Abstract
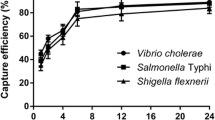
Yersinia enterocolitica is an important zoonotic pathogen, which seriously endangers food-safety risk. In this study, the recombinant outer membrane protein OmpF and its antibody were prepared and coupled with immunomagnetic beads (IMBs) to capture Y. enterocolitica in food samples, combining the quantitative PCR detection with primers of virulence factor gene foxA for Yersinia enterocolitica contamination. The results showed that the capture efficiency of approximately 80% using anti-OmpF antibody-immunomagnetic beads and linearly dependent capture under 101–105 CFU/mL Y. enterocolitica compared with less than 10% capture of other bacteria. The detection limit of 64 CFU/mL was obtained based on foxA gene PCR detection combined with capture of the anti-OmpF antibody-immunomagnetic beads to detect Yersinia enterocolitica in artificially contaminated milk and pork samples. Compared to the culture method, the developed IMBs–qPCR method has higher consistency, was less time consuming, which taken together provides an effective alternative method for rapid detection of Y. enterocolitica in food.






Similar content being viewed by others
References
Agrimonti C, Bottari B, Sardaro MLS, Marmiroli N (2019) Application of real-time PCR (qPCR) for characterization of microbial populations and type of milk in dairy food products. Crit Rev Food Sci Nutr 59:423–442. https://doi.org/10.1080/10408398.2017.1375893
Bliska JB, Falkow S (1992) Bacterial resistance to complement killing mediated by the Ail protein of Yersinia enterocolitica. Proc Natl Acad Sci U S A 89:3561–3565. https://doi.org/10.1073/pnas.89.8.3561
Chen J, Park B (2018) Effect of immunomagnetic bead size on recovery of foodborne pathogenic bacteria. Int J Food Microbiol 267:1–8. https://doi.org/10.1016/j.ijfoodmicro.2017.11.022
Chen Q, Li Y, Tao T, Bie X, Lu F, Lu Z (2017) Development and application of a sensitive, rapid, and reliable immunomagnetic separation-PCR detection method for Cronobacter spp. J Dairy Sci 100:961–969. https://doi.org/10.3168/jds.2016-11087
Delcour AH (2009) Outer membrane permeability and antibiotic resistance. Biochim Biophys Acta 1794:808–816. https://doi.org/10.1016/j.bbapap.2008.11.005
Fabrega A, Vila J (2012) Yersinia enterocolitica: pathogenesis, virulence and antimicrobial resistance. Enferm Infecc Microbiol Clin 30:24–32. https://doi.org/10.1016/j.eimc.2011.07.017
Fairman JW, Noinaj N, Buchanan SK (2011) The structural biology of beta-barrel membrane proteins: a summary of recent reports. Curr Opin Struct Biol 21:523–531. https://doi.org/10.1016/j.sbi.2011.05.005
Fredriksson-Ahomaa M, Stolle A, Korkeala H (2006) Molecular epidemiology of Yersinia enterocolitica infections. FEMS Immunol Med Microbiol 47:315–329. https://doi.org/10.1111/j.1574-695X.2006.00095.x
Galluzzi L, Ceccarelli M, Diotallevi A, Menotta M, Magnani M (2018) Real-time PCR applications for diagnosis of leishmaniasis. Parasit Vectors 11:273. https://doi.org/10.1186/s13071-018-2859-8
Garrido-Maestu A, Azinheiro S, Carvalho J, Prado M (2018) Rapid and sensitive detection of viable Listeria monocytogenes in food products by a filtration-based protocol and qPCR. Food Microbiol 73:254–263. https://doi.org/10.1016/j.fm.2018.02.004
Huang Y et al (2010) Possible use of ail and foxA polymorphisms for detecting pathogenic Yersinia enterocolitica. BMC Microbiol 10:211. https://doi.org/10.1186/1471-2180-10-211
Huang W et al (2016) Immunization with a 22-kDa outer membrane protein elicits protective immunity to multidrug-resistant Acinetobacter baumannii. Sci Rep 6:20724. https://doi.org/10.1038/srep20724
Kang HZDH, Shen ML et al (2015) Superantigenicity analysis of staphylococcal enterotoxins SElK and SElQ in a mouse model. RSC Adv 5:29684–29692
Kasturi KN, Drgon T (2017) Real-time PCR method for detection of salmonella spp in environmental samples. Appl Environ Microbiol. https://doi.org/10.1128/aem.00644-17
Koebnik R, Locher KP, Van Gelder P (2000) Structure and function of bacterial outer membrane proteins: barrels in a nutshell. Mol Microbiol 37:239–253. https://doi.org/10.1046/j.1365-2958.2000.01983.x
Kornreich-Leshem H et al (2005) Ferrioxamine B analogues: targeting the FoxA uptake system in the pathogenic Yersinia enterocolitica. J Am Chem Soc 127:1137–1145. https://doi.org/10.1021/ja035182m
Lim MC et al (2016) Biological preparation of highly effective immunomagnetic beads for the separation, concentration, and detection of pathogenic bacteria in milk. Colloids Surf B Biointerfaces 145:854–861. https://doi.org/10.1016/j.colsurfb.2016.05.077
Mikula KM, Kolodziejczyk R, Goldman A (2012) Yersinia infection tools-characterization of structure and function of adhesins. Front Cell Infect Microbiol 2:169. https://doi.org/10.3389/fcimb.2012.00169
Morse DL, Shayegani M, Gallo RJ (1984) Epidemiologic investigation of a Yersinia camp outbreak linked to a food handler. Am J Public Health 74:589–592. https://doi.org/10.2105/ajph.74.6.589
Perry RD, Brubaker RR (1979) Accumulation of iron by yersiniae. J Bacteriol 137:1290–1298
Pierson DE, Falkow S (1993) The ail gene of Yersinia enterocolitica has a role in the ability of the organism to survive serum killing. Infect Immun 61:1846–1852
Rosner BM, Stark K, Werber D (2010) Epidemiology of reported Yersinia enterocolitica infections in Germany, 2001–2008. BMC Public Health 10:337. https://doi.org/10.1186/1471-2458-10-337
Rusak LA et al (2018) Rapid detection of Yersinia enterocolitica serotype O:3 using a duplex PCR assay. J Microbiol Methods 154:107–111. https://doi.org/10.1016/j.mimet.2018.10.014
Saraka D et al (2017) Yersinia enterocolitica, a neglected cause of human enteric infections in Cote d’Ivoire. PLoS Negl Trop Dis 11:e0005216. https://doi.org/10.1371/journal.pntd.0005216
Schrammel B, Petzold M, Cervero-Arago S, Sommer R, Luck C, Kirschner A (2018) Persistent presence of outer membrane epitopes during short- and long-term starvation of five Legionella pneumophila strains. BMC Microbiol 18:75. https://doi.org/10.1186/s12866-018-1220-x
Shaban H, Na I, Kislichkina AA, Dentovskaya SV, Anisimov AP, Uversky VN (2017) Effect of natural polymorphism on structure and function of the Yersinia pestis outer membrane porin F (OmpF protein): a computational study. J Biomol Struct Dyn 35:2588–2603. https://doi.org/10.1080/07391102.2016.1224734
Shayegani M, Morse D, DeForge I, Root T, Parsons LM, Maupin PS (1983) Microbiology of a major foodborne outbreak of gastroenteritis caused by Yersinia enterocolitica serogroup O:8. J Clin Microbiol 17:35–40
Song X, Shukla S, Kim M (2018) Detection of Cronobacter species in powdered infant formula using immunoliposome-based immunomagnetic concentration and separation assay. Food Microbiol 72:23–30. https://doi.org/10.1016/j.fm.2017.11.002
Srisa-Art M, Boehle KE, Geiss BJ, Henry CS (2018) Highly sensitive detection of salmonella typhimurium using a colorimetric paper-based analytical device coupled with Immunomagnetic Separation. Anal Chem 90:1035–1043. https://doi.org/10.1021/acs.analchem.7b04628
Stenkova AM, Isaeva MP, Shubin FN, Rasskazov VA, Rakin AV (2011) Trends of the major porin gene (ompF) evolution: insight from the genus Yersinia. PLoS ONE 6:e20546. https://doi.org/10.1371/journal.pone.0020546
Thoerner P et al (2003) PCR detection of virulence genes in Yersinia enterocolitica and Yersinia pseudotuberculosis and investigation of virulence gene distribution. Appl Environ Microbiol 69:1810–1816. https://doi.org/10.1128/aem.69.3.1810-1816.2003
Tsang TM, Wiese JS, Felek S, Kronshage M, Krukonis ES (2013) Ail proteins of Yersinia pestis and Y. pseudotuberculosis have different cell binding and invasion activities. PLoS ONE 8:e83621. https://doi.org/10.1371/journal.pone.0083621
Vital PG, Van Ha NT, Tuyet LT, Widmer KW (2017) Application of quantitative real-time PCR compared to filtration methods for the enumeration of Escherichia coli in surface waters within Vietnam. J Water Health 15:155–162. https://doi.org/10.2166/wh.2016.173
Wang JZ et al (2014) Real-time TaqMan PCR for Yersinia enterocolitica detection based on the ail and foxA genes. J Clin Microbiol 52:4443–4444. https://doi.org/10.1128/jcm.02528-14
Wang E et al (2018a) Molecular characterization, phylogenetic, expression, and protective immunity analysis of OmpF, a promising candidate immunogen against yersinia ruckeri infection in channel catfish. Front Immunol 9:2003. https://doi.org/10.3389/fimmu.2018.02003
Wang J et al (2018b) Rapid detection of food-borne Salmonella contamination using IMBs–qPCR method based on pagC gene. Braz J Microbiol 49:320–328. https://doi.org/10.1016/j.bjm.2017.09.001
Wen Z et al (2016) Recombinant expression of Chlamydia trachomatis major outer membrane protein in E. Coli outer membrane as a substrate for vaccine research. BMC Microbiol 16:165. https://doi.org/10.1186/s12866-016-0787-3
Xiong Q, Cui X (2014) Development of an immunomagnetic separation method for efficient enrichment of Escherichia coli O157:H7. Food Control 37:41–45
Zhang X et al (2018) Immunization with pseudomonas aeruginosa outer membrane vesicles stimulates protective immunity in mice. Vaccine 36:1047–1054. https://doi.org/10.1016/j.vaccine.2018.01.034
Zhu P et al (2011) Detection of E. coli O157:H7 by immunomagnetic separation coupled with fluorescence immunoassay. Biosens Bioelectron 30:337–341. https://doi.org/10.1016/j.bios.2011.09.029
Acknowledgements
This project was supported by National Key Research and Development Program of China (2018YFD0500501), and Key Projects of Science and Technology Support Grant of Tianjin in China (20YFZCSN00340).
Author information
Authors and Affiliations
Contributions
Conceived and designed the experiments: JHH. Performed the experiments: JXS, HC, APC, YNS, MZ, LLZ and FZX. Analyzed the data: APC and JHH. Contributed reagents/materials /analysis tools: JHH. Wrote the paper: JXS, HC, and JHH.
Corresponding authors
Ethics declarations
Conflict of interest
We declare that we have no competing interests.
Additional information
Communicated by Erko Stackebrandt.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Shi, J., Chi, H., Cao, A. et al. Development of IMBs–qPCR detection method for Yersinia enterocolitica based on the foxA gene. Arch Microbiol 203, 4653–4662 (2021). https://doi.org/10.1007/s00203-021-02459-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00203-021-02459-4