Abstract
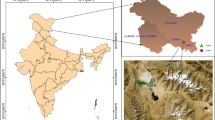
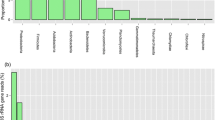
The Kyzyl-Kum Desert extends over an area of 300,000 Km2, in the region bordering Kazakhstan, Uzbekistan and Turkmenistan and is mainly covered by sand dunes. The Kyzyl-Kum desert is also known for its large deposits of minerals of economic interests, the exploitation of which is affecting the local ecosystem and increasing the desertification. We examined the bacterial biodiversity of surface sand samples from several sites from the Kyzyl-Kum desert using pyrosequencing of PCR amplified bacterial 16S rRNA genes from total extracted soil DNA. We also examined several physicochemical parameters of the sand samples to investigate any possible correlations between bacterial community structure and environmental drivers. The predominant bacterial phyla present in the samples were found to belong to members of the Actinobacteria, Proteobacteria and Bacteroidetes. The most abundant genera in our samples were found to belong to the Arthrobacter, Adhaeribacter and Roseomonas genera. We found that the relative abundance of members belonging to the Actinobacteria phylum, commonly found in desertic areas, increase in abundance in sites with higher content of organic matter and sulfur, whereas members of the Proteobacteria and Bacteroidetes phyla seems to diminish in abundance in coarse silt and fine-grained soils and those rich in magnesium, suggesting that those parameters might influence the bacterial community composition in this desertic area. This study is the first to provide new insights into the prokaryotic community composition from this unusual desert site.






Similar content being viewed by others
Data availability
The datasets generated during and/or analysed during the current study are available in the Short Read Archive (accession number: SRP074819).
References
Alivisatos AP, Blaser MJ, Brodie EL, Chun M, Dangl JL, Donohue TJ, Dorrestein PC, Gilbert JA, Green JL, Jansson JK, Knight R, Maxon ME, McFall-Ngai MJ, Miller JF, Pollard KS, Ruby EG, Taha SA (2015) Unified microbiome initiative consortium. A unified initiative to harness Earth’s microbiomes. Science 350:507–508. https://doi.org/10.1126/science.aac8480
Alsharif W, Saad MM, Hirt H (2020) Desert microbes for boosting sustainable agriculture in extreme environments. Front Microbiol 11:1666. https://doi.org/10.3389/fmicb.2020.01666
An S, Couteau C, Luo F, Neveu J, DuBow MS (2013) Bacterial diversity of surface sand samples from the Gobi and Taklamaken deserts. Microb Ecol 66:850–860. https://doi.org/10.1007/s00248-013-0276-2
An S, DuBow MS (2015) Modification of atmospheric sand-associated bacterial communities during Asian sandstorms in China and South Korea. Heredity 114:460–467
Andrews S (2010) FastQC: a quality control tool for high throughput sequence data. Available online at: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/
Bashan Y, Salazar BG, Moreno M, Lopez BR, Linderman RG (2012) Restoration of eroded soil in the Sonoran Desert with native leguminous trees using plant growth-promoting microorganisms and limited amounts of compost and water. J Environ Manage 102:26–36
Bi B, Yuan Y, Zhang H, Wu Z, Wang Y, Han F (2022) Rhizosphere soil metabolites mediated microbial community changes of Pinus Sylvestris var. mongolica across stand ages in the Mu Us Desert. Appl Soil Ecol 169:104222
Binladen J, Gilbert MT, Bollback JP, Panitz F, Bendixen C, Nielsen R, Willerslev E (2007) The use of coded PCR primers enables high-throughput sequencing of multiple homolog amplification products by 454 parallel sequencing. PLoS ONE 2:e197. https://doi.org/10.1371/journal.pone.0000197
Calleja-Cervantes ME, Menendez S, Fernandez-Gonzalez AJ, Irigoyen I, Cibri-ain-Sabalza JF, Toro N, Aparicio-Tejo PM, Fernandez-Lopez M (2015) Changes in soil nutrient content and bacterial community after 12 years oforganic amendment application to a vineyard. Eur J Soil Sci 66:802e812. https://doi.org/10.1111/ejss.12261
Carbonetto B, Rascovan N, Álvarez R, Mentaberry A, Vázquez MP (2014) Structure, Composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine pampas. PLoS ONE 9(6):e99949. https://doi.org/10.1371/journal.pone.0099949
Chuvochina MS, Marie D, Chevaillier S, Petit JR, Normand P, Alekhina IA, Bulat SA (2011) Community variability of bacteria in alpine snow (Mont Blanc) containing Saharan dust deposition and their snow colonisation potential. Microbes Environ 26:237–247. https://doi.org/10.1264/jsme2.me11116
Dubilier N, McFall-Ngai M, Zhao L (2015) Create a global microbiome effort. Nature 526:631–634. https://doi.org/10.1038/526631a
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Favet JL, Lapanje A, Giongo A, Kennedy S, Aung YY, Cattaneo A, Davis-Richardson AG, Brown CT, Kort R, Brumsack HJ, Schnetger B, Chappell A, Kroijenga J, Beck A, Schwibbert K, Mohamed AH, Kirchner T, de Quadros PD, Triplett EW, Broughton WJ, Gorbushina AA (2013) Microbial hitchhikers on intercontinental dust: catching a lift in Chad. ISME J 7:850–867. https://doi.org/10.1038/ismej.2012.152
Fierer N, Jackson RB (2006) The diversity and biogeography of soil bacterial communities. Proc Natl Acad Sci USA 103:626–631. https://doi.org/10.1073/pnas.0507535103
Goudie AS (2014) Desert dust and human health disorders. Environ Int 63:101–113. https://doi.org/10.1016/j.envint.2013.10.011
Gundlapally SR, Garcia-Pichel F (2006) The community and phylogenetic diversity of biological soil crusts in the Colorado Plateau studied by molecular fingerprinting and intensive cultivation. Microb Ecol 52:345–357. https://doi.org/10.1007/s00248-006-9011-6
Hamady M, Walker JJ, Harris JK, Gold NJ, Knight R (2008) Error correcting barcoded primers for pyrosequencing hundreds of samples in multiplex. Nat Methods 5:235–237. https://doi.org/10.1038/nmeth.1184
Indoitu R, Orlovsky L, Orlovsky N (2012) Dust storms in Central Asia: spatial and temporal variations. J Arid Environ 85:62–70. https://doi.org/10.2495/ECO090331
Indoitu R, Kozhoridze G, Batyrbaeva M, Vitkovskaya I, Orlovsky N, Blumberg D, Orlovsky, (2015) Dust emission and environmental changes in the dried bottom of the Aral Sea. Aeolian Res 17:101–115. https://doi.org/10.1016/j.aeolia.2015.02.004
Izhitskiy AS, Zavialov PO, Sapozhnikov PV, Kirillin GB, Grossart HP, Kalinina OY, Zalota AK, Goncharenko IV, Kurbaniyazov AK (2016) Present state of the Aral Sea: diverging physical and biological characteristics of the residual basins. SciRep 6:23906. https://doi.org/10.1038/srep23906
Laiz L, Miller AZ, Jurado V, Akatova E, Sanchez-Moral S, Gonzalez JM, Dionisio A, Macedo MF, Saiz-Jimenez C (2009) Isolation of five Rubrobacter strains from biodeteriorated monuments. Naturwissenschaften 96:71–79. https://doi.org/10.1007/s00114-008-0452-2
Lauber CL, Hamady M, Knight R, Fierer N (2009) Pyrosequencing-based assessment of soil pH as a predictor of soil bacterial community structure at the continental scale. Appl Environ Microbiol 75:5111–5120. https://doi.org/10.1128/AEM.00335-09
Limin Fu, Niu B, Zhu Z, Sitao Wu, Li W (2012) CD-HIT: accelerated for clustering the next generation sequencing data. Bioinformatics 28:3150–3152. https://doi.org/10.1093/bioinformatics/bts565
Love MI, Huber W, Anders S (2014) Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol 15:550. https://doi.org/10.1186/s13059-014-0550-8
McMurdie PJ, Holmes S (2013) Phyloseq: An R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
McMurdie PJ, Holmes S (2014) Waste not, want not: why rarefying microbiome data is inadmissible. PLoS Comput Biol 10:e1003531. https://doi.org/10.1371/journal.pcbi.1003531
Mukhtar S, Mehnaz S, Malik KA (2021) Comparative study of the rhizosphere and root endosphere microbiomes of Cholistan Desert plants. Front Microbiol 12:618742
Neilson JW, Quade J, Ortiz M, Nelson WM, Legatzki A, Tian F, LaComb M, Betancourt JL, Wing RA, Soderlund CA, Maier RM (2012) Life at the hyperarid margin: novel bacterial diversity in arid soils of the Atacama Desert. Chile Extremophiles 16:553–566. https://doi.org/10.1007/s00792-012-0454-z
Nemergut DR, Costello EK, Hamady M, Lozupone C, Jiang L, Schmidt SK, Fierer N, Townsend AR, CLeveland CC, Stanish L, Knight R, (2010) Global patterns in the biogeography of bacterial taxa. Environ Microbiol 13:135–144. https://doi.org/10.1111/j.1462-2920.2010.02315.x
Ofek M, Hadar Y, Minz D (2012) Ecology of root colonizing massilia (Oxalobacteraceae). PLoS ONE 7:e40117. https://doi.org/10.1371/journal.pone.0040117
Oksanen J, Blanchet FG, Kindt R, Legendre P, O’Hara RB et al (2011) Vegan: community ecology package. R Pack Version 1:17
Osman JR, Fernandes G, Regeard C, DuBow MS (2017) Bacterial diversity of the rhizosphere and nearby surface soil of rice (Oryza sativa) growing in the Camargue (France). Rhizosphere 3:112–122
Osman JR, Fernandes G, Regeard C, Jaubert C, DuBow MS (2018) Examination of the bacterial biodiversity of coastal eroded surface soils from the Padza de Dapani (Mayotte Island). Geomicrobiol J 35:355–365
Osman JR, Wang Y, Jaubert C, Nguyen TN, Fernandes GR, DuBow M (2021) The bacterial communities of surface soils from the desert sites in the eastern Utah (USA) portion of the Colorado Plateau. Microbiol Res 244:126664
Opp C, Groll M, Aslanov I, Lotz T, Vereshagina N (2016) Aeolian dust deposition in the southern Aral Sea region (Uztbekistan): Ground-based monitoring results from the LUCA project. Quat Int 429:86–99. https://doi.org/10.1016/j.quaint.2015.12.103
R Core Team (2022) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna, Austria
Ragon M, Restoux G, Moreira D, Møller AP, Lopez-Garcia P (2011) Sunlight-exposed biofilm microbial communities are naturally resistant to Chernobyl ionizing-radiation levels. PLoS ONE 6:e21764. https://doi.org/10.1371/journal.pone.0021764
Sabloff PLW (2011) Mapping mongolia: situating mongolia in the world from geologic time to the present. University of Pennsylvania Press. 10:1934536180
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Schmitz L, Yan Z, Schneijderberg M et al (2022) Synthetic bacterial community derived from a desert rhizosphere confers salt stress resilience to tomato in the presence of a soil microbiome. ISME J 16:1907–1920
Sghaier H, Hezbri K, Ghodhbane-Gtari F, Pujic P, Sen A, Daffonchio D, Gtari M (2016) Stone-dwelling actinobacteria Blastococcus saxobsidens, Modestobacter marinus and Geodermatophilus obscurus proteogenomes. ISME J 10:21–29. https://doi.org/10.1038/ismej.2015.108
Shen C, Xiong J, Zhang H, Feng Y, Lin X, Li X, Chu LW, H, (2013) Soil pH drives the spatial distribution of bacterial communities along elevation on Changbai Mountain. Soil Biol Biochem 57:204–211. https://doi.org/10.1016/j.soilbio.2012.07.013
Smeds L, Künstner A (2011) ConDeTri—a content dependent read trimmer for illumina data. PLoS ONE 6(10):e26314
Sun H, Zhang T, Yu L, Sen K, Zhang Y (2015) Ubiquity, diversity and physiological characteristics of geodermatophilaceae in shapotou national desert ecological reserve. Front Microbiol 6:1059. https://doi.org/10.3389/fmicb.2015.01059
Toderich KN, Tsukatani T, Black CC, Takabe K, Katayama Y (2002) Adaptations of plants to metal/salt contained environments: glandular structure and salt excretion. Discuss Paper. 2002:552
Touchon M, Hoede C, Tenaillon O, Barbe V, Baeriswyl S, Bidet P, Bingen E, Bonacorsi S, Bouchier C, Bouvet O, Calteau A, Chiapello H, Clermont O, Cruveiller S, Danchin A et al (2009) Organised genome dynamics in the escherichia coli species results in highly diverse adaptive paths. PLoS Genet 5(1):e1000344
Uksa M, Schloter M, Endesfelder D, Kublik S, Engel M, Kautz T, Köpke U, Fischer D (2015) Prokaryotes in subsoil evidence for a strong spatial separation of different phyla by analysing co-occurrence etworks. Front Microbiol 6:1269. https://doi.org/10.3389/fmicb.2015.01269
Uroz S, Calvaruso C, Turpault MP, Frey-Klett P (2009) Mineral weathering by bacteria: ecology, actors and mechanisms. Trends in Microbiol 17:378–387. https://doi.org/10.1016/j.tim.2009.05.004
Uroz S, Oger P, Lepleux C, Collignon C, Frey-Klett P, Turpault MP (2011) Bacterial weathering and its contribution to nutrient cycling in temperate forest ecosystems. Res Microbiol 9:820–831. https://doi.org/10.1016/j.resmic.2011.01.013
Vásquez-Dean J, Maza F, Morel I, Pulgar R, González M (2020) Microbial communities from arid environments on a global scale. Syst Rev Biol Res 53:29
Vu V (2015) ggbiplot: A ggplot2 based biplot. R package. http://github.com/vqv/ggbiplot
Weyant RS, Whitney AM (2015) Roseomonas. Bergey’s Manual of Systematics of Archaea and Bacteria. Wiley, New York USA, pp 1–9
Wright ES, Yilmaz LS, Noguera DR (2012) DECIPHER, a search-based approach to chimera identification for 16S rRNA sequences. Appl Environ Microbiol 78:717–725. https://doi.org/10.1128/AEM.06516-11
Yasir M, Azhar EI, Khan I, Bibi Baabdullah RF, Al-Zahrani IA, Al-Ghamdi AK (2015) Composition of soil microbiome along elevation gradients in southwestern highlands of Saudi Arabia. BMC Microbiol 15:65. https://doi.org/10.1186/s12866-015-0398-4
Zhang Z, Tariq A, Zeng F, Graciano C, Sun F, Chau X, Ahmed Z (2021) Nitrogen and water addition regulate fungal community and microbial co-occurrence network complexity in the rhizosphere of Alhagi sparsifolia seedlings. Appl Soil Ecol 164:103940
Acknowledgements
We thank all the members of the Laboratoire de Génomique et Biodiversité Microbienne des Biofilms (LGBMB) of the Institute for Integrative Biology of the Cell (I2BC) and the Editor and three anonymous Reviewers for their interesting discussions, comments and suggestions. JO was a doctoral scholarship recipient from CONICYT Becas Chile (2012-2016). GR was a postdoctoral scholarship recipient from Brazil. This work was supported by the Centre National de la Recherche Scientifique (CNRS), France.
Author information
Authors and Affiliations
Contributions
Osman JR and DuBow MS designed the experiments, review and editing; Kamilova E collected the sand samples; Fernandes GR and Osman JR conducted the experiments; Fernandes GR analyzed the data; Fernandes GR, Osman JR and Dubow MS wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Conflict of interest
None.
Additional information
# In memory of Dr. Michael S. DuBow. On behalf of the authors, we thank him for his support as a tutor and scientific collaborator.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Osman, J.R., Fernandes, G.R., Kamilova, E. et al. Genomic microbiome analyses of surface sand samples from the Kyzyl-Kum Desert (Uzbekistan): characterization and comparative study. Arch Microbiol 205, 90 (2023). https://doi.org/10.1007/s00203-023-03432-z
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00203-023-03432-z