Abstract
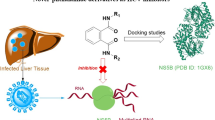
Hepatitis C virus (HCV) genotype 4a (GT4a) is prevalent in Egypt. It did not gain the necessary scientific focus despite its high resistance. Since the crystal structure NS5B (RNA-dependent RNA polymerase) of HCV GT4a has not been resolved until now, homology modeling was conducted to build and validate the 3D model of the enzyme. Ligand binding sites including the allosteric thumb II pocket were detected and used in lead optimization. Sixty new 4-thiazolidinone derivatives have been virtually designed and docked into thumb II site of HCV NS5B GT4a using rigid docking approach. Eighteen compounds (7a–r) that show good docking scores were synthesized and tested in vitro against NS5B GT4a. Compounds 7b and 7n showed the best inhibitory activity (IC50 = 0.338 and 0.342 µM, respectively). Compounds 7a, 7b, 7c, 7d, 7k, 7n, 7q, and 7r that have IC50 values less than 2 µM were assessed for cellular anti-HCV GT4a activity using human hepatoma cell line (Huh 7.5). The percentages of viral growth inhibition are between 79.67 and 94.77%. Compound 7b is the most active in the in vitro and cellular assays and could be considered a potential new lead for future anti-HCV studies.














Similar content being viewed by others
References
WHO. Hepatitis C fact sheet. 2019. https://www.who.int/news-room/fact-sheets/detail/hepatitis-c. Accessed July 29 2020.
Jafri SM, Gordon SC. Epidemiology of hepatitis C. Clin Liver Dis. 2018;12:140–2. https://doi.org/10.1002/cld.783
Alexopoulou A, Karayiannis P. Interferon-based combination treatment for chronic hepatitis C in the era of direct acting antivirals. Ann Gastroenterol. 2015;28:55–65.
Smith K. Sofosbuvir: a new milestone in HCV treatment? Nat Rev Gastroenterol Hepatol. 2013;10:258- https://doi.org/10.1038/nrgastro.2013.60
Götte M, Feld JJ. Direct-acting antiviral agents for hepatitis C: structural and mechanistic insights. Nat Rev Gastroenterol Hepatol. 2016;13:338–51. https://doi.org/10.1038/nrgastro.2016.60
Legrand-Abravanel F, Henquell C, Le Guillou-Guillemette H, Balan V, Mirand A, Dubois M, et al. Naturally occurring substitutions conferring resistance to hepatitis C virus polymerase inhibitors in treatment-naive patients infected with genotypes 1-5. Antivir Ther. 2009;14:723–30.
El-Tahan RR, Ghoneim AM, Zaghloul H. Dissection of two drug-targeted regions of Hepatitis C virus subtype 4a infecting Egyptian patients. Virus Genes. 2020. https://doi.org/10.1007/s11262-020-01776-y.
Bartenschlager R, Lohmann V, Penin F. The molecular and structural basis of advanced antiviral therapy for hepatitis C virus infection. Nat Rev Microbiol. 2013;11:482–96. https://doi.org/10.1038/nrmicro3046
Di Maio VC, Cento V, Mirabelli C, Artese A, Costa G, Alcaro S, et al. Hepatitis C virus genetic variability and the presence of NS5B resistance-associated mutations as natural polymorphisms in selected genotypes could affect the response to NS5B inhibitors. Antimicrob Agents Chemother. 2014;58:2781–97. https://doi.org/10.1128/aac.02386-13
Watkins WJ. Evolution of HCV NS5B non-nucleoside inhibitors. In: Sofia M, editor. HCV: the journey from discovery to a cure. Cham: Springer; 2019. p. 171–91.
Li H, Tatlock J, Linton A, Gonzalez J, Borchardt A, Dragovich P, et al. Identification and structure-based optimization of novel dihydropyrones as potent HCV RNA polymerase inhibitors. Bioorg Med Chem Lett. 2006;16:4834–8. https://doi.org/10.1016/j.bmcl.2006.06.065
Li P, Dorsch W, Lauffer DJ, Bilimoria D, Chauret N, Court JJ, et al. Discovery of novel allosteric HCV NS5B inhibitors. 2. lactam-containing thiophene carboxylates. ACS Med Chem Lett. 2017;8:251–5. https://doi.org/10.1021/acsmedchemlett.6b00479
Yan S, Appleby T, Larson G, Wu JZ, Hamatake R, Hong Z, et al. Structure-based design of a novel thiazolone scaffold as HCV NS5B polymerase allosteric inhibitors. Bioorg Med Chem Lett. 2006;16:5888–91. https://doi.org/10.1016/j.bmcl.2006.08.056
Li H, Tatlock J, Linton A, Gonzalez J, Jewell T, Patel L, et al. Discovery of (R)-6-cyclopentyl-6-(2-(2,6-diethylpyridin-4-yl)ethyl)-3-((5,7-dimethyl-[1,2,4]triazolo[1,5-a]pyrimidin-2-yl)methyl)-4-hydroxy-5,6-dihydropyran-2-one (PF-00868554) as a potent and orally available hepatitis C virus polymerase inhibitor. J Med Chem. 2009;52:1255–8. https://doi.org/10.1021/jm8014537
Lazerwith SE, Lew W, Zhang J, Morganelli P, Liu Q, Canales E, et al. Discovery of GS-9669, a thumb site II non-nucleoside inhibitor of NS5B for the treatment of genotype 1 chronic hepatitis C infection. J Med Chem. 2014;57:1893–901. https://doi.org/10.1021/jm401420j
Shi ST, Herlihy KJ, Graham JP, Nonomiya J, Rahavendran SV, Skor H, et al. Preclinical characterization of PF-00868554, a potent nonnucleoside inhibitor of the hepatitis C virus RNA-dependent RNA polymerase. Antimicrob Agents Chemother. 2009;53:2544–52. https://doi.org/10.1128/aac.01599-08
Ding Y, Smith KL, Varaprasad CVNS, Chang E, Alexander J, Yao N. Synthesis of thiazolone-based sulfonamides as inhibitors of HCV NS5B polymerase. Bioorg Med Chem Lett. 2007;17:841–5. https://doi.org/10.1016/j.bmcl.2006.08.104
Babaoglu K, Page MA, Jones VC, McNeil MR, Dong C, Naismith JH, et al. Novel inhibitors of an emerging target in Mycobacterium tuberculosis; substituted thiazolidinones as inhibitors of dTDP-rhamnose synthesis. Bioorg Med Chem Lett. 2003;13:3227–30. https://doi.org/10.1016/S0960-894X(03)00673-5
Küçükgüzel G, Kocatepe A, De Clercq E, Şahin F, Güllüce M. Synthesis and biological activity of 4-thiazolidinones, thiosemicarbazides derived from diflunisal hydrazide. Eur J Med Chem. 2006;41:353–9. https://doi.org/10.1016/j.ejmech.2005.11.005
Ottanà R, Carotti S, Maccari R, Landini I, Chiricosta G, Caciagli B, et al. In vitro antiproliferative activity against human colon cancer cell lines of representative 4-thiazolidinones. Part I. Bioorg Med Chem Lett. 2005;15:3930–3. https://doi.org/10.1016/j.bmcl.2005.05.093
Kumar A, Rajput CS, Bhati SK. Synthesis of 3-[4’-(p-chlorophenyl)-thiazol-2’-yl]-2-[(substituted azetidinone/thiazolidinone)-aminomethyl]-6-bromoquinazolin-4-ones as anti-inflammatory agent. Bioorg Med Chem. 2007;15:3089–96. https://doi.org/10.1016/j.bmc.2007.01.042.
Rawal RK, Tripathi R, Katti SB, Pannecouque C, De Clercq E. Design, synthesis, and evaluation of 2-aryl-3-heteroaryl-1,3-thiazolidin-4-ones as anti-HIV agents. Bioorg Medicinal Chem. 2007;15:1725–31. https://doi.org/10.1016/j.bmc.2006.12.003
Çakır G, Küçükgüzel İ, Guhamazumder R, Tatar E, Manvar D, Basu A, et al. Novel 4-thiazolidinones as non-nucleoside inhibitors of hepatitis C virus NS5B RNA-dependent RNA polymerase. Arch Pharm. 2015;348:10–22. https://doi.org/10.1002/ardp.201400247
Küçükgüzel İ, Satılmış G, Gurukumar KR, Basu A, Tatar E, Nichols DB, et al. 2-Heteroarylimino-5-arylidene-4-thiazolidinones as a new class of non-nucleoside inhibitors of HCV NS5B polymerase. Eur J Med Chem. 2013;69:931–41. https://doi.org/10.1016/j.ejmech.2013.08.043
UniProt Consortium. UniProt: the universal protein knowledgebase. Nucleic Acids Res. 2018;46:2699- https://doi.org/10.1093/nar/gky092
Xu D, Zhang Y. Improving the physical realism and structural accuracy of protein models by a two-step atomic-level energy minimization. Biophys J. 2011;101:2525–34. https://doi.org/10.1016/j.bpj.2011.10.024
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, et al. SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res. 2018;46:W296–W303. https://doi.org/10.1093/nar/gky427
Barreca ML, Iraci N, Manfroni G, Cecchetti V. Allosteric inhibition of the hepatitis C virus NS5B polymerase: in silico strategies for drug discovery and development. Future Med Chem. 2011;3:1027–55. https://doi.org/10.4155/fmc.11.53
Shih M-H, Wu C-L. Efficient syntheses of thiadiazoline and thiadiazole derivatives by the cyclization of 3-aryl-4-formylsydnone thiosemicarbazones with acetic anhydride and ferric chloride. Tetrahedron. 2005;61:10917–25. https://doi.org/10.1016/j.tet.2005.08.107
Vicini P, Geronikaki A, Anastasia K, Incerti M, Zani F. Synthesis and antimicrobial activity of novel 2-thiazolylimino-5-arylidene-4-thiazolidinones. Bioorg Medicinal Chem. 2006;14:3859–64. https://doi.org/10.1016/j.bmc.2006.01.043
Vicini P, Geronikaki A, Incerti M, Zani F, Dearden J, Hewitt M. 2-Heteroarylimino-5-benzylidene-4-thiazolidinones analogues of 2-thiazolylimino-5-benzylidene-4-thiazolidinones with antimicrobial activity: synthesis and structure–activity relationship. Bioorg Med Chem. 2008;16:3714–24. https://doi.org/10.1016/j.bmc.2008.02.001
Behbehani H, Ibrahim HM. 4-Thiazolidinones in heterocyclic synthesis: synthesis of novel enaminones, azolopyrimidines and 2-arylimino-5-arylidene-4-thiazolidinones. Molecules. 2012;17:6362–85.
Cousins KR. ChemDraw Ultra 9.0. CambridgeSoft, 100 CambridgePark Drive, Cambridge, MA 02140. www. cambridgesoft.com. See Web site for pricing options. J Am Chem Soc. 2005;127:4115–6. https://doi.org/10.1021/ja0410237.
Tu G, Li S, Huang H, Li G, Xiong F, Mai X, et al. Novel aminopeptidase N inhibitors derived from 1,3,4-thiadiazole scaffold. Bioorg Med Chem. 2008;16:6663–8. https://doi.org/10.1016/j.bmc.2008.05.081
Sun J-M, Kim S-J, Kim G-W, Rhee J-K, Kim ND, Jung H, et al. Inhibition of hepatitis C virus replication by monascus pigment derivatives that interfere with viral RNA polymerase activity and the mevalonate biosynthesis pathway. J Antimicrob Chemother. 2011;67:49–58. https://doi.org/10.1093/jac/dkr432
Hassan GS, Georgey HH, Mohammed EZ, Omar FA. Anti-hepatitis-C virus activity and QSAR study of certain thiazolidinone and thiazolotriazine derivatives as potential NS5B polymerase inhibitors. Eur J Med Chem. 2019;184:111747–62. https://doi.org/10.1016/j.ejmech.2019.111747
Howe AYM, Bloom J, Baldick CJ, Benetatos CA, Cheng H, Christensen JS, et al. Novel nonnucleoside inhibitor of hepatitis C virus RNA-dependent RNA polymerase. Antimicrob Agents Chemother. 2004;48:4813–21. https://doi.org/10.1128/aac.48.12.4813-4821.2004
Ye J, McGinnis S, Madden TL. BLAST: improvements for better sequence analysis. Nucleic acids Res. 2006;34:6–9. https://doi.org/10.1093/nar/gkl164
Nero TL, Parker MW, Morton CJ. Protein structure and computational drug discovery. Biochem Soc Trans. 2018;46:1367–79. https://doi.org/10.1042/bst20180202
Hawkins PCD, Skillman AG, Warren GL, Ellingson BA, Stahl MT. Conformer generation with OMEGA: Algorithm and validation using high quality structures from the protein databank and cambridge structural database. J Chem Inf Model. 2010;50:572–84. https://doi.org/10.1021/ci100031x
McGann M. FRED and HYBRID docking performance on standardized datasets. J Comput Aided Mol Des. 2012;26:897–906. https://doi.org/10.1007/s10822-012-9584-8
BIOVIA. Discovery studio Visualizer v19.1.0.18287. San Diego: Dassault Systèmes 2019.
DeLano WL. Pymol: an open-source molecular graphics tool. CCP4 Newsl Protein Crystallogr. 2002;40:82–92.
Acknowledgements
We are grateful for OpenEye Scientific Software for supporting the academic license granted to MAE laboratory. We also thank Dr. Khaled M. Elokely, Tanta University, for his help in analyzing the computational data and Dr. Ahmed E. Goda, Tanta University, for his help in the biological studies.
Author contributions
MAE contributed to conceptualization and design. ASA-B collected the data and carried out the experimental section. All authors analyzed the data, wrote, and revised the manuscript.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
About this article
Cite this article
Al-Behery, A.S., Elberembally, K.M. & Eldawy, M.A. Synthesis, docking, and biological evaluation of thiazolidinone derivatives against hepatitis C virus genotype 4a. Med Chem Res 30, 1151–1165 (2021). https://doi.org/10.1007/s00044-021-02721-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-021-02721-w