Abstract
Somatic mutations originate from both exogenous (e.g. UV radiation, chemical agents) and endogenous (e.g., DNA replication, defective DNA repair) sources and can have significant impacts on an animal’s reproductive success. This may be especially true for haploid organisms that are susceptible to any deleterious alleles inherited from their parent and any that arise over their lifetime. Unfortunately, little is known about the rate of somatic mutation accumulation across individuals and tissues of haplodiploid animal populations, the functional processes through which they arise, and their distribution across tissues and the genome. Here, we generated short-read whole-genome sequencing data for four tissues of haploid honey bee males. We paired this with estimates of telomere length and tissue-specific DNA content to address three major questions: is there variance in somatic mutational load across haploid individuals and specific tissues therein, does increased DNA content in a tissue contribute to somatic mutational load, and does telomere length correlate with mutational load? Our results suggest that variance in somatic mutational load is better captured across individuals than across tissues, that tissue-specific DNA content is not associated with somatic mutation load, and that variance in telomere length does not correlate with somatic mutation loads across tissues. To our knowledge, this is the first observational study on somatic mutational load in Apoidea and likely Hymenoptera. It serves as a useful advent for additional studies understanding the processes through which haploids tolerate or repair somatic mutations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Mutations drive evolution. They may alter protein sequence, change binding affinity, alter gene expression, or do nothing at all (Chatterjee and Walker, 2017; Friedman, 2022; Rockman and Kruglyak, 2006) but it is their change in frequency that, by basic definition, is evolution. Most mutations have little to no effect on an organism’s reproductive success and mutational frequency in a population changes as a result of drift and linked selection (Lynch et al., 2016; Ohta and Kimura, 1969). However, when a mutation does impact fitness, its frequency and the frequencies of any surrounding linked mutations can rapidly change in a population (Achatz and Zambetti, 2016; Inusa et al., 2019). This Neo-Darwinian view of evolution can be applied to any heritable mutation (Campbell and Eichler, 2013; Gerlinger et al., 2014).
All mutations arise through endogenous or exogenous processes. DNA replication is one such endogenous process. Each time a copy of the genome is made, errors can accrue, albeit at a low frequency (Alberts, 2003; Kunkel, 2004). Aging is an amalgamation of endogenous and exogenous stressors through time (Yousefzadeh et al., 2021). As an animal ages, its DNA has undergone many rounds of replication and has also likely been subjected to additional mutagens like UV radiation or oxidizing chemicals (Chatterjee and Walker, 2017; Hosamani and Muralidhara, 2013). Any copy of the genome in any cell of any animal can be mutated through these endogenous and exogenous processes. Therefore, most mutations arise as evolutionary ‘dead ends’ in somatic tissue that will not be inherited since only a small fraction of most animal cells are in the germ line (Surani et al., 2004) and most animals do not reproduce through fission (Howe et al., 2022); for a discussion on fission-inherited somatic mutations, see (Hanlon et al., 2019)
Germ line mutations can be passed from parent to offspring and therefore are the basis for most genetic work to date. Germ line mutations are the fuel for evolutionary change via drift or selection (Fan et al., 2008; Lynch et al., 2016), but see Hanlon et al., 2019. Heritable diseases have in part been extensively studied due to their ease of detection: germ line mutations arise pre-fertilization and therefore proliferate into every cell of an organism (Campbell & Eichler, 2013). Somatic mutations are rarely inherited in animals (but see these examples: (Bosch et al., 2009; Howe et al., 2022; Vasquez Kuntz et al., 2022) and may be less important to evolution; however, they do play critical roles in health, such as in cancer and age-related diseases (Greenman et al., 2007; Ikebe et al., 1995). Somatic mutations in animals, especially non-humans, have received little attention for their functional relevance. Not only does the soma have a greater proportion of cells able to undergo mutation compared to germ line, but also has an overall higher mutation rate: in both humans and mice, it has been shown that the mutation rate in the soma greatly exceeds that in the germ line (Milholland et al., 2017). The disposable soma theory predicts a trade-off between an organism’s health and its production of viable offspring (Lucas and Keller, 2014). It could explain an accumulation of mutations in somatic tissue, treating the soma as a repository for mutations, while preserving the germ line.
The endogenous and exogenous processes that generate somatic mutations may also lead to non-random mutation distributions within an individual. In humans, both the age and differential exogenous mutagenic exposure are correlated with variance in somatic mutation loads (Chatterjee and Walker, 2017; Lucas and Keller, 2014; Yousefzadeh et al., 2021). Sun-exposed skin, for example, accumulates more somatic mutations than covered skin and both have more mutations than hippocampal cells (García-Nieto et al., 2019). Further, replicating cells accumulate and then pass somatic mutations to their replicants, causing highly replicating tissues to accumulate more mutations (Bae et al., 2018; Cui et al., 2012).
Our understanding of somatic mutations in wild and managed animals is lacking: is there variance in the accumulation of somatic mutations across tissue types? Does DNA replication correlate with somatic mutation load? Does somatic mutation load increase with age? We proposed to investigate these questions using the honey bee Apis mellifera as a model. As with all Hymenoptera, A. mellifera are haplodiploid: males typically result from unfertilized eggs, and females result from fertilized eggs. Males (called drones) are large-bodied, readily collected in the field, and easily dissected. These haploid males provide a mutation identification advantage: as little as 7× coverage is needed to be confident in a male honey bee genotype from short-read sequencing (Wragg et al., 2016).
One potential source of mutation is through DNA replication: every round of DNA replication increases the chance that an error is made. The cell cycle state of some insect tissues is readily known (Britton and Edgar, 1998) and at least one tissue in honey bees, the thoracic muscle, undergoes early developmental endoreplication and becomes diploid (Aron et al., 2005; Glastad et al., 2014). Endoreplication is the cellular process of increasing DNA content through delineated doublings of the genome without division at the end of the cell cycle (Lee et al., 2009). Endoreplication works to generate function in differentiated cells, (Lee et al., 2009), but may also lead to an increased potential of DNA error.
Another potential source of mutation accumulation is through age-related processes. Aging is correlated with increased environmental exposure as well as general decline of maintenance machineries (Yousefzadeh et al., 2021). The short lifespan of the honey bee drone allows for the rapid analysis of aging (Currie, 1987). A honey bee’s lifespan varies with the seasons. Generally, queens are the longest living, with a lifespan of approximately 1–3 years, followed by drones, living between 13 and 38 days, and then workers are the shortest lived, with a summer lifespan of approximately 25–35 days (Amdam and Omholt, 2002).
In this study, we hypothesized that different tissues in an individual undergo different cellular processes, particularly in haplodiploid species undergoing tissue-specific endoreplication, and as such, predicted that endoreplication would lead to increased variance of somatic mutational load across tissues. We also hypothesized that individuals would experience different life histories, such as lifespan and exposure to oxidative stressors, and as such would lead to an increased variance of somatic mutation burden across individuals in a population. Here we present a unique survey study to examine some of the patterns and processes associated with somatic mutation burden in haploid individuals. Our work begins to investigate the variance of somatic mutations an individual has accumulated throughout development and early stages of life.
Methods
Sampling and DNA sequencing
Adult Apis mellifera drones were randomly sampled from multiple colonies (up to 6) at the Purdue Research Apiary as they returned from mating flights at colony entrances. The apiary was designed as a common garden and contains honey bees from various genetic backgrounds sourced from across the United States. While most drones of flying age are estimated to be between 5 and 16 days of age (Reyes et al., 2019), we did not track the precise ages of drones in order to obtain a random sample and assess the standing somatic variation. The drones were immediately flash-frozen and transported to an − 80 °C freezer. Flash-frozen samples were dissected in cold ethanol separating the brain, thoracic muscle, midgut, and reproductive tracts (Fig. 1). The reproductive tract of dissected drones included testes and accessory glands. DNA from individual tissues was extracted via a phenol/chloroform protocol (Martinson et al., 2011) and optimized for HMW DNA through the use of wide-bore pipette tips and gentle inversion by hand when mixing. Six individual drones (4 tissues each) were selected for sequencing based on DNA extraction quality from Qubit and Nanodrop 2000 readings. Following library preparation, tissue samples were independently sequenced to an average of 40× depth with pair ended, 150 bp reads on an Illumina NovaSeq 6000 (Novogene, CA).
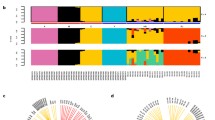
Somatic variance across tissues and individuals. A Dot plot of the number of somatic mutations for each of the 24 samples. Black lines represent the median across all tissues for individual drones. The number of mutations is separated by drone and color coded by tissue type. There were significant differences among drones (AOV; p < 0.05; TukeyHSD; B: A adj. p = 0.034, C: A adj. p = 0.012, D: A adj. p = 0.006, F: A adj. p = 0.019). B Bar plot of the reference allele frequency across bins for each tissue type. There was no significant difference between tissue types (AOV; p > 0.05). C Dendrogram illustrating allele frequency across individuals and tissues. Clustering occurs by individuals (A–F) as opposed to tissue type
Alignment and variant calling
Somatic mutations can be difficult to identify with short-read sequence technology. To identify tissue- and drone-specific somatic mutations, we used a pipeline developed by Hanlon et al (2019). Each single tissue genome sequence was aligned to the most recent honey bee reference genome (Amel_HAv3.1; (Wallberg et al., 2019) using BWA-MEM version 0.7.5a (Li, 2013). Aligned reads were then formatted, sorted, and indexed with SAMTOOLS version 1.16 (Li et al., 2009) prior to marking duplicate reads through PICARDTOOLS version 2.26.2. We identified variant sites in two ways. In our first method, we identified those that represent the majority of alleles within each tissue. We did this by joint variant calling with GATK’s haplotypecaller and genotypegvcf version 4.1.4.1 (McKenna et al., 2010). Each tissue contained diploid cells, so samples were treated as diploids (see results). To this data set, we applied several hard filters to remove low-quality sites and genotypes: GATK’s variant filtration filter QD2, QUAL30, SOR3, FS60, MQ40, MQRankSum-12.5, and ReadPosRankSum-20. These hard filters were chosen based on previous honey bee genomic work (Dogantzis et al., 2021). A total of 38,690 single tissue somatic mutations were observed after GATK hard filters and VCFTOOLS singletons identification (Table 1; Danecek et al., 2011). Because SNP variants may be skewed when overlapping with insertion–deletion mutation regions, we corrected for sites located near indels by removing somatic mutations ± 10 base pairs around indels. Mean read depth was filtered through VCFTOOLS version 0.1.16 (Danecek et al., 2011). Allelic depth was filtered on reference and alternate alleles with SAMTOOLS (Kofler et al., 2011). We also selected for somatic mutations with a minimum mean read depth of 20, a maximum mean read depth of 1000, and minimum genotype quality of 30. We employed a second method to call somatic mutations that used a pooled sequencing approach and called sites using POPOOLATION2 version 1.201 (Kofler et al., 2011). We calculated the reference allele frequency at all sites with between 14 and 100 reads coverage using popoolation2helper. This second approach allows us to estimate allele frequencies at each variant site for any number of alleles. Alignments and variant calls were manually checked across several megabases using IGV (Robinson et al., 2011). We use the individual somatic mutation dataset (first method) to represent high-confidence somatic mutations throughout, unless otherwise stated.
Telomere length estimation, functional prediction, and spatial distribution
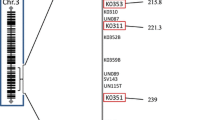
Telomere length can be used as a tool to indirectly measure aging and stress in an individual (Mather et al., 2011; Shalev et al., 2013). We estimated telomere length using TELSEQ (Ding et al., 2014) using the canonical telomeric repeat TTAGG that is present in honey bees. We optimized the number of required canonical repeats (k) by comparing telomere length to values k 1–15. A canonical repeat length of six was selected to predict telomere length in each tissue, as this was where the predicted length of the telomeres began to plateau in relation to repeat number. Functional predictions for all variants and transition/transversion ratios were determined by SNPEFF version 5.1 (Cingolani et al., 2012). SNPEFF was used to determine the features of single tissue mutations and which regions they occupy across the genome (Table 2; (Cingolani et al., 2012). Fisher’s pairwise tests were used to determine the significance of the number of somatic mutations across a region as well as their functionality (missense and silent). Pairwise tests were conducted between each tissue pairing: 2 × 2 contingency tables were built for testing the functional variants (S1) and regional variants (S2) and significant p values were adjusted for the 12 pairwise tests across functions and 24 pairwise tests across regions. The density of single tissue mutations was determined across each chromosome in 1 Mb windows across each of the four tissues.
Flow cytometry
Nineteen additional live honey bee drones and workers were collected randomly from the Purdue Research Apiary, as above, and kept live until dissection for flow cytometry analysis. In total, we live-dissected brain (N = 4 drones; 7 workers), thoracic muscle (N = 5; 5), midgut (N = 4; 6), and reproductive tract (N = 4 drone). Nuclei were prepared guided by the procedure outlined by Aron et al (2003). Individual live-dissected tissue samples were suspended in 450 µl Vindelov’s PI stain (Aron et al., 2003). Samples were homogenized (10 X mechanical grinding with pestle in 1.5 ml tube), added to an additional 450 µl of Vindelov’s PI stain, and passed through a 30 µm mesh to separate the nuclei from debris. The filtrate containing nuclei was topped to 1 mL with Vindelov’s PI stain and the samples were left on ice in darkness for 1 h before being quantified using an Attune Nxt Flow Cytometer. Samples were drawn at an acquisition volume of 650 µl and either 20,000 events or the entire volume was collected. Samples were discriminated on PI rather than on scatter to further reduce noise caused by unstained debris. To accurately compare among samples, haploid peaks were placed at channels 100–150 using a drone brain as a positive haploid control (Aron et al., 2005). Our procedure could not adequately resolve the ploidy of the midgut or reproductive tissues. This is a frequent challenge in the field (Aron et al., 2005) and we opted to remove them from our analyses. Data were initially visualized using Purdue flow cytometry software Xploid (Patsekin, 2019) prior to raw data being exported and analyzed.
Statistical analyses and data availability
Statistical analysis was performed using R version 4.2.2 through RStudio 2022 (R Core Team, 2021). All data were munged and processed using tidyverse (Wickham and Vaughan, 2023), Reshape2 (Wickham, 2007), lsr (Navarro, 2015), statpsych (Bonett and Calin-Jageman, 2023), and plyr (Wickham, 2011). Clustering allele frequencies across sites and samples was performed using the Euclidean distance and hierarchical clustering plotted with ggdendrogram, and ggdendro version 0.1.22 (Armitage et al., 1973; de Vries & Ripley, 2020). Functional prediction data were analyzed using R package Rstatix (Alboukadel, 2023). Flow cytometry data were processed using packages Flowcore (Hahne et al., 2009), FlowAI (Monaco et al., 2016), ggcyto (Van et al., 2018), and ggpubr (Kassambra, 2023). Specific statistical analyses are present in the results. Raw sequence data is available through NCBI BioProject PRJNA957323. Supplemental Table 1 provides a summary of data used for analyses and Supplemental Table 2 is a principle coordinate analysis among drone brains (Purcell et al., 2007) .
Results
Drones harbor many tissue-specific mutations
To identify somatic mutations in adult Apis mellifera drones, we analyzed whole-genome sequence data from four tissues: brain, muscle, midgut, and reproductive tract across six randomly-sampled drones. Samples were randomly collected from multiple colonies across the Purdue Research Apiary, which include a variety of honey bee breeds (Supplemental Table 2). Six individual drones were sequenced based on DNA extraction quality. We used two genotyping approaches to ensure we captured high-quality and representative single nucleotide polymorphisms (SNPs). Our first approach identified the major single mutations within tissues, likely to be those shared across most of the tissue and reflective of mutations accrued during development (Cui et al., 2012). Second, we quantified the allele frequency at each SNP site across individual genomes (e.g. treating the sample as pool-seq data; (Kofler et al., 2011). We discovered 13,632 putative tissue-specific mutations (average per drone per tissue ranges from 510 to 714; Table 1).
Endoreplication does not impact the number of mutations in a tissue and variation in mutational load is drone specific
While drones develop from unfertilized (haploid) eggs, the ploidy of their tissues varies: muscle tissue in drones endoreplicates and becomes diploid (Aron et al., 2005; Rangel et al., 2015). We predicted that endoreplicated tissue (muscle) will harbor more somatic mutations than haploid tissue when comparing tissues in the same cell cycle. We specifically tested if the number of somatic mutations varied between brain and muscle considering that they are in the same cell cycle (Britton and Edgar, 1998) and we could accurately estimate ploidy in both tissues (Fig. 2). We also predicted that tissues within drones would experience more consistent mutational inputs and predicted significant but consistent differences in the number of mutations among tissues. Given these two predictions, we tested for variance in mutational load across tissues and found no significant differences in the number of mutations between brains and muscles (Fig. 1A; AOV; F1,10 = 2.76, p = 0.064, adj. η2 = 0.138, 95% CI [0.14,0.53]) and nor for any other tissue comparison (Fig. 1A; AOV; F3,15 = 2.04, p = 0.152, adj. ηp2 = 0.148, 95% CI [0,0.49]). Similarly, using the pool-seq approach, we found no differences in allele frequency bins across tissues suggesting that there is an equivalent number of ‘new’ alleles across tissues within randomly selected individuals (Fig. 1B). We discovered no differences in any mutational frequency bin across tissues via the pool-seq approach (Fig. 1B; AOV; p > 0.05). Allele frequency did not cluster by tissue type, but rather by individual (Fig. 1C). We hypothesized drones would experience differential mutational inputs over their lifespans, and therefore we predicted significant differences in the number of mutations among drones. We found significant differences in the number of mutations among drones (Fig. 1A; AOV; F5,15 = 5.01, p = 0.00675, adj. ηp2 = 0.501, 95% CI [0.095, 0.72]).
Polyploidization across worker and drone tissues. A Heat map of DNA content for worker brain and muscle and drone brain and muscle. This combines all individuals in the dataset and observes the proportion of nuclei at each respective ploidy peak. The highest proportion of nuclei in each brain tissue of drones remained at 1C DNA content, indicative of a haploid peak. The highest proportion of nuclei in each muscle tissue of drones fell under 2C DNA content, indicative of a diploid peak. The relative proportion of nuclei in 2C DNA content of drone muscles matched that of what was seen in the nuclei of workers. B–E Histograms of the density of nuclei at each fluorescence peak for a single individual. The black dotted line indicates the haploid channel, set at channel 100–150. B Drone brain tissue remains haploid. C Drone muscle tissue endoreplicates. D, E Worker brain and muscle tissue remain diploid. The initial large peaks represent the G0/G1 phase whereas the smaller peaks represent nuclei in the G2 phase of the cell cycle and polyploidization (Aron et al., 2005)
Somatic mutations have the same functional composition as germ line mutations
Tissues vary in the mutational burden they can carry (García-Nieto et al., 2019) and so we predicted that there would be variation among tissues in the functional role of somatic mutations. We found that, across all tissues, the location of somatic mutations reflected the expected overall genomic expectation: most mutations fell into intronic and intergenic regions and fewest in exons (Table 2; Amel_HAv3.1; (Milholland et al., 2017; Wallberg et al., 2019). Within exons, we identified amino-acid changing mutations (missense mutations) and silent mutations. We found a significant difference in the number of missense mutations between brain and midgut (Fisher's exact test; p = 0.00109), midgut and reproductive (Fisher’s exact test; p = 5.36e-06), and between muscle and reproductive tissue (Fisher's exact test; p = 0.000129). Midgut tissue had the most mutational variance compared to other tissues and the least number of missense mutations. (S1 and S2). Aside from evaluating functional composition, we compared somatic mutation density within windows across all tissues for all our samples (Yin et al., 2021); S3). We found no significant evidence for differences in density among tissues following a Bonferroni correction (adj. p > 0.05 for all comparisons).
We also quantified the number of transitions and transversions by each tissue type to investigate the distribution of nucleotide substitutions across the genome. In general, there were more transitions than transversions (Fig. 3). Transitions did vary significantly by drone (Fig. 3A; AOV; F5,15 = 3.08, p = 0.0414, adj. ηp2 = 0.342, 95% CI [0,0.624]), while transversions did not (Fig. 3A; AOV; F5,15 = 1.68, p = 0.199, adj. ηp2 = 0.146, 95% CI [0,0.498]). It was revealed that drone A had a significantly higher number of transitions than drone D (Fig. 3A; TukeyHSD, p = 0.0205). Neither transitions (Fig. 3A; AOV; F3,15 = 2.3, p = 0.1187, adj. ηp2 = 0.178, 95% CI [0,0.514]) nor transversions (Fig. 3A; AOV; F3,15 = 2.57, p = 0.0933, adj. ηp2 = 0.207, 95% CI [0,0.534]) varied significantly by tissue type (Fig. 3B). Errors in Illumina sequencing platforms may lead to false mutations in a dataset and disproportionately arise in specific transitions or transversions (Kirsch and Klein, 2012). To investigate somatic mutations arising due to sequencing error, we compared ratios of CT:AG (t test; p = 0.9159) and GT:AC (t test; p = 0.945) in our somatic mutation dataset to SNPs and found no significant differences.
Transitions and transversions type. A Bar plot of the average number and type of transitions and transversions across tissues in each drone. B Bar plot of the average number and type of transitions and transversions in a specific tissue across all drones. Far-right columns summarize transitions (Ts) versus transversions (Tv). Ts significantly varied by the individual where drone A was significantly higher than drone D (Fig. 3A; AOV < 0.05; TukeyHSD, p = 0.0205)
Telomere length does not correlate with mutational load
In some animals, and indeed in some insects, telomere length is correlated with both organismal age and past stress (Jemielity et al., 2007; Mather et al., 2011). Telomeres act as a protective barrier for replication (and thus age) associated sequence loss (Jacobs, 2013). Telomere-initiated cellular senescence describes the process of shortening telomeres during rounds of incomplete DNA replication (Harley et al., 1990). Despite the activity of telomerase, telomeres tend to shorten over many rounds of DNA replication. In honey bees, early work suggested that telomere length is highly variable across individuals and that telomerase activity generally reduces as an individual ages (Korandová and Frydrychová, 2016). There has yet to be a definitive exploration of telomere length dynamics in honey bees; however, we hypothesized that if indeed telomeres reflect past age and stress in individual bees, then there should be a negative relationship between telomere length and the number of somatic mutations in a tissue. As with previous studies, we found substantial variation in telomere length (here, predicted length) among individuals (Fig. 4A; AOV; F5,15 = 5.596, p = 0.00417, ηp2 = 0.535, 95% CI [0.131,0.737]; (Jemielity et al., 2007). We did not find a significant variation in telomere length across tissue type (AOV; F3,15 = 2.218, p = 0.128, ηp2 = 0.169, 95% CI [0,0.508]). We found that there was no significant variance in the telomere length when compared to mutation accumulation (Fig. 4B; log10-corrected r22 = − 0.121, p > 0.05). We repeated this analysis to see if the number of mutations within each allele frequency bin correlated with telomere length but found no significant association (Pearson correlation; p > 0.05).
Average predicted telomere lengths and mutation accumulations across drones. A Dot plot of the telomere length of each of the 24 drone tissue samples. The black line indicates the median telomere length of each drone. There is significant variation of telomere length across individuals (AOV < 0.05; TukeyHSD; B: F adj. p = 0.0337, B: D adj. p = 0.0109, C: D adj p = 0.0331). B Scatter plot of the number of somatic variants and telomere lengths for all individuals, where distinct colors represent tissue type and diverse shapes denote individual drones. Overall, there was no significant correlation between the number of mutations and telomere length (log10 r22 = -0.121, p > 0.05)
Discussion
Here, we have developed the first tissue and individual atlas of somatic mutations for male honey bees. Somatic mutagenesis is understudied in animals due to several factors: a historical focus on heritable diseases, complications with methodologies to differentiate soma from germ line, and misconceptions about the fitness effects of somatic mutations outside cancer in humans (Campbell and Eichler, 2013; Greenman et al., 2007; Milholland et al., 2017). Our preliminary study shows that Apis mellifera drones harbor hundreds of tissue-specific mutations by the time they reach adulthood. With this data, we extracted somatic mutations from a population that had already undergone selection for the healthiest individuals (i.e., drones that have accrued lethal mutations during development would not be present in adult populations). Our first method of identifying tissue-specific somatic mutations aimed to identify variants that arose early in development. This method created a robust sample set of somatic mutations, successfully eliminating mutational artifacts; problematically this technique also eliminated low-frequency somatic mutations that arose during adulthood. To counter the removal of potentially novel low-frequency mutations, we performed a pooled sequencing analysis to call SNPs that arose during an individual’s lifetime post-development. While sequencing error can skew somatic mutation data, errors in Illumina sequencing technology typically lead to an excess of C > T transitions compared to A > G and G > T transversions relative to A > C (Kirsch and Klein, 2012). If our putative single tissue somatic mutations arose due to sequencing error, we might expect similarly elevated levels across our transitions and transversions. We did not find any significant differences in our ratios comparing our putative somatic mutations to all other SNPs in the dataset, and this increases our confidence that mutations did not arise due to sequencing error.
Through our two methods of calling single tissue somatic mutations, we provide insight into the somatic mutational landscape both within and across honey bee drones. We observed differences in mutational burden among individuals. This led us to explore the possible avenues that would lead to an increase in mutational load in drone A compared to most other individuals. We hypothesized that individuals would experience different life histories, and as such predicted to see variance in mutation number across our six individual drones. Older drones would have been subjected to increased rounds of DNA replication as well as increased environmental and metabolism stressors. We may predict that mutational load differences may be a consequence of any combination of these stressors, and subsequent research should be conducted to understand the exact mechanism. While we did observe mutational burden differences across individuals, we did not observe significant variance in mutational burden across tissues despite tissues of the same type sharing common development across individuals and undergoing differential development between tissue types (Fig. 1; (Newman, 1994; Strand et al., 2010). We do not think this is a result of sampling adult drones. If drones harboring deleterious alleles died prior to our sampling, and thus skewed our data, we would still expect to observe variance in neutral mutations across tissues (Table 2). On the one hand, we may propose that mutations are more likely to arise during development and cells share common developmental pathways. A study comparing human skeletal muscle, kidney tubules, blood, and fat progenitors found that many somatic mutations emerged from common cellular activities (Franco et al., 2019). On the other hand, the disposable soma theory hypothesizes that the soma of an organism is no longer pertinent after sexual maturity (Kirkwood, 1977). The disposable soma theory, thus, supports the prediction that an entire organism would accumulate mutations with age, regardless of tissue type. We did, however, find that midgut had a significantly lower number of mutations in terms of functional mutations, but a variable number of mutations across genomic regions (S1 and S2). The nature of the midgut, shedding and renewing cells continuously could explain its variance in somatic mutations (Zhang & Edgar, 2022). Future investigations should track mutational rate over drone development.
We hypothesized one potential contributor to somatic mutation load to be the endogenous process of endoreplication. A. mellifera males undergo endoreplication in at least one tissue. In Drosophila and likely A. mellifera, both neurogenesis and myogenesis become quiescent post-embryogenesis (Britton and Edgar, 1998). Since they are in the same cell state, we were able to accurately compare brain and muscle tissue as our representative non-endoreplicated and endoreplicated tissues (Aron et al., 2005). As expected, brain tissue was confirmed haploid and muscle was confirmed diploid (Fig. 2A–C). Surprisingly, we found no significant increase in mutation load in endoreplicated tissue. This may be due to the nature of mutation repair in haploids. Haploid organisms lack a template for homology-directed repair pathways (Chatterjee and Walker, 2017; Liang et al., 1998; Mourrain and Boissonneault, 2021), allowing for the potential for unrepaired mutations to accumulate by means of less efficient pathways. This includes the potential for haploid brain tissue to accumulate mutations through a lack of efficient repair systems, such as non-homologous end joining (Mourrain and Boissonneault, 2021). In muscle tissue, with increased cycles of endoreplication, there is the potential for increased chances of error (Lee et al., 2009). The net result of brain being haploid and muscle-cell endoreplication may be no difference in mutations at adulthood. We know little about how mutations are repaired in haplodiploid insects and even less about how repair may vary by tissue. In addition, while we did not observe significantly higher number of mutations in muscle tissue as compared to brain tissue, there was a large effect size (see Results). Although our sample size for this observational study was limited, studies to search for somatic mutations may use single cells from a single organism (Milholland et al., 2017) or across samples from many individuals (Cagan et al., 2022). Increasing sample size of number of individuals may have given the precision necessary to observe variance across tissues. For example, somatic mutation screens in humans have used hundreds of individuals and tissues therein to find subtle differences in the mutational landscape among individuals (Ren et al., 2022). Ultimately, more work needs to be done to study repair mechanisms in haploids, especially to investigate the burden of non-homologous repair (Caldecott, 2008; Chatterjee and Walker, 2017; Mourrain and Boissonneault, 2021).
In many animals, as an individual ages, telomere length shortens, leading to potential DNA damage in aging organisms (Mather et al., 2011) and a correlation between telomere length and the number of somatic mutations genome wide. We used a novel telomere length prediction algorithm (Ding et al., 2014) within tissues and found that using canonical honey bee telomere sequence, we obtained telomere length estimates within the range expected for bees (Korandová and Frydrychová, 2016). Our approach found that shorter telomeres do not directly correlate with higher somatic mutation loads (Fig. 4B). Accordingly, telomeres may not be a good indicator of age in insects; there is a high variability of telomere length and lifespan across many species (Walter et al., 2007; Wright and Shay, 2000). Perhaps telomere length is a better indicator of developmental stages, as it has been shown to decrease quickly during development and remain largely unchanged once an insect has reached adulthood (Jemielity et al., 2007). As with mutational accumulation variance across tissue type, we again had a large effect size when comparing mutation accumulation to telomere length. To accurately elucidate the link between somatic mutation accumulation and aging, this experiment would best be repeated to corroborate telomere length with an increased number of aged individuals or those that have been stressed.
This study has helped us to better understand the distribution of somatic mutations across and within haploid insects and provides a leaping-off point for additional work into somatic mutations. While somatic mutations have a significant impact on disease, cancer, and reproductive success (Greenman et al., 2007; Inusa et al., 2019; Lucas and Keller, 2014), the distribution of somatic mutations across and within organisms remains largely uncharacterized. We were unable to answer questions regarding somatic distribution, mechanisms of mutagenesis, sampling bias, somatic compensation, and mechanisms of somatic repair due to how little exploration has been conducted in the field. We lack study on a robust model in which we can easily elucidate somatic from germ line variants. The western honey bee, A. mellifera, provides a powerful model system to build an atlas of somatic mutations. Prior studies have suggested or refuted that there are tissue-specific differences, but little had been done to directly compare mutation accumulation within an individual (Franco et al., 2019; Lucas and Keller, 2014). While tissues have little variance in somatic mutation accumulation, further investigation should be conducted on a cell-specific level. Our results also have interesting implications for genomics in social insects more broadly. Variation in the accumulation of mutations across individuals and their tissues could lead to erroneous population genomic conclusions, especially for rare mutations. Understanding somatic mutation distribution in haploid drones has laid the foundation for future research into tissue-specific repair mechanisms. DNA repair varies significantly between germ line and soma. Mutation repair could be influenced by tissue-specific processes, undergo differential DNA damage response when exposed to stressors. Future studies on DNA damage and repair may have implications for therapies involving longevity and disease.
Data availability
Raw sequence data are available through NCBI BioProject PRJNA957323.
References
Achatz MI, Zambetti GP (2016) The inherited p53 mutation in the brazilian population. Cold Spring Harb Perspect Med 6(12):a026195. https://doi.org/10.1101/cshperspect.a026195
Alberts B (2003) DNA replication and recombination. Nature. https://doi.org/10.1038/nature01407
Alboukadel K (2023). rstatix: pipe-friendly framework for basic statistical tests (0.7.2) [Computer software]. https://rpkgs.datanovia.com/rstatix/. Accessed 24 Aug 2023
Amdam GV, Omholt SW (2002) The regulatory anatomy of honeybee lifespan. J Theor Biol 216(2):209–228. https://doi.org/10.1006/jtbi.2002.2545
Armitage P, Miller A, Rodger CD, Tregear RT (1973) The structure and function of insect muscle. Cold Spring Harb Symp Quant Biol 37:379–387. https://doi.org/10.1101/SQB.1973.037.01.048
Aron S, De Menten L, Van Bockstaele D (2003) Brood sex ratio determination by flow cytometry in ants. Mol Ecol Notes 3(3):471–475. https://doi.org/10.1046/j.1471-8286.2003.00488.x
Aron S, de Menten L, Van Bockstaele DR, Blank SM, Roisin Y (2005) When hymenopteran males reinvented diploidy. Curr Biol 15(9):824–827. https://doi.org/10.1016/j.cub.2005.03.017
Bae T, Tomasini L, Mariani J, Zhou B, Roychowdhury T, Franjic D, Pletikos M, Pattni R, Chen B-J, Venturini E, Riley-Gillis B, Sestan N, Urban AE, Abyzov A, Vaccarino FM (2018) Different mutational rates and mechanisms in human cells at pregastrulation and neurogenesis. Science 359(6375):550–555. https://doi.org/10.1126/science.aan8690
Bonett D, Calin-Jageman R (2023) statpsych: statistical methods for psychologists (1.4.0) [Computer software]. https://CRAN.R-project.org/package=statpsych. Accessed 10 Aug 2023
Bosch TCG, Anton-Erxleben F, Hemmrich G, Khalturin K (2009) The Hydra polyp: nothing but an active stem cell community: the hydra polyp: a stem cell community. Dev Growth Differ 52(1):15–25. https://doi.org/10.1111/j.1440-169X.2009.01143.x
Britton JS, Edgar BA (1998) Environmental control of the cell cycle in Drosophila: nutrition activates mitotic and endoreplicative cells by distinct mechanisms. Development 125(11):2149–2158. https://doi.org/10.1242/dev.125.11.2149
Cagan A, Baez-Ortega A, Brzozowska N, Abascal F, Coorens THH, Sanders MA, Lawson ARJ, Harvey LMR, Bhosle S, Jones D, Alcantara RE, Butler TM, Hooks Y, Roberts K, Anderson E, Lunn S, Flach E, Spiro S, Januszczak I, Martincorena I (2022) Somatic mutation rates scale with lifespan across mammals. Nature. https://doi.org/10.1038/s41586-022-04618-z
Caldecott KW (2008) Single-strand break repair and genetic disease. Nature Rev Genet. https://doi.org/10.1038/nrg2380
Campbell CD, Eichler EE (2013) Properties and rates of germline mutations in humans. Trends Genet 29(10):575–584. https://doi.org/10.1016/j.tig.2013.04.005
Chatterjee N, Walker GC (2017) Mechanisms of DNA damage, repair and mutagenesis. Environ Mol Mutagen 58(5):235–263. https://doi.org/10.1002/em.22087
Cingolani P, Platts A, Wang LL, Coon M, Nguyen T, Wang L, Land SJ, Lu X, Ruden DM (2012) A program for annotating and predicting the effects of single nucleotide polymorphisms, SnpEff: SNPs in the genome of Drosophila melanogaster strain w 1118; iso-2; iso-3. Fly 6(2):80–92. https://doi.org/10.4161/fly.19695
Cui P, Ding F, Lin Q, Zhang L, Li A, Zhang Z, Hu S, Yu J (2012) Distinct contributions of replication and transcription to mutation rate variation of human genomes. Genomics Proteomics Bioinform 10(1):4–10. https://doi.org/10.1016/S1672-0229(11)60028-4
Currie RW (1987) The biology and behaviour of drones. Bee World 68(3):129–143. https://doi.org/10.1080/0005772X.1987.11098922
Danecek P, Auton A, Abecasis G, Albers CA, Banks E, DePristo MA, Handsaker RE, Lunter G, Marth GT, Sherry ST, McVean G, Durbin R (2011) The variant call format and VCFtools. Bioinformatics 27(15):2156–2158. https://doi.org/10.1093/bioinformatics/btr330
Ding Z, Mangino M, Aviv A, Spector T, Durbin R (2014) Estimating telomere length from whole genome sequence data. Nucleic Acids Res 42(9):e75. https://doi.org/10.1093/nar/gku181
Dogantzis KA, Tiwari T, Conflitti IM, Dey A, Patch HM, Muli EM, Garnery L, Whitfield CW, Stolle E, Alqarni AS, Allsopp MH, Zayed A (2021) Thrice out of Asia and the adaptive radiation of the western honey bee. Sci Adv 7(49):2151. https://doi.org/10.1126/sciadv.abj2151
Fan W, Waymire KG, Narula N, Li P, Rocher C, Coskun PE, Vannan MA, Narula J, MacGregor GR, Wallace DC (2008) A mouse model of mitochondrial disease reveals germline selection against severe mtDNA mutations. Science 319(5865):958–962. https://doi.org/10.1126/science.1147786
Franco I, Helgadottir HT, Moggio A, Larsson M, Vrtačnik P, Johansson A, Norgren N, Lundin P, Mas-Ponte D, Nordström J, Lundgren T, Stenvinkel P, Wennberg L, Supek F, Eriksson M (2019) Whole genome DNA sequencing provides an atlas of somatic mutagenesis in healthy human cells and identifies a tumor-prone cell type. Genome Biol 20(1):285. https://doi.org/10.1186/s13059-019-1892-z
Friedman R (2022) Computational studies of protein–drug binding affinity changes upon mutations in the drug target. WIREs Comput Mole Sci 12(1):e1563. https://doi.org/10.1002/wcms.1563
García-Nieto PE, Morrison AJ, Fraser HB (2019) The somatic mutation landscape of the human body. Genome Biol 20(1):298. https://doi.org/10.1186/s13059-019-1919-5
Gerlinger M, McGranahan N, Dewhurst SM, Burrell RA, Tomlinson I, Swanton C (2014) Cancer: evolution within a lifetime. Annu Rev Genet 48(1):215–236. https://doi.org/10.1146/annurev-genet-120213-092314
Glastad KM, Hunt BG, Yi SV, Goodisman MAD (2014) Epigenetic inheritance and genome regulation: is DNA methylation linked to ploidy in haplodiploid insects? Proc Royal Soc B: Biol Sci 281(1785):20140411. https://doi.org/10.1098/rspb.2014.0411
Greenman C, Stephens P, Smith R, Dalgliesh GL, Hunter C, Bignell G, Davies H, Teague J, Butler A, Stevens C, Edkins S, O’Meara S, Vastrik I, Schmidt EE, Avis T, Barthorpe S, Bhamra G, Buck G, Choudhury B, Stratton MR (2007) Patterns of somatic mutation in human cancer genomes. Nature. https://doi.org/10.1038/nature05610
Hahne F, LeMeur N, Brinkman RR, Ellis B, Haaland P, Sarkar D, Spidlen J, Strain E, Gentleman R (2009) flowCore: a bioconductor package for high throughput flow cytometry. BMC Bioinform 10(1):106. https://doi.org/10.1186/1471-2105-10-106
Hanlon VCT, Otto SP, Aitken SN (2019) Somatic mutations substantially increase the per-generation mutation rate in the conifer Picea sitchensis. Evol Lett 3(4):348–358. https://doi.org/10.1002/evl3.121
Harley CB, Futcher AB, Greider CW (1990) Telomeres shorten during ageing of human fibroblasts. Nature. https://doi.org/10.1038/345458a0
Hosamani R, Muralidhara. (2013) Acute exposure of Drosophila melanogaster to paraquat causes oxidative stress and mitochondrial dysfunction. Arch Insect Biochem Physiol 83(1):25–40. https://doi.org/10.1002/arch.21094
Howe J, Rink JC, Wang B, Griffin AS (2022) Multicellularity in animals: the potential for within-organism conflict. Proc Natl Acad Sci 119(32):e2120457119. https://doi.org/10.1073/pnas.2120457119
Ikebe S, Tanaka M, Ozawa T (1995) Point mutations of mitochondrial genome in Parkinson’s disease. Mol Brain Res 28(2):281–295. https://doi.org/10.1016/0169-328X(94)00209-W
Inusa BP, Hsu LL, Kohli N, Patel A, OminuEvbota K, Anie KA, Atoyebi W (2019) Sickle cell disease—genetics, pathophysiology, clinical presentation and treatment. Int J Neonatal Screening. https://doi.org/10.3390/ijns5020020
Jacobs J (2013) Loss of telomere protection: consequences and opportunities. Front Oncol. https://doi.org/10.3389/fonc.2013.00088
Jemielity S, Kimura M, Parker KM, Parker JD, Cao X, Aviv A, Keller L (2007) Short telomeres in short-lived males: what are the molecular and evolutionary causes? Aging Cell 6(2):225–233. https://doi.org/10.1111/j.1474-9726.2007.00279.x
Kassambara A (2023) ggpubr: ‘ggplot2’ Based Publication Ready Plots. R package version 0.6.0. https://rpkgs.datanovia.com/ggpubr/. Accessed 1 Mar 2023
Kirkwood TBL (1977) Evolution of ageing. Nature. https://doi.org/10.1038/270301a0
Kirsch S, Klein CA (2012) Sequence error storms and the landscape of mutations in cancer. Proc Natl Acad Sci USA 109(36):14289–14290. https://doi.org/10.1073/pnas.1212246109
Kofler R, Pandey RV, Schlötterer C (2011) PoPoolation2: Identifying differentiation between populations using sequencing of pooled DNA samples (Pool-Seq). Bioinformatics 27(24):3435–3436. https://doi.org/10.1093/bioinformatics/btr589
Korandová M, Frydrychová RČ (2016) Activity of telomerase and telomeric length in Apis mellifera. Chromosoma 125(3):405–411. https://doi.org/10.1007/s00412-015-0547-4
Kunkel TA (2004) DNA Replication Fidelity *. J Biol Chem 279(17):16895–16898. https://doi.org/10.1074/jbc.R400006200
Lee HO, Davidson JM, Duronio RJ (2009) Endoreplication: polyploidy with purpose. Genes Dev 23(21):2461–2477. https://doi.org/10.1101/gad.1829209
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25(16):2078–2079. https://doi.org/10.1093/bioinformatics/btp352
Li, H. (2013). Aligning sequence reads, clone sequences and assembly contigs with BWA-MEM (arXiv:1303.3997). arXiv. http://arxiv.org/abs/1303.3997
Liang F, Han M, Romanienko PJ, Jasin M (1998) Homology-directed repair is a major double-strand break repair pathway in mammalian cells. Proc Natl Acad Sci 95(9):5172–5177. https://doi.org/10.1073/pnas.95.9.5172
Lucas ER, Keller L (2014) Ageing and somatic maintenance in social insects. Current Opinion in Insect Science 5:31–36. https://doi.org/10.1016/j.cois.2014.09.009
Lynch, M., Ackerman, M. S., Gout, J.-F., Long, H., Sung, W., Thomas, W. K., & Foster, P. L. (2016). Genetic drift, selection and the evolution of the mutation rate. Nature Reviews Genetics, 17(11), Article 11. https://doi.org/10.1038/nrg.2016.104
Martinson VG, Danforth BN, Minckley RL, Rueppell O, Tingek S, Moran NA (2011) A simple and distinctive microbiota associated with honey bees and bumble bees. Mol Ecol 20(3):619–628. https://doi.org/10.1111/j.1365-294X.2010.04959.x
Mather KA, Jorm AF, Parslow RA, Christensen H (2011) Is telomere length a biomarker of aging? A review. J Gerontol: Series A 66A(2):202–213. https://doi.org/10.1093/gerona/glq180
McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, Garimella K, Altshuler D, Gabriel S, Daly M, DePristo MA (2010) The Genome analysis toolkit: a mapreduce framework for analyzing next-generation DNA sequencing data. Genome Res 20(9):1297–1303. https://doi.org/10.1101/gr.107524.110
Milholland B, Dong X, Zhang L, Hao X, Suh Y, Vijg J (2017) Differences between germline and somatic mutation rates in humans and mice. Nature Commun. https://doi.org/10.1038/ncomms15183
Monaco G, Chen H, Poidinger M, Chen J, de Magalhães JP, Larbi A (2016) flowAI: automatic and interactive anomaly discerning tools for flow cytometry data. Bioinformatics 32(16):2473–2480. https://doi.org/10.1093/bioinformatics/btw191
Mourrain L, Boissonneault G (2021) DNA Repair in Haploid Context. Int J Mole Sci. https://doi.org/10.3390/ijms222212418
Navarro D (2015) Learning statistics with R: A tutorial for psychology students and other beginners (0.6) [Computer software]. https://learningstatisticswithr.com. Accessed 4 Aug 2023
Newman SA (1994) Generic physical mechanisms of tissue morphogenesis: a common basis for development and evolution. J Evol Biol 7(4):467–488. https://doi.org/10.1046/j.1420-9101.1994.7040467.x
Ohta T, Kimura M (1969) Linkage disequilibrium at steady state determined by random genetic drift and recurrent mutation. Genetics 63(1):229–238
Patsekin V (2019) Xploid (2.13) [Computer software]. http://www.cyto.purdue.edu/Xploid. Accessed 16 Jun 2023
Purcell S, Neale B, Todd-Brown K, Thomas L, Ferreira MAR, Bender D, Maller J, Sklar P, de Bakker PIW, Daly MJ, Sham PC (2007) PLINK: a tool set for whole-genome association and population-based linkage analyses. Am J Hum Genet 81(3):559–575
Rangel J, Strauss K, Seedorf K, Hjelmen CE, Johnston JS (2015) Endopolyploidy changes with age-related polyethism in the honey bee Apis mellifera. PLoS ONE 10(4):e0122208. https://doi.org/10.1371/journal.pone.0122208
Ren P, Dong X, Vijg J (2022) Age-related somatic mutation burden in human tissues. Front Aging. https://doi.org/10.3389/fragi.2022.1018119
Reyes M, Crauser D, Prado A, Le Conte Y (2019) Flight activity of honey bee (Apis mellifera) drones. Apidologie 50(5):669–680. https://doi.org/10.1007/s13592-019-00677-w
Robinson JT, Thorvaldsdóttir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26. https://doi.org/10.1038/nbt.1754
Rockman MV, Kruglyak L (2006) Genetics of global gene expression. Nature Rev Genet. https://doi.org/10.1038/nrg1964
Shalev I, Entringer S, Wadhwa PD, Wolkowitz OM, Puterman E, Lin J, Epel ES (2013) Stress and telomere biology: a lifespan perspective. Psychoneuroendocrinology 38(9):1835–1842. https://doi.org/10.1016/j.psyneuen.2013.03.010
Strand DW, Franco OE, Basanta D, Anderson ARA, Hayward SW (2010) Perspectives on tissue interactions in development and disease. Curr Mol Med 10(1):95–112. https://doi.org/10.2174/156652410791065363
Surani MA, Ancelin K, Hajkova P, Lange U, Payer B, Western P, Saitou M (2004) Mechanism of mouse germ cell specification: a genetic program regulating epigenetic reprogramming. Cold Spring Harb Symp Quant Biol 69:1–9. https://doi.org/10.1101/sqb.2004.69.1
Van P, Jiang W, Gottardo R, Finak G (2018) ggCyto: next generation open-source visualization software for cytometry. Bioinformatics 34(22):3951–3953. https://doi.org/10.1093/bioinformatics/bty441
Vasquez Kuntz KL, Kitchen SA, Conn TL, Vohsen SA, Chan AN, Vermeij MJA, Page C, Marhaver KL, Baums IB (2022) Inheritance of somatic mutations by animal offspring. Sci Adv 8(35):eabn0707. https://doi.org/10.1126/sciadv.abn0707
de Vries A, Ripley BD (2024) ggdendro: Create Dendrograms and Tree Diagrams Using ‘ggplot2’. R package version 0.2.0. https://andrie.github.io/ggdendro/. Accessed 1 Mar 2023
Wallberg A, Bunikis I, Pettersson OV, Mosbech M-B, Childers AK, Evans JD, Mikheyev AS, Robertson HM, Robinson GE, Webster MT (2019) A hybrid de novo genome assembly of the honeybee, Apis mellifera, with chromosome-length scaffolds. BMC Genomics 20(1):275. https://doi.org/10.1186/s12864-019-5642-0
Walter MF, Biessmann MR, Benitez C, Török T, Mason JM, Biessmann H (2007) Effects of telomere length in Drosophila melanogaster on life span, fecundity, and fertility. Chromosoma 116(1):41–51. https://doi.org/10.1007/s00412-006-0081-5
Wickham H (2007) Reshaping data with the reshape package. J Stat Softw 21:1–20. https://doi.org/10.18637/jss.v021.i12
Wickham H (2011) The split-apply-combine strategy for data analysis. J Stat Softw 40:1–29. https://doi.org/10.18637/jss.v040.i01
Wickham H, Vaughan M (2023) tidyr: Tidy Messy data. https://tidyr.tidyverse.org/. Accessed 31 Jan 2023
Wragg D, Marti-Marimon M, Basso B, Bidanel J-P, Labarthe E, Bouchez O, Le Conte Y, Vignal A (2016) Whole-genome resequencing of honeybee drones to detect genomic selection in a population managed for royal jelly. Sci Rep 6(1):27168. https://doi.org/10.1038/srep27168
Wright WE, Shay JW (2000) Telomere dynamics in cancer progression and prevention: fundamental differences in human and mouse telomere biology. Nature Med. https://doi.org/10.1038/78592
Yin L, Zhang H, Tang Z, Xu J, Yin D, Zhang Z, Yuan X, Zhu M, Zhao S, Li X, Liu X (2021) rMVP: a memory-efficient, visualization-enhanced, and parallel-accelerated tool for genome-wide association study. Genom Proteom Bioinform 19(4):619–628. https://doi.org/10.1016/j.gpb.2020.10.007
Yousefzadeh M, Henpita C, Vyas R, Soto-Palma C, Robbins P, Niedernhofer L (2021) DNA damage how and why we age? Elife 10:62852. https://doi.org/10.7554/eLife.62852
Zhang P, Edgar BA (2022) Insect gut regeneration. Cold Spring Harb Perspect Biol 14(2):a040915. https://doi.org/10.1101/cshperspect.a040915
Acknowledgements
We would like to acknowledge Dr. Serge Aron for his expertise in flow cytometry techniques in Hymenoptera.
Funding
Funding for this project came in part from Purdue AgSEED, Start-up funds, and Project Apis m awarded to BAH.
Author information
Authors and Affiliations
Contributions
BAH and RRS developed hypotheses and the study’s goals. KR, AC, and RRS developed the technique for individual honey bee tissue flow cytometry. RRS and KR performed flow cytometry analysis. RRS performed dissections and DNA extractions. BAH and RRS developed the pipeline for the analysis of tissue-specific somatic variants. RRS developed the analysis pipeline. JPR and BAH provided experimental guidance and tools. All authors have read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no competing interests.
Ethical approval
Not applicable.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Supplementary Information
Below is the link to the electronic supplementary material.
40_2024_948_MOESM1_ESM.pdf
Supplementary file1 S1: Accumulation of functional mutations across tissues. Light blue boxes indicate significant variance (P < 0.00417) in mutation accumulation between tissue couplings. (PDF 16 KB)
40_2024_948_MOESM2_ESM.pdf
Supplementary file2 S2: Accumulation of mutations across genomic regions and tissues. Light blue boxes indicate significant variance (P < 0.00208) in mutation accumulation between tissue couplings. (PDF 27 KB)
40_2024_948_MOESM3_ESM.pdf
Supplementary file3 S3: Somatic mutation density plot by tissue type. A plot of the density of mutations across each chromosome in 1 Mb windows. There was no significant difference in mutation density among tissues (adj. P > 0.05). (PDF 1923 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Shultz, R.R., Carey, A., Ragheb, K.E. et al. On the distribution and diversity of tissue-specific somatic mutations in honey bee (Apis mellifera) drones. Insect. Soc. 71, 97–108 (2024). https://doi.org/10.1007/s00040-024-00948-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00040-024-00948-5