Abstract
Key message
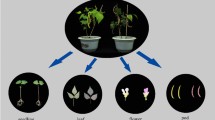
The bHLH transcription factor, PPLS1, interacts with SiMYB85 to control the color of pulvinus and leaf sheath by regulating anthocyanin biosynthesis in foxtail millet (Setaria italica).
Abstract
Foxtail millet (Setaria italica), a self-pollinated crop with numerous small florets, is difficult for cross-pollination. The color of pulvinus and leaf sheath with purple being dominant to green is an indicative character and often used for screening authentic hybrids in foxtail millet crossing. Deciphering molecular mechanism controlling this trait would greatly facilitate genetic improvement of cultivars in foxtail millet. Here, using the F2 bulk specific-locus amplified fragment sequencing approach, we mapped the putative causal gene for the purple color of pulvinus and leaf sheath (PPLS) trait to a 100 Kb region on chromosome 7. Expression analyses of the 15 genes in this region revealed that Seita.7G195400 (renamed here as PPLS1) was differentially expressed between purple and green cultivars. PPLS1 encodes a bHLH transcription factor and is localized in the nucleus with a transactivation activity. Furthermore, we observed that expression of a MYB transcription factor gene, SiMYB85 (Seita.4G086300) involved in anthocyanin biosynthesis, shows a totally positive association with that of PPLS1. Heterologous co-expression of both PPLS1 and SiMYB85 in tobacco leaves led to elevated anthocyanin accumulation and expression of some anthocyanin-related genes. Furthermore, PPLS1 physically interacts with SiMYB85. Taken together, our results suggest that PPLS1 interacts with SiMYB85 to control the color of pulvinus and leaf sheath by regulating anthocyanin biosynthesis in foxtail millet.








Similar content being viewed by others
References
Abe A, Kosugi S, Yoshida K, Natsume S, Takagi H, Kanzaki H, Matsumura H, Yoshida K, Mitsuoka C, Tamiru M, Innan H, Cano L, Kamoun S, Terauchi R (2012) Genome sequencing reveals agronomically important loci in rice using MutMap. Nat Biotechnol 30:174–178
Bai H, Cao YH, Quan JZ, Dong L, Li ZY, Zhu YB, Zhu LH, Dong ZP, Li DY (2013) Identifying the genome-wide sequence variations and developing new molecular markers for genetics research by re-sequencing a landrace cultivar of foxtail millet. PLoS ONE 8:e73514
Bart R, Chern M, Park CJ, Bartley L, Ronald PC (2006) A novel system for gene silencing using siRNAs in rice leaf and stem-derived protoplasts. Plant Methods 2:13
Bennetzen JL, Schmutz J, Wang H, Percifield R, Hawkins J, Pontaroli AC, Estep M, Feng L, Vaughn JN, Grimwood J, Jenkins J, Barry K, Lindquist E, Hellsten U, Deshpande S, Wang X, Wu X, Mitros T, Triplett J, Yang X, Ye CY, Mauro-Herrera M, Wang L, Li P, Sharma M, Sharma R, Ronald PC, Panaud O, Kellogg EA, Brutnell TP, Doust AN, Tuskan GA, Rokhsar D, Devos KM (2012) Reference genome sequence of the model plant Setaria. Nat Biotechnol 30:555–561
Broun P (2005) Transcriptional control of flavonoid biosynthesis: a complex network of conserved regulators involved in multiple aspects of differentiation in Arabidopsis. Curr Opin Plant Biol 8:272–279
Chandler VL, Radicella JP, Robbins TP, Chen J, Turks D (1989) Two regulatory genes of the maize anthocyanin pathway are homologous: isolation of B utilizing R genomic sequences. Plant Cell 1:1175–1183
Chen H, Zou Y, Shang Y, Lin H, Wang Y, Cai R, Tang X, Zhou JM (2008) Firefly luciferase complementation imaging assay for protein-protein interactions in plants. Plant Physiol 146:368–376
D'Amelia V, Aversano R, Batelli G, Caruso I, Castellano Moreno M, Castro-Sanz AB, Chiaiese P, Fasano C, Palomba F, Carputo D (2014) High AN1 variability and interaction with basic helix-loop-helix co-factors related to anthocyanin biosynthesis in potato leaves. Plant J 80:527–540
Dixon RA, Liu CG, Jun JH (2013) Metabolic engineering of anthocyanins and condensed tannins in plants. Curr Opin Biotechnol 24:329–335
Dooner HK, Robbins TP, Jorgensen RA (1991) Genetic and developmental control of anthocyanin biosynthesis. Annu Rev Genet 25:173–219
Doust AN, Kellogg EA, Devos KM, Bennetzen JL (2009) Foxtail millet: a sequence-driven grass model system. Plant Physiol 149:137–141
Feller A, Hernandez JM, Grotewold E (2006) An ACT-like domain participates in the dimerization of several plant basic-helix-loop-helix transcription factors. J Biol Chem 281:28964–28974
Feng K, Xu ZS, Que F, Liu JX, Wang F, Xiong AS (2018) An R2R3-MYB transcription factor, OjMYB1, functions in anthocyanin biosynthesis in Oenanthe javanica. Planta 247:301–315
Gao DY, He B, Zhou YH, Sun LH (2011) Genetic and molecular analysis of a purple sheath somaclonal mutant in japonica rice. Plant Cell Rep 30:901–911
Goff SA, Cone KC, Chandler VL (1992) Functional analysis of the transcriptional activator encoded by the maize B gene: evidence for a direct functional interaction between two classes of regulatory proteins. Genes Dev 6:864–875
Grotewold E, Sainz MB, Tagliani L, Hernandez JM, Bowen B, Chandler VL (2000) Identification of the residues in the Myb domain of maize C1 that specify the interaction with the bHLH cofactor R. Proc Natl Acad Sci USA 97:13579–13584
Heim MA, Jakoby M, Werber M, Martin C, Weisshaar B, Bailey PC (2003) The basic helix–loop–helix transcription factor family in plants: a genome-wide study of protein structure and functional diversity. Mol Biol Evol 20:735–747
Hichri I, Barrieu F, Bogs J, Kappel C, Delrot S, Lauvergeat V (2011) Recent advances in the transcriptional regulation of the flavonoid biosynthetic pathway. J Exp Bot 62:2465–2483
Jia GQ, Huang XH, Zhi H, Zhao Y, Zhao Q, Li WJ, Chai Y, Yang LF, Liu KY, Lu HY, Zhu CR, Lu YQ, Zhou CC, Fan DL, Weng QJ, Guo YL, Huang T, Zhang L, Lu TT, Feng Q, Hao HF, Liu HK, Lu P, Zhang N, Li YH, Guo EH, Wang SJ, Wang SY, Liu JR, Zhang WF, Chen GQ, Zhang BJ, Li W, Wang YF, Li HQ, Zhao BH, Li JY, Diao XM, Han B (2013) A haplotype map of genomic variations and genome-wide association studies of agronomic traits in foxtail millet (Setaria italica). Nat Genet 45:957–961
Jiang WH, Liu TX, Nan WZ, Chamila Jeewani D, Niu YL, Li CL, Wang Y, Shi X, Wang C, Wang JH, Li Y, Gao X, Wang ZH (2018) Two transcription factors TaPpm1 and TaPpb1 co-regulate the anthocyanin biosynthesis in purple pericarp of wheat. J Exp Bot 69:2555–2567
Kang YH, Song SK, Schiefelbein J, Lee MM (2013) Nuclear trapping controls the position-dependent localization of CAPRICE in the root epidermis of Arabidopsis. Plant Physiol 163:193–204
Karki S, Rizal G, Quick WP (2013) Improvement of photosynthesis in rice (Oryza sativa L.) by inserting the C4 pathway. Rice (N Y) 6:28
Kent WJ (2002) BLAT—the BLAST-like alignment tool. Genome Res 12:656–664
Khlestkina EK (2013) Genes determining the coloration of different organs in wheat. Russ J Genet Appl Res 3:54–65
Koes R, Verweij W, Quattrocchio F (2005) Flavonoids: a colorful model for the regulation and evolution of biochemical pathways. Trends Plant Sci 10:236–242
Kumar S, Tamura K, Jakobsen IB, Nei M (2001) MEGA2: molecular evolutionary genetics analysis software. Bioinformatics 17:1244–1245
Lata C, Gupta S, Prasad M (2013) Foxtail millet: a model crop for genetic and genomic studies in bioenergy grasses. Crit Rev Biotechnol 33:328–343
Li YH (1979) Morphology and anatomy of gramineae crops. Shanghai Scientific and Technical Publishers, Shanghai, pp 379–393
Li ZY, Wang N, Dong L, Bai H, Quan JZ, Liu L, Dong ZP (2015) Differential gene expression in foxtail millet during incompatible interaction with Uromyces setariae-italicae. PLoS ONE 10:e0123825
Li YQ, Shan XT, Gao RF, Yang S, Wang SC, Gao X, Wang L (2016) Two IIIf clade-bHLHs from freesia hybrida play divergent roles in flavonoid biosynthesis and trichome formation when ectopically expressed in Arabidopsis. Sci Rep 6:30514
Lim SH, Kim DH, Kim JK, Lee JY, Ha SH (2017) A radish basic helix-loop-helix transcription factor, RsTT8 acts a positive regulator for anthocyanin biosynthesis. Front Plant Sci 8:1917
Ludwig SR, Habera LF, Dellaporta SL, Wessler SR (1989) Lc, a member of the maize R gene family responsible for tissue-specific anthocyanin production, encodes a protein similar to transcriptional activators and contains the myc-homology region. Proc Natl Acad Sci USA 86:7092–7096
Michelmore RW, Paran I, Kesseli RV (1991) Identification of markers linked to disease-resistance genes by bulked segregant analysis: a rapid method to detect markers in specific genomic regions by using segregating populations. Proc Natl Acad Sci USA 88:9828–9832
Muthamilarasan M, Prasad M (2015) Advances in Setaria genomics for genetic improvement of cereals and bioenergy grasses. Theor Appl Genet 128:1–14
Nesi N, Debeaujon I, Jond C, Pelletier G, Caboche M, Lepiniec L (2000) The TT8 gene encodes a basic helix-loop-helix domain protein required for expression of DFR and BAN genes in Arabidopsis siliques. Plant Cell 12:1863–1878
Pant SR, Irigoyen S, Doust AN, Scholthof KB, Mandadi KK (2016) Setaria: a food crop and translational research model for C4 grasses. Front Plant Sci 7:1885
Payyavula RS, Singh RK, Navarre DA (2013) Transcription factors, sucrose, and sucrose metabolic genes interact to regulate potato phenylpropanoid metabolism. J Exp Bot 64:5115–5131
Petroni K, Tonelli C (2011) Recent advances on the regulation of anthocyanin synthesis in reproductive organs. Plant Sci 181(3):219–229
Radicella JP, Turks D, Chandler VL (1991) Cloning and nucleotide sequence of a cDNA encoding B-Peru, a regulatory protein of the anthocyanin pathway in maize. Plant Mol Biol 17:127–130
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schwinn K, Venail J, Shang Y, Mackay S, Alm V, Butelli E, Oyama R, Bailey P, Davies K, Martin C (2006) A small family of MYB-regulatory genes controls floral pigmentation intensity and patterning in the genus Antirrhinum. Plant Cell 18:831–851
Spelt C, Quattrocchio F, Mol JN, Koes R (2000) anthocyanin1 of petunia encodes a basic helix-loop-helix protein that directly activates transcription of structural anthocyanin genes. Plant Cell 12:1619–1632
Sun XW, Liu DY, Zhang XF, Li WB, Liu H, Hong WG, Jiang CB, Guan N, Ma CX, Zeng HP, Xu CH, Song J, Huang L, Wang CM, Shi JJ, Wang R, Zheng XH, Lu CY, Wang XW, Zheng HK (2013) SLAF-seq: an efficient method of large-scale de novo SNP discovery and genotyping using high-throughput sequencing. PLoS ONE 8:e58700
Sun XM, Zhang ZY, Chen C, Wu W, Ren NN, Jiang CH, Yu JP, Zhao Y, Zheng XM, Yang QW, Zhang HL, Li JJ, Li ZC (2018) The C-S-A gene system regulates hull pigmentation and reveals evolution of anthocyanin biosynthesis pathway in rice. J Exp Bot 69:1485–1498
Winkel-Shirley B (2001) Flavonoid biosynthesis. A colorful model for genetics, biochemistry, cell biology, and biotechnology. Plant Physiol 126:485–493
Xiao H, Wang Y, Liu DF, Wang WM, Li XB, Zhao XF, Xu JC, Zhai WX, Zhu LH (2003) Functional analysis of the rice AP3 homologue OsMADS16 by RNA interference. Plant Mol Biol 52:957–966
Xu WJ, Dubos C, Lepiniec L (2015) Transcriptional control of flavonoid biosynthesis by MYB-bHLH-WDR complexes. Trends Plant Sci 20:176–185
Xu ZS, Yang QQ, Feng K, Xiong AS (2019) Changing carrot color: insertions in DcMYB7 alter the regulation of anthocyanin biosynthesis and modification. Plant Physiol 181:195–207
Xue W, Xing Y, Weng X, Zhao Y, Tang W, Wang L, Zhou H, Yu S, Xu C, Li X, Zhang Q (2008) Natural variation in Ghd7 is an important regulator of heading date and yield potential in rice. Nat Genet 40:761–767
Yan C, An G, Zhu T, Zhang W, Zhang L, Peng L, Chen J, Kuang H (2019) Independent activation of the BoMYB2 gene leading to purple traits in Brassica oleracea. Theor Appl Genet 132:895–906
Zhang HZ (1980) Study on the natural hybridization rate and the acquisition of sexual hybrids of millet. Shanxi Agric Sci 12–13
Zhang GY, Liu X, Quan ZW, Cheng SF, Xu X, Pan SK, Xie M, Zeng P, Yue Z, Wang WL, Tao Y, Bian C, Han CL, Xia QJ, Peng XH, Cao R, Yang XH, Zhan DL, Hu JC, Zhang YX, Li HN, Li H, Li N, Wang JY, Wang CC, Wang RY, Guo T, Cai YJ, Liu CZ, Xiang HT, Shi QX, Huang P, Chen QC, Li YR, Wang J, Zhao ZH, Wang J (2012) Genome sequence of foxtail millet (Setaria italica) provides insights into grass evolution and biofuel potential. Nat Biotechnol 30:549–554
Zhang Y, Butelli E, Martin C (2014) Engineering anthocyanin biosynthesis in plants. Curr Opin Plant Biol 19:81–90
Zhang P, Zhu YQ, Wang LL, Chen LP, Zhou SJ (2015) Mining candidate genes associated with powdery mildew resistance in cucumber via super-BSA by specific length amplified fragment (SLAF) sequencing. BMC Genomics 16:1058
Zhao TT, Jiang JB, Liu G, He SS, Zhang H, Chen XL, Li JF, Xu XY (2016) Mapping and candidate gene screening of tomato Cladosporium fulvum-resistant gene Cf-19, based on high-throughput sequencing technology. BMC Plant Biol 16:51
Zhao Y, Ma J, Li M, Deng L, Li G, Xia H, Zhao S, Hou L, Li P, Ma C, Yuan M, Ren L, Gu J, Guo B, Zhao C, Wang X (2019) Whole-genome resequencing-based QTL-seq identified AhTc1 gene encoding a R2R3-MYB transcription factor controlling peanut purple testa colour. Plant Biotechnol J 18:96–105
Zhu ZX, Lu YQ (2016) Plant color mutants and the anthocyanin pathway. Chin Bull Bot 51:107–119
Zhu ZX, Wang HL, Wang YT, Guan S, Wang F, Tang JY, Zhang RJ, Xie LL, Lu YQ (2015) Characterization of the cis elements in the proximal promoter regions of the anthocyanin pathway genes reveals a common regulatory logic that governs pathway regulation. J Exp Bot 66:3775–3789
Acknowledgements
We thank Dr. Yunyuan Xu (Institute of Botany, Chinese Academy of Sciences), Dr. Hong Luo (Clemson University), Dr. Maoyin Li (Donald Danforth Plant Science Center), Dr. Tianhua He (Curtin University), and Dr. Xu Li (North Carolina State University) for their critical reading and helpful suggestions for the manuscript. We also thank Drs Zhuangzhi Zhou, Zhao Liu, Weixia Lv and Yameng Peng for their technical assistance. This work was supported by the National Key R&D Program of China (2018YFD1000703 and 2018YFD1000700), the National Natural Science Foundation of China (31872880), HAAFS Agriculture Science and Technology Innovation Project (Grant number: 2019-4-2-3), and China Agriculture Research System (CARS-06-13.5-A25).
Author information
Authors and Affiliations
Contributions
HB, DL and ZD conceived and designed the experiments; HB, ZS, YZ, ZL, XL, JM and XW performed the experiments; YW contributed materials; JQ constructed the F2 populations; HB, ZS, DL, ML, JZ and ZD analyzed the data; HB, ZS and DL wrote the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Hai-Chun Jing.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Bai, H., Song, Z., Zhang, Y. et al. The bHLH transcription factor PPLS1 regulates the color of pulvinus and leaf sheath in foxtail millet (Setaria italica). Theor Appl Genet 133, 1911–1926 (2020). https://doi.org/10.1007/s00122-020-03566-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03566-4