Abstract
Key message
Using a fixed RIL population derived from a widely used foxtail millet backbone breeding line and an elite cultivar, we constructed a high-density bin map and identified six novel multi-environment effect QTLs and seven candidate genes for dwarf phenotype.
Abstract
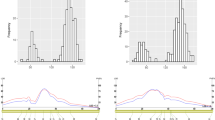
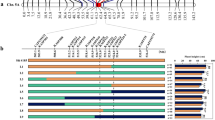
Plant height is an important trait that determines tradeoffs between competition and resource allocation, which is crucial for yield potential. To improve the C4 model plant foxtail millet (Setaria italica) productivity, it is necessary to isolate plant height-related genes that contribute to ideal plant architecture in breeding. In the present study, we generated a foxtail millet population of 333 recombinant inbred lines (RILs) derived from a cross between a backbone line Ai 88 and an elite cultivar Liaogu 1. We evaluated plant height in 13 environmental conditions across 4 years, the mean plant height of the RIL population ranged from 89.5 to 149.9 cm. Using deep re-sequencing data, we constructed a high-density bin map with 3744 marker bins. Quantitative trait locus (QTL) mapping identified 26 QTLs significantly associated with plant height. Of these, 13 QTLs were repeatedly detected under multiple environments, including six novel QTLs that have not been reported before. Seita.1G242300, a gene encodes gibberellin 2-oxidase-8, which was detected in nine environments in a 1.54-Mb interval of qPH1.3, was considered as an important candidate gene. Moreover, other six genes involved in GA biosynthesis or signaling pathways, and fifteen genes encode F-box domain proteins which might function as E3 ligases, were also considered as candidate genes in different QTLs. These QTLs and candidate genes identified in this study will help to elucidate the genetic basis of foxtail millet plant height, and the linked markers will be useful for marker-assistant selection of varieties with ideal plant architecture and high yield potential.


Similar content being viewed by others
References.
Bennetzen JL, Schmutz J, Wang H, Percifield R, Hawkins J, Pontaroli AC, Estep M, Feng L, Vaughn JN, Grimwood J (2012) Reference genome sequence of the model plant Setaria. Nat Biotechnol 30:555–561
Bettinger RL, Barton L, Morgan C (2010) The origins of food production in north china: a different kind of agricultural revolution. Evol Anthropol 19:9–21
Cassani E, Bertolini E, Badone CF, Landoni M, Gavina D, Sirizzotti A, Pilu R (2009) Characterization of the first dominant dwarf maize mutant carrying a single amino acid insertion in the VHYNP domain of the dwarf8 gene. Mol Breed 24:375–385
Cheng R, Liu Z, Shi Z, Xia X, Yang W (2006) Breeding of foxtail millet cultivar Jigu19. J Hebei Agron Sci 10:80–81 (In Chinese)
Diao X (2016) Production and genetic improvement of minor cereals in China. Crop J 5:103–114
Diao X, Cheng R (2017) Current breeding situation of foxtail millet and common millet in china as revealed by exploitation of 15 years regional adaptation test data. Sci Agric Sin 50(23):4469–4474
Diao X, Jia G (2017) Foxtail millet breeding in China. In: Diao X (ed) Doust A. Genetics and genomics of setaria. plant genetics and genomics, Crops and Models, Springer, Cham, pp 93–113
Diao X, Schnable J, Bennetzen JL, Jiayang L (2014) Initiation of Setaria as a model plant. Front Agr Sci Eng 1:16–20
Dill A, Thomas SG, Hu J, Steber CM, Sun T-P (2004) The Arabidopsis F-box protein SLEEPY1 targets gibberellin signaling repressors for gibberellin-induced degradation. Plant Cell 16:1392–1405
Doust A, Kellogg E, Devos K, Bennetzen J (2009) Foxtail millet: a sequence-driven grass model system. Plant Physiol 149:137–141
Evans LT (1998) Feeding the ten billion plant and population growth. Cambridge University Press, Cambridge
Evenson Gollin RD (2003) Assessing the impact of the green revolution, 1960 to 2000. Science 300(5620):758–762
Fan X, Tang S, Zhi H, He M, Ma W, Jia Y, Zhao B, Jia G, Diao X (2017) Identification and fine mapping of SiDWARF (D3), a pleiotropic locus controlling environment-independent dwarfism in foxtail millet. Crop Sci 57:2431–2442
Fang X, Dong K, Wang X, Liu T, He J, Ren R, Zhang L, Liu R, Liu X, Li M, Huang M, Zhang Z, Yang TJBG (2016) A high density genetic map and QTL for agronomic and yield traits in Foxtail millet Setaria italica L P. Beauv. BMC Genom 17:336. https://doi.org/10.1186/s12864-016-2628-z
Feldman MJ, Paul RE, Banan D, Barrett JF, Genetics IBJP (2016) Time dependent genetic analysis links field and controlled environment phenotypes in the model C4 grass Setaria. PLoS Genet 13:e1006841
Fujioka S, Yamane H, Spray CR, Katsumi M, Phinney BO, Gaskin P, MacMillan J, Takahashi N (1988) The dominant non-gibberellin-responding dwarf mutant (D8) of maize accumulates native gibberellins. Proc Natl Acad Sci USA 85:9031–9035
Hedden P (2003) The genes of the green revolution. Trends Genet 19:5–9
Hong Z, Ueguchi-Tanaka M, Fujioka S, Takatsuto S, Yoshida S, Hasegawa Y, Ashikari M, Kitano H, Matsuoka M (2005) The Rice brassinosteroid-deficient dwarf2 mutant, defective in the rice homolog of Arabidopsis DIMINUTO/DWARF1, is rescued by the endogenously accumulated alternative bioactive brassinosteroid, dolichosterone. Plant cell 17:2243–2254
Hong Z, Ueguchi-Tanaka M, Umemura K, Uozu S, Fujioka S, Takatsuto S, Yoshida S, Ashikari M, Kitano H, Matsuoka M (2003) A rice brassinosteroid-deficient mutant, ebisu dwarf (d2), is caused by a loss of function of a new member of cytochrome P450. Plant Cell 15:2900–2910
Huang X, Feng Q, Qian Q, Zhao Q, Wang L, Wang A, Guan J, Fan D, Weng Q, Huang T, Dong G, Sang T, Han B (2009) High-throughput genotyping by whole-genome resequencing. Genome Res 19:1068–1076
Jia G, Huang X, Zhi H, Zhao Y, Zhao Q, Li W, Chai Y, Yang L, Liu K, Lu H (2013) A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nat Genet 45:957–961
Jia X, Zhang Z, Liu Y, Zhang C, Shi Y, Song Y, Wang T, Li Y (2009) Development and genetic mapping of SSR markers in foxtail millet [Setaria italica (L.) P. Beauv.]. Theor Appl Genet 118:821–829
Kir G, Ye H, Nelissen H, Neelakandan AK, Kusnandar AS, Luo A, Inzé D, Sylvester AW, Yin Y, Becraft PW (2015) RNA interference knockdown of BRASSINOSTEROID INSENSITIVE1 in maize reveals novel functions for brassinosteroid signaling in controlling plant architecture. Plant Physiol 169:826–839
Lata C, Gupta S, Prasad M (2013) Foxtail millet: a model crop for genetic and genomic studies in bioenergy grasses. Crit Rev Biotechnol 33:328–343
Li H, Durbin R (2009) Fast and accurate short read alignment with burrows-wheeler transform. Bioinformatics 25:1754–1760
Li H, Wang L, Liu M, Dong Z, Li Q, Fei S, Xiang H, Liu B, Jin W (2020) Maize plant architecture is regulated by the ethylene biosynthetic gene ZmACS7. Plant Physiol 183:1184–1199
Li L, Sun W, Cheng Y (2005) Breeding report of a new high-yield and quality millet variety Liaogu 1. Rain fed crops 25:307–307 (In Chinese)
Li P, Brutnell TP (2011) Setaria viridis and Setaria italica, model genetic systems for the Panicoid grasses. J Exp Bot 62:3031–3037
Li S, Hu X, Li J, Wang Y (2010) Breeding and cultivation techniques of a new high-quality, dwarf foxtail millet variety Gongai 6. China Sci Technol Adv 11:11–11 (In Chinese)
Liu X, Gao S, Yang M, Li J (2005) Selection of a good quality high yield and new type foxtail millet variety Gongai 2. Hortic Seed 25:305–306 (In Chinese)
Liu XT, Tang S, Jia GQ, Schnable JC, Su HX, Tang CJ, Zhi H, Diao XM (2016) The C-terminal motif of SiAGO1b is required for the regulation of growth, development and stress responses in foxtail millet (Setaria italica (L.) P. Beauv). J Exp Bot 67:3237–3249
Lu H, Zhang J, Liu K, Wu N, Li Y, Zhou K, Ye M, Zhang T, Zhang H, Yang X, Shen L, Xu D, Li Q (2009) Earliest domestication of common millet (Panicum miliaceum) in East Asia extended to 10,000 years ago. Proc Natl Acad Sci USA 106:7367–7372
Lu P (2006) Criteria for descriptions of morphological characteristics of foxtail millet germplasm. China Agricultural Press, Beijing (In Chinese)
Mace E, Innes D, Hunt C, Wang X, Tao Y, Baxter J, Hassall M, Hathorn A, Jordan D (2019) The Sorghum QTL Atlas: a powerful tool for trait dissection, comparative genomics and crop improvement. Theor Appl Genet 132:751–766
Multani DS, Briggs SP, Chamberlin MA, Blakeslee JJ, Murphy AS, Johal GS (2003) Loss of an MDR transporter in compact stalks of maize br2 and sorghum dw3 mutants. Science 302:81–84
Ni X, Xia Q, Zhang H, Cheng S, Li H, Fan G, Guo T, Huang P, Xiang H, Chen Q, Li N, Zou H, Cai X, Lei X, Wang X, Zhou C, Zhao Z, Zhang G, Du G, Cai W, Quan Z (2017) Updated foxtail millet genome assembly and gene mapping of nine key agronomic traits by resequencing a RIL population. Giga Sci 6:1–8
Peng J, Carol P, Richards DE, King KE, Cowling RJ, Murphy GP, Harberd NP (1997) The Arabidopsis GAI gene defines a signaling pathway that negatively regulates gibberellin responses. Genes Dev 11:3194–3205
Peng J, Richards DE, Hartley NM, Murphy GP, Devos KM, Flintham JE, Beales J, Fish LJ, Worland AJ, Pelica F, Sudhakar D, Christou P, Snape JW, Gale MD, Harberd NP (1999) “Green revolution” genes encode mutant gibberellin response modulators. Nature 400:256–261
Qian J, Jia G, Hui Z, Wei L, Wang Y, Li H, Shang Z, Doust AN, Diao X (2012) Sensitivity to gibberellin of dwarf foxtail millet varieties. Crop Sci 52:1068–1075
Quinby JR, Karper PE (1954) Inheritance of height in sorghum. In abstracts of the annual meetings of the American society of agronomy, Dallas, Texas 98–99
Sasaki A, Ashikari M, Ueguchi-Tanaka M, Itoh H, Nishimura A, Datta S, Ishiyama K, Saito T, Kobayashi M, Khush S, G, Kitano H, Matsuoka M, (2002) A mutant gibberellin–synthesis gene in rice. Nature 416:701–702
Sasaki A, Itoh H, Gomi K, Ueguchi-Tanaka M, Ishiyama K, Kobayashi M, Jeong DH, An G, Kitano H, Ashikari Matsuoka M (2003) Accumulation of phosphorylated repressor for gibberellin signaling in an F-box mutant. Science 299:1896–1898
Schomburg F, Bizzell M, C, Lee DJ, A D Zeevaart J, M Amasino R, (2003) Overexpression of a novel class of gibberellin 2-oxidases decreases gibberellin levels and creates dwarf plants. Plant Cell 15:151–163
Schnable J, Zang Y, Ngu DWC (2016) Pan-grass syntenic gene set sorghum reference. figshare https://doi.org/10.6084/m9.figshare.3113488.v1
Spielmeyer W, Ellis MH, Chandler PM (2002) Semidwarf (sd-1), “green revolution” rice, contains a defective gibberellin 20-oxidase gene. Proc Natl Acad Sci USA 99:9043–9048
Sun TP (2011) The molecular mechanism and evolution of the GA-GID1-DELLA signaling module in plants. Curr Biol 21:R338–R345
Tanabe S, Ashikari M, Fujioka S, Takatsuto S, Yoshida S, Yano M, Yoshimura A, Kitano H, Matsuoka M, Fujisawa Y, Kato H, Iwasaki Y (2005) A novel cytochrome P450 is implicated in brassinosteroid biosynthesis via the characterization of a rice dwarf mutant, dwarf11 with reduced seed length. Plant Cell 17(3):776–790
Teng F, Zhai L, Liu R, Bai W, Wang L, Huo D, Tao Y, Zheng Y, Zhang Z (2013) ZmGA3ox2, a candidate gene for a major QTL, qPH3.1, for plant height in maize. Plant J 73:405–416
Ueguchi-Tanaka M, Ashikari M, Nakajima M, Itoh H, Katoh E, Kobayashi M, Chow T, Hsing Y, Kitano H, Yamaguchi I, Matsuoka M (2005) GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 437:693–698
Wang J, Wang Z, Du X, Yang H, Han F, Han Y, Yuan F, Zhang L, Peng S, Guo E (2017) A high-density genetic map and QTL analysis of agronomic traits in foxtail millet Setaria italica L P. Beauv. using RAD-seq. PLoS ONE. https://doi.org/10.1371/journal.pone.0179717
Wang S, Xie H, Xing L, Wei M, Wang S, Liu H (2016) Breeding of a high quality and yield variety yugu25 derived from xiagu. China Seed Ind 11:58–59 (In Chinese)
Wang Z, Wang J, Peng J, Du X, Jiang M, Li Y, Han F, Du G, Yang H, Lian S, Yong J, Cai W, Cui J, Han K, Yuan F, Chang F, Yuan G, Zhang W, Zhang L, Peng S, Zou H, Guo E (2019) QTL mapping for 11 agronomic traits based on a genome-wide Bin-map in a large F2 population of foxtail millet Setaria italica L. P Beauv Mol Breed. https://doi.org/10.1007/s11032-019-0930-6
Winkler RG, Helentjaris T (1995) The maize Dwarf3 gene encodes a cytochrome P450-mediated early step in gibberellin biosynthesis. Plant Cell 7:1307–1317
Wu J, Jiang H (1994) The first upright leaf foxtail millet cultivar-Ai88. Beijing Agric 026:18–19 (In Chinese)
Xing A, Gao Y, Ye L, Zhang W, Cai L, Ching A, Llaca V, Johnson B, Liu L, Yang X, Kang D, Yan J, Li J (2015) A rare SNP mutation in Brachytic2 moderately reduces plant height and increases yield potential in maize. J Exp Bot 66:3791–3802
Xue C, Zhi H, Fang X, Liu X, Tang S, Chai Y, Zhao B, Jia G, Diao X (2016) Characterization and fine mapping of SiDWARF2 (D2) in foxtail millet. Crop Sci 56:95–103
Zhang K, Fan G, Zhang X, Zhao F, Wei W, Du G, Feng X, Wang X, Wang F, Song G, Zou H, Zhang X, Li S, Ni X, Zhang G, Zhao Z (2017) Identification of QTLs for agronomically important traits in Setaria italica based on SNPs generated from high-throughput sequencing. G3-Gene Gen Genet 7(5):1587–1594
Zhao M, Zhi H, Zhang X, Jia G, Diao X (2019) Retrotransposon-mediated DELLA transcriptional reprograming underlies semi-dominant dwarfism in foxtail millet. Crop J 7:458–468
Acknowledgement
We thank Dr. Kun Xie helped in data analysis. This research was supported by the National Key R&D Program of China (2018YFD1000706, 2018YFD1000700), the National Natural Science Foundation of China (31871630, 31771807), China agricultural research system (CARS-06-13.5) and the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences.
Funding
This research was supported by the National Key R&D Program of China (2018YFD1000706, 2018YFD1000700), the National Natural Science Foundation of China (31871630, 31771807), China Agricultural Research System (CARS-06–13.5) and the Agricultural Science and Technology Innovation Program of the Chinese Academy of Agricultural Sciences.
Author information
Authors and Affiliations
Contributions
QH and HZ did data analysis and drafted the manuscript. Xd and HZ designed the experiment, developed the RIL population, and revised the manuscript. Jun Liu helped in the data analysis and discussion. ST, LX, SW, HW, AZ, YL, MG, HZ, GC, SD, JL, JY, HL, WZ, YJ, SL, JL, ZQ, EG, and GJ collected the phenotype. All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest
Additional information
Communicated by Ian D Godwin.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
He, Q., Zhi, H., Tang, S. et al. QTL mapping for foxtail millet plant height in multi-environment using an ultra-high density bin map. Theor Appl Genet 134, 557–572 (2021). https://doi.org/10.1007/s00122-020-03714-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03714-w