospholipids. J Biol Chem. 1972;247(9):2747–54.
Vallabhapurapu SD, Blanco VM, Sulaiman MK, Vallabhapurapu SL, Chu Z, Franco RS, Qi X. Variation in human cancer cell external phosphatidylserine is regulated by flippase activity and intracellular calcium. Oncotarget. 2015;6(33):34375–88. https://doi.org/10.18632/oncotarget.6045.
Martin-Molina A, Rodriguez-Beas C, Faraudo J. Effect of calcium and magnesium on phosphatidylserine membranes: experiments and all-atomic simulations. Biophys J. 2012;102(9):2095–103.
Aussel C, Pelassy C, Mary D, Breittmayer JP, Cousin JL, Rossi B. Calcium-dependent regulation of phosphatidylserine synthesis in control and activated Jurkat T cells. J Lipid Mediat Cell Signal. 1991;3(3):267–81.
Zinrajh D, Horl G, Jurgens G, Marc J, Sok M, Cerne C. Increased phosphatidylethanolamine N-methyltransferase gene expression in non-small-cell lung cancer tissue predicts shorter patient survival. Oncol Lett. 2014;7(6):2175–9.
Muralikrishna Adibhatla R, Hatcher JF, Larsen EC, Chen X, Tsao FHC. CDP-choline significantly restores phosphatidylcholine levels by differentially affecting phospholipase A2 and CTP: phosphocholine cytidylyltransferase after stroke. J Biol Chem. 2005;281:6718–25.
Nishida A, Emoto K, Shimizu M, Uozumi T, Yamawaki S. Brain ischemia decreases phosphatidylcholine-phospholipase D but not phosphatidylinositol-phospholipase C in rats. Stroke. 1994;25(6):1247–51.
Sabogal-Guaqueta AM, Villamil-Ortiz JG, Arias-Londono JD, Cardona-Gomez GP. Inverse phosphatidylcholine/phosphatidylinositol levels as peripheral biomarkers and phosphatidylcholine/lysophosphoethanolamine-phosphatidylserine as hippocampal indicator of postischemic cognitive impairment in rats. Front Neurosci. 2018;21. https://doi.org/10.3389/fnins.2018.00989.
Mousavi SA, Khorvash F, Hoseini T. The efficacy of citroline in the treatment of ischemic stroke and primary hypertensive intracereal hemorrhage; a review article. ARYA Atherosclerosis. 2010;6(3):122–5.
Moto A, Hirashima Y, Endo S, Takaku A. Changes in lipid metabolites and enzymes in rat brain due to ischemia and recirculation. Mol Chem Neuropathol. 1991;14(1):35–51.
Hattori T, Nishimura Y, Sakai N, Yamada H, Kameyama Y, Nozawa Y. Effects of pentobarbital on brain lipid metabolism during global ischemia. Neurol Surg. 1986;38(6):585–91.
Matthys E, Patel Y, Kreisberg J, Stewart JH, Venkatachalam M. Lipid alterations induced by renal ischemia: pathogenic factor in membrane damage. Kidney Int. 1984;26(2):153–61.
Sun GY, Lu FL, Lin SE, Ko MR. Decapitation ischemia-induced release of free fatty acids in mouse brain. Relationship with diacylglycerols and lysophospholipids. Mol Chem Neuropathol. 1992;17(1):39–50.
van der Veen JN, Kennelly JP, Wan S, Vance JE, Vance DE, Jacobs RL. The critical role of phosphatidylcholine and phosphatidylethanolamine metabolism in health and disease. Biochim Biophys Acta Biomembr. 2017;1859(9):1558–72.
Nilanjana M, Kagan VE, Tyurin VA, Das DK. Redistribution of phosphatidylethanolamine and phosphatidylserine precedes reperfusion-induced apoptosis. Am J Physiol Heart Circ Physiol. 1998:H242–8.
Kawai H, Chaudhry F, Shekhar A, Petrov A, Nakahara T, Tanimoto T, et al. Molecular imaging of apoptosis in ischemia reperfusion injury with radiolabeled duramycin targeting phosphatidylethanolamine. Effective target uptake and reduced nontarget organ radioation burden. J Am Coll Cardiol Img. 2018;11(12). https://doi.org/10.1016/j.jcmg.2017.11.037.
Schabitz WR, Giuffrida A, Berger C, Aschoff A, Schwaninger M, Schwab S, et al. Release of fatty acid amides in a patient with hemispheric stroke, a microdialysis study. Stroke. 2002;33:2112–4.
Post JA, Bivelt JJ, Verkleij AJ. Phosphatidylethanolamine and sarcolemmal damage during ischemia or metabolic inhibition of heart myocytes. Am J Physiol Heart Circ Physiol. 1995;268(2):H773–80.
Hirabayashi T, Larson TJ, Dowan W. Membrane-associated phosphatidylglycerophosphate synthetase from Escherichia coli: purification by substrate affinity chromatography on cytidine 5′-diphospho-1,2-diacyl-sn-glycerol sepharose. Biochemistry. 1976;15(24):5205–11.
Morita SY, Terada T. Enzymatic measurement of phosphadiylglycerol and cardiolipin in cultured cells and mitochondria. Sci Rep. 2015;5. https://doi.org/10.1038/srep11737.
Gebert N, Joshi AS, Kutik S, Becker T, McKenzie M, Guan XL, et al. Mitochondrial cardiolipin involved in outer-membrane protein biogenesis: implications for Barth syndrome. Curr Biol. 2009;19(24):2133–9.
Osman C, Haag M, Wieland FT, Brugger B, Langer T. A mitochondrial phosphatase required for cardiolipin biosynthesis: the PGP phosphatase Gep4. EMBO J. 2010;29(12):1976–87.
Gross A, Yin XM, Wang K, Wei MC, Jockel J, Milliman C, et al. Caspase cleaved BID targets mitochondria and is required for cytochrome c release, while BCL-XL prevents this release but not tumor necrosis factor-R1/Fas death. J Biol Chem. 1999;274(2):1156–63.
Orrenius S, Zhivotovsky B. Cardiolipin oxidation sets cytochrome c free. Nat Chem Biol. 2005;1(4):188–9.
Choi SY, Gonzalvez F, Jenkins GM, Slominanny C, Chretien D, Arnoult D, et al. Cardiolipin deficiency releases cytochrome c from the inner mitochondrial membrane and accelerates stimuli-elicited apoptosis. Cell Death Differ. 2007;14:597–606.
Acknowledgments
ABH was supported by NIGMS award R01-GM110406 and WTA was supported by NIA R21-AG062144. The authors would like to acknowledge the Campus Chemical Instrument Center at the Ohio State University for providing instrument usage, as well as Arpad Somogyi for helpful discussions and support. The 15 T Bruker SolariX FT-ICR instrument was supported by NIH Award Number Grant S10 OD018507.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Published in the topical collection featuring Female Role Models in Analytical Chemistry.
Electronic supplementary material
ESM 1
(PDF 1.06 mb)
Rights and permissions
About this article
Cite this article
Andrews, W.T., Donahue, D., Holmes, A. et al. In situ metabolite and lipid analysis of GluN2D−/− and wild-type mice after ischemic stroke using MALDI MSI. Anal Bioanal Chem 412, 6275–6285 (2020). https://doi.org/10.1007/s00216-020-02477-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-020-02477-z