Abstract
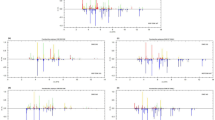
An untargeted multi-criteria approach was used to select the best extraction method among freeze-thawing in methanol (FTM), boiling ethanol (BE) and chloroform-methanol (CM) for gas chromatography mass spectrometry (GC-MS) metabolic fingerprinting of Lactobacillus paracasei subsp. paracasei (CRL-431®). The following results were obtained: (i) coverage and efficiency, measured by the number of features extracted and the sum of feature intensities, showed that FTM extraction resulted in the largest compound coverage with a total number of features 8.9 × 103 ± 0.5 × 103, while merely 6.6 × 103 ± 0.9 × 103 and 7.9 × 103 ± 0.8 × 103 were detected in BE or CM, respectively; (ii) the similarity of extraction methods, measured by common features, demonstrated that FTM yielded the most complementary information to BE and CM; i.e. 17 and 33 % of the features of FTM extracted were unique compared to CM and BE, respectively; and (iii) a clear-cut separation according to extraction method was demonstrated by assessment of the metabolic fingerprints by pixel-based data analysis. Indications of metabolite degradation were observed under the elevated temperature for BE extraction. A superior coverage of FTM together with a high repeatability over nearly the whole range of GC-amenable compounds makes this the extraction method of choice for metabolic fingerprinting of L. paracasei.




Similar content being viewed by others
References
Ruiz L, Margolles A, Sanchez B (2013) Bile resistance mechanisms in Lactobacillus and Bifidobacterium. Front Microbiol 4:396
Duportet X, Aggio RBM, Carneiro S, Villas-Boas SG (2012) The biological interpretation of metabolomic data can be misled by the extraction method used. Metabolomics 8:410
De Koning W, Van Dam K (1992) A method for the determination of changes of glycolytic metabolites in yeast on a subsecond time scale using extraction at neutral pH. Anal Biochem 204:118
Gonzalez B, Francois J, Renaud M (1997) A rapid and reliable method for metabolite extraction in yeast using boiling buffered ethanol. Yeast 13:1347
Hiller J, Franco-Lara E, Weuster-Botz D (2007) Metabolic profiling of Escherichia coli cultivations: evaluation of extraction and metabolite analysis procedures. Biotechnol Lett 29:1169
Winder CL, Dunn WB, Schuler S, Broadhurst D, Jarvis R, Stephens GM, Goodacre R (2008) Global metabolic profiling of Escherichia coli cultures: an evaluation of methods for quenching and extraction of intracellular metabolites. Anal Chem 80:2939
Maharjan R (2003) Global metabolite analysis: the influence of extraction methodology on metabolome profiles of Escherichia coli. Anal Biochem 313:145
Faijes M, Mars AE, Smid EJ (2007) Comparison of quenching and extraction methodologies for metabolome analysis of Lactobacillus plantarum. Microb Cell Factories 6:27
Shin MH, Lee DY, Liu K, Fiehn O, Kim KH (2010) Evaluation of sampling and extraction methodologies for the global metabolic profiling of Saccharophagus degradans. Anal Chem 82:6660
Kim S, Lee DY, Wohlgemuth G, Park HS, Fiehn O, Kim KH (2013) Evaluation and optimization of metabolome sample preparation methods for Saccharomyces cerevisiae. Anal Chem 85:2169
Jernejc K (2004) Comparison of different methods for metabolite extraction from Aspergillus niger mycelium. Acta Chim Slov 51:567
Li X, Long D, Ji J, Yang W, Zeng Z, Guo S, Ji Z, Qi G, Chen S (2013) Sample preparation for the metabolomics investigation of poly-gamma-glutamate-producing Bacillus licheniformis by GC-MS. J Microbiol Methods 94:61
Meyer H, Liebeke M, Lalk M (2010) A protocol for the investigation of the intracellular Staphylococcus aureus metabolome. Anal Biochem 401:250
Park C, Yun S, Lee SY, Park K, Lee J (2012) Metabolic profiling of klebsiella oxytoca: evaluation of methods for extraction of intracellular metabolites using UPLC/Q-TOF-MS. Appl Biochem Biotechnol 167:425
Canelas AB, ten Pierick A, Ras C, Seifar RM, van Dam JC, van Gulik WM, Heijnen JJ (2009) Quantitative evaluation of intracellular metabolite extraction techniques for yeast metabolomics. Anal Chem 81:7379
Rabinowitz JD, Kimball E (2007) Acidic acetonitrile for cellular metabolome extraction from Escherichia coli. Anal Chem 79:6167
Villas-Boas S, Hojer-Pedersen J, Akesson M, Smedsgaard J, Nielsen J (2005) Global metabolite analysis of yeast: evaluation of sample preparation methods. Yeast 22:1155
van der Werf MJ, Overkamp KM, Muilwijk B, Coulier L, Hankemeier T (2007) Microbial metabolomics: toward a platform with full metabolome coverage. Anal Biochem 370:17
Putri SP, Yamamoto S, Tsugawa H, Fukusaki E (2013) Current metabolomics: technological advances. J Biosci Bioeng 116:9
Honore AH, Thorsen M, Skov T (2013) Liquid chromatography-mass spectrometry for metabolic footprinting of co-cultures of lactic and propionic acid bacteria. Anal Bioanal Chem 405:8151
Mashego MR, Rumbold K, De Mey M, Vandamme E, Soetaert W, Heijnen JJ (2007) Microbial metabolomics: past, present and future methodologies. Biotechnol Lett 29:1
Amigo JM, Popielarz MJ, Callejon RM, Morales ML, Troncoso AM, Petersen MA, Toldam-Andersen TB (2010) Comprehensive analysis of chromatographic data by using PARAFAC2 and principal components analysis. J Chromatogr 1217:4422
Petersen IL, Tomasi G, Sørensen H, Boll ES, Hansen HCB, Christensen JH (2011) The use of environmental metabolomics to determine glyphosate level of exposure in rapeseed (Brassica napus L.) seedlings. Environ Pollut 159:3071
Smith C, Want E, O’Maille G, Abagyan R, Siuzdak G (2006) XCMS: processing mass spectrometry data for metabolite profiling using nonlinear peak alignment, matching, and identification. Anal Chem 78:779
Katajamaa M, Orešič M (2007) Data processing for mass spectrometry-based metabolomics. J Chromatogr A 1158:318
Lommen A (2009) MetAlign: interface-driven, versatile metabolomics tool for hyphenated full-scan mass spectrometry data preprocessing. Anal Chem 81:3079
Aggio R, Villas-Boas SG, Ruggiero K (2011) Metab: an R package for high-throughput analysis of metabolomics data generated by GC-MS. Bioinformatics 27:2316
Christensen JH, Tomasi G (2007) Practical aspects of chemometrics for oil spill fingerprinting. J Chromatogr A 1169:1
Danielsson R, Bylund D, Markides K (2002) Matched filtering with background suppression for improved quality of base peak chromatograms and mass spectra in liquid chromatography-mass spectrometry. Anal Chim Acta 454:167
Ullsten S, Danielsson R, Backstrom D, Sjoberg P, Bergquist J (2006) Urine profiling using capillary electrophoresis-mass spectrometry and multivariate data analysis. J Chromatogr A 1117:87
Christensen J, Mortensen J, Hansen A, Andersen O (2005) Chromatographic preprocessing of GC-MS data for analysis of complex chemical mixtures. J Chromatogr A 1062:113
Savitzky A, Golay M (1964) Smoothing + differentiation of data by simplified least squares procedures. Anal Chem 36:1627
Tomasi G, van den Berg F, Andersson C (2004) Correlation optimized warping and dynamic time warping as preprocessing methods for chromatographic data. J Chemom 18:231
Skov T, van den Berg F, Tomasi G, Bro R (2006) Automated alignment of chromatographic data. J Chemom 20:484
Christensen J, Tomasi G, Hansen A (2005) Chemical fingerprinting of petroleum biomarkers using time warping and PCA. Environ Sci Technol 39:255
Soleimani M, Farhoudi M, Christensen JH (2013) Chemometric assessment of enhanced bioremediation of oil contaminated soils. J Hazard Mater 254:372
Gallotta FDC, Christensen JH (2012) Source identification of petroleum hydrocarbons in soil and sediments from Iguaçu River Watershed, Paraná, Brazil using the CHEMSIC method (CHEMometric analysis of Selected Ion Chromatograms). J Chromatogr A 1235:149
Folch J, Lees M, Stanley G (1957) A simple method for the isolation and purification of total lipides from animal tissues. J Biol Chem 226:497
More N, Daniel R, Petach H (1995) The effect of low-temperatures on enzyme-activity. Biochem J 305:17
Pohekar SD, Ramachandran M (2004) Application of multi-criteria decision making to sustainable energy planning—a review. Renew Sust Energ Rev 8:365
Furbo S, Hansen AB, Skov T, Christensen JH (2014) Pixel-based analysis of comprehensive two-dimensional gas chromatograms (color plots) of petroleum: a tutorial. Anal Chem 86:7160
Acknowledgments
We thank Chr. Hansen A/S and University of Copenhagen for financial support.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jäpelt, K.B., Nielsen, N.J., Wiese, S. et al. Metabolic fingerprinting of Lactobacillus paracasei: a multi-criteria evaluation of methods for extraction of intracellular metabolites. Anal Bioanal Chem 407, 6095–6104 (2015). https://doi.org/10.1007/s00216-015-8783-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00216-015-8783-2