Abstract
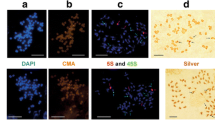
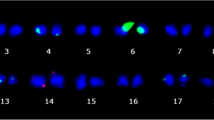
The extent of 5S and 45S ribosomal DNA (rDNA) variation was investigated in wild and domesticated common beans (Phaseolus vulgaris) chosen to represent the known genetic diversity of the species. 5S and 45S rDNA probes were localized on mitotic chromosomes of 37 accessions by fluorescent in situ hybridization (FISH). The two 5S rDNA loci were largely conserved within the species, whereas a high variation in the number of 45S rDNA loci and changes in position of loci and number of repeats per locus were observed. Domesticated accessions from the Mesoamerican gene pool frequently had three 45S rDNA loci per haploid genome, and rarely four. Domesticated accessions from Andean gene pool, particularly from the race Peru, showed six, seven, eight or nine loci, but seven loci were found in all three races of this gene pool. Between three and eight loci were observed in accessions resulting from crosses between Andean and Mesoamerican genotypes. The presence of two to eight 45S rDNA loci in wild common beans from different geographic locations indicates that the 45S rDNA amplification observed in the Andean lineage took place before domestication. Our data suggest that ectopic recombination between terminal chromosomal regions might be the mechanism responsible for this variation.



Similar content being viewed by others
References
Adams SP, Leitch IJ, Bennett MD, Chase MW, Leitch AR (2000) Ribosomal DNA evolution and phylogeny in Aloe (Asphodelaceae). Am J Bot 87:1578–1583
Amirul Islam FM, Beebe S, Munoz M, Tohme J, Redden RJ, Basford KE (2004) Using molecular markers to assess the effect of introgression on quantitative attributes of common bean in the Andean gene pool. Theor Appl Genet 108:243–252
Arnheim N, Krystal M, Schmickel R, Wilson G, Ryder O, Zimmer E (1980) Molecular evidence for genetic exchanges among ribosomal genes on non-homologous chromosomes in man and apes. Proc Natl Acad Sci USA 77:7323–7327
Becerra Velasquez VL, Gepts P (1994) RFLP diversity of common bean (Phaseolus vulgaris) in its centres of origin. Genome 37:256–263
Beebe S, Skroch PW, Tohme J, Duque MC, Pedraza F, Nienhuis J (2000) Structure of genetic diversity among common bean landraces of Middle American origin based on correspondence analysis of RAPD. Crop Sci 40:264–273
Beebe S, Rengifo J, Gaitán E, Duque MC, Tohme J (2001) Diversity and origin of Andean landraces of common bean. Crop Sci 41:854–862
Blair MW, Pedraza F, Buendia HF, Gaitán-Solís E, Beebe SE, Gepts P, Tohme J (2003) Development of a genome-wide anchored microsatellite map for common bean (Phaseolus vulgaris L.). Theor Appl Genet 107:1362–1374
Chácon MI, Pickersgill SB, Debouck DG (2005) Domestication patterns in common bean (Phaseolus vulgaris L.) and the origin of the Mesoamerican and Andean cultivated races. Theor Appl Genet 110:432–444
Delgado-Salinas A, Turley T, Richman A, Lavin M (1999) Phylogenetic analysis of the cultivated and wild species of Phaseolus (Fabaceae). Syst Bot 24:438–460
Dubcovsky J, Dvorak J (1995) Ribosomal RNA multigene loci: nomads of the Triticeae genomes. Genetics 140:1367–1377
Frello S, Heslop-Harrison JS (2000) Chromosomal variation in Crocus vernus Hill (Iridaceae) investigated by in situ hybridization of rDNA and tandemly repeated sequence. Ann Bot 86:317–322
Freyre R, Skroch PW, Geffroy V, Adam-Blondon A-F, Shirmohamadali A, Johnson WC, Llaca V, Nodari RO, Pereira PA, Tsai S-M, Tohme J, Dron M, Nienhuis J, Vallejos CE, Gepts P (1998) Towards an integrated linkage map of common bean. 4. Development of a core linkage map and alignment of RFLP maps. Theor Appl Genet 97:847–856
Gepts P (1998) Origin and evolution of common bean: past events and recent trends. HortScience 33:1124–1130
Guerra M, Kenton A (1996) Distribution of telomere DNA in mitotic and polytene nuclei of the anther tapetum of a tetraploid hybrid bean Phaseolus vulgaris × P. acutifolius. Brazil J Genet 19:313–318
Guerra M, Kenton A, Bennett M (1996) rDNA sites in mitotic and polytene chromosomes of Vigna unguiculata (L.) Walp. and Phaseolus coccineus L. revealed by in situ hybridization. Ann Bot 78:157–161
Hanson RE, Islam-Faridi MN, Percival EA, Crane CF, Ji Y, McKnight TD, Stelly DM, Price HJ (1996) Distribution of 5S and 18S-28S rDNA loci in a tetraploid cotton (Gossypium hirsutum L.) and its putative diploid ancestors. Chromosoma 105:55–61
Hayasaki M, Morikawa T, Legget JM (2001) Intraspecific variation of 18S-5.8S-26S rDNA sites revealed by FISH and RFLP in wild oat, Avena agadiriana. Genes Genet Syst 76:9–14
Hayashi M, Miyahara A, Sato S, Kato T, Yoshikawa M, Taketa M, Hayashi M, Pedrosa A, Onda R, Imaizumi-Anraku H, Bachmair A, Sandal N, Stougaard J, Murooka Y, Tabata S, Kawasaki S, Kawaguchi M, Harada K (2001) Construction of a genetic linkage map of the model legume Lotus japonicus using an intraspecific F2 population. DNA Res 8:301–310
Heslop-Harrison JS, Schwarzacher T, Anamthawat-Jónsson K, Leitch AR, Shi M, Leitch IJ (1991) In situ hybridization with automated chromosome denaturation. Technique 3:109–115
Kami J, Becerra Velasquez V, Debouck DG, Gepts P (1995) Identification of presumed ancestral DNA sequences of phaseolin in Phaseolus vulgaris. Proc Natl Acad Sci USA 92:1101–1104
Koenig R, Gepts P (1989) Allozyme diversity in wild Phaseolus vulgaris: further evidence for two major centers of genetic diversity. Theor Appl Genet 78:809–817
Marcon AB, Barros IC, Guerra M (2005) Variation in chromosome numbers, CMA bands and 45S rDNA sites in species of Selaginella (Pteridophyta). Ann Bot 95:271–276
Melo NF, Guerra M (2003) Variability of the 5S and 45S rDNA sites in Passiflora L. species with distinct base chromosome numbers. Ann Bot 92:309–316
Moscone EA, Klein F, Lambrou M, Fuchs J, Schweizer D (1999) Quantitative karyotyping and dual-color FISH mapping of 5S and 18S-25S rDNA probes in the cultivated Phaseolus species (Leguminosae). Genome 42:1224–1233
Navratilova A, Neumann P, Macas J (2003) Karyotype analysis of four Vicia species using in situ hybridization with repetitive sequences. Ann Bot 91:921–926
Papa R, Gepts P (2003) Asymmetry of gene flow and differential geographical structure of molecular diversity in wild and domesticated common bean (Phaseolus vulgaris L.) from Mesoamerica. Theor Appl Genet 106:239–250
Pedrosa A, Jantsch MF, Moscone EA, Ambros PF, Schweizer D (2001) Characterisation of pericentromeric and sticky intercalary heterochromatin in Ornithogalum longibracteatum (Hyacinthaceae). Chromosoma 110:203–213
Pedrosa A, Sandal N, Stougaard J, Schweizer D, Bachmair A (2002) Chromosomal map of the model legume Lotus japonicus. Genetics 161:1661–1672
Pedrosa A, Vallejos CE, Bachmair A, Schweizer D (2003) Integration of common bean (Phaseolus vulgaris L.) linkage and chromosomal maps. Theor Appl Genet 106:205–212
Pich U, Fuchs J, Schubert I (1996) How do Alliaceae stabilize their chromosome ends in the absence of TTTAGGG sequences? Chromosome Res 4:207–213
Raskina O, Belyayev A, Nevo E (2004a) Activity of the En/Spm-like transposons in meiosis as a base for chromosome repatterning in a small, isolated, peripheral population of Aegilops speltoides Tausch. Chromosome Res 12:153–161
Raskina O, Belyayev A, Nevo E (2004b) Quantum speciation in Aegilops: molecular cytogenetic evidence from rDNA cluster variability in natural populations. Proc Natl Acad Sci USA 101:14818–14823
Schubert I, Wobus U (1985) In situ hybridization confirms jumping nucleolus organizing regions in Allium. Chromosoma 92:143–148
Sharma S, Raina SN (2005) Organization and evolution of highly repeated satellite DNA sequences in plant chromosomes. Cytogenet Genome Res 109:15–26
Shishido R, Sano Y, Fukui K (2000) Ribosomal DNAs: an exception to the conservation of gene order in rice genomes. Mol Gen Genet 263:586–591
Singh SP, Gepts P, Debouck DG (1991) Races of common bean (Phaseolus vulgaris, Fabaceae). Econ Bot 45:379–396
Sonnante G, Stockton T, Nodari RO, Becerra Velásquez VL, Gepts P (1994) Evolution of genetic diversity during the domestication of common-bean (Phaseolus vulgaris L.). Theor Appl Genet 89:629–635
Tohme J, González DO, Beebe S, Duque MC (1996) AFLP analysis of gene pools of a wild bean core collection. Crop Sci 36:1375–1384
Vaio M, Speranza P, Valls JF, Guerra M, Mazzella C (2005) Localization of the 5S and 45S rDNA sites and cpDNA sequence analysis in species of the Quadrifaria group of Paspalum (Poaceae, Paniceae). Ann Bot. DOI 10.1093/aob/mci168
Vallejos CE, Sakiyama NS, Chase CD (1992) A molecular marker-based linkage map of Phaseolus vulgaris L. Genetics 131:733–740
Wanzenböck E-M, Schöfer C, Schweizer D, Bachmair A (1997) Ribosomal transcription units integrated via T-DNA transformation associate with the nucleolus and do not require upstream repeat sequences for activity in Arabidopsis thaliana. Plant J 11:1007–1016
Wendel JF, Schnabel A, Seelanan T (1995) Bidirectional interlocus concerted evolution following allopolyploid speciation in cotton (Gossypium). Proc Natl Acad Sci USA 92:280–284
Zhang D, Sang T (1999) Physical mapping of ribosomal RNA genes in the peonies (Paeonia, Paeoniaceae) by fluorescent in situ hybridization: implications for phylogeny and concerted evolution. Am J Bot 86:735–740
Zheng J, Irifune K, Hirai K, Nakata M, Tanaka R, Morikawa H (1994) In situ hybridization to metaphase chromosomes in six species of Phaseolus and Vigna using ribosomal DNA as the probe. J Plant Res 107:365–369
Acknowledgements
A.P-H. was partially supported by a grant from the Conselho Nacional de Desenvolvimento Científico e Tecnológico (CNPq), Brazil, and by the Hertha-Firnberg program from the Austrian Science Fund (FWF) on behalf of the Federal Ministry for Education, Science and Culture (BMBWK), Austria. C.C.S.A. was supported by a grant from CNPq, Brazil. The work was partially supported by a grant to D.S. from the Gregor Mendel Institute of Molecular Plant Biology (GMI), Austria.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by J. S. Heslop-Harrison
Rights and permissions
About this article
Cite this article
Pedrosa-Harand, A., de Almeida, C.C.S., Mosiolek, M. et al. Extensive ribosomal DNA amplification during Andean common bean (Phaseolus vulgaris L.) evolution. Theor Appl Genet 112, 924–933 (2006). https://doi.org/10.1007/s00122-005-0196-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-005-0196-8