Abstract
Parkinson’s disease (PD) is a common and incurable neurodegenerative disorder that arises from the loss of dopaminergic neurons in the substantia nigra and is mainly characterized by progressive loss of motor function. Monogenic familial PD is associated with highly penetrant variants in specific genes, notably the PRKN gene, where homozygous or compound heterozygous loss-of-function variants predominate. PRKN encodes Parkin, an E3 ubiquitin-protein ligase important for protein ubiquitination and mitophagy of damaged mitochondria. Accordingly, Parkin plays a central role in mitochondrial quality control but is itself also subject to a strict protein quality control system that rapidly eliminates certain disease-linked Parkin variants. Here, we summarize the cellular and molecular functions of Parkin, highlighting the various mechanisms by which PRKN gene variants result in loss-of-function. We emphasize the importance of high-throughput assays and computational tools for the clinical classification of PRKN gene variants and how detailed insights into the pathogenic mechanisms of PRKN gene variants may impact the development of personalized therapeutics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Parkinson’s disease (PD) is the most common movement disorder and the second most common neurodegenerative disorder after Alzheimer’s disease [1,2], affecting 2–3% of the population aged ≥ 65 years [1]. The etiology of PD is still not clear, although research suggests that the cause is multifactorial and linked to environmental agents, gene variants and aging [1].
Pathologically, PD is the result of the progressive loss of ventral mesencephalic dopaminergic neurons, which causes striatal dopamine deficiency and impairment of motor control [3]. Accordingly, there is an estimated 30% loss of midbrain dopaminergic neurons at the onset of motor symptoms [4]. The primary motor symptoms include bradykinesia, rigidity, tremor and at later stages postural instability [3]. In addition, PD patients also display a broad spectrum of non-motor features that typically develop gradually for years before any motor symptoms appear, including autonomic dysfunction (e.g. constipation), loss of the sense of smell and sleep impairment [5].
Currently, efficient therapies are available, making PD the first neurodegenerative disorder to be successfully managed and increasing the quality of life for patients for many years after disease onset [1]. However, the treatments are not curative and as a result the disease will slowly progress and eventually cause severe disabilities. The main therapeutic approaches include dopamine-related pharmacological treatment for PD motor symptoms, the most common being Levodopa (l-DOPA) [1,5]. However, l-DOPA treatment is complicated by l-DOPA-induced dyskinesia, motor fluctuations, and l-DOPA-resistant motor function [1]. For all PD patients the effect of medication wears off over time, hence only alleviating the symptoms for a limited number of years after diagnosis.
Often, the affected neurons in PD accumulate misfolded and aggregated α-synuclein into intracellular inclusions termed Lewy bodies, which appear to contribute to PD pathogenesis. Thus, both degeneration of dopaminergic neurons and the presence of Lewy bodies, constitute the hallmarks of PD [5]. However, since Lewy bodies are also present in other diseases such as dementia with Lewy bodies (DLB) [6] and are rarely seen in certain familial subtypes of PD [7], their formation does not solely account for the neuronal cell loss observed in PD. Another important mechanism contributing to PD, as well as other neurodegenerative diseases such as Huntington’s disease and amyotrophic lateral sclerosis, is mitochondrial dysfunction [8,9]. The initial evidence of altered mitochondrial function associated with PD emerged when mitochondrial toxins were observed to induce acute PD-like symptoms [10]. Later, this link was supported by the identification of genes associated with monogenic forms of PD. Here, the associated genes were found to encode proteins involved in mitochondrial quality control and the degradation of damaged mitochondria [11], supporting mitochondrial dysfunction to be implicated in the pathology of PD and a likely mechanism contributing to the neuronal cell loss [12].
Although a large majority of diagnosed PD cases are idiopathic, autosomal dominant and recessive familial forms have been identified [13]. Common genetic variability at more than 90 loci has been linked to PD, accounting for 16–36% of the heritable component of the disease. However, individually, each of these loci only has a small effect size. Conversely, rare, but highly penetrant, genetic alterations in SNCA (PARK1/4), LRRK2 (PARK8), VPS35 (PARK17), DJ-1 (PARK7), PINK1 (PARK6) and PRKN (PARK2) have been linked to monogenic familial PD [13,14]. Obviously, the identification of disease-linked germline variants in these genes is paramount for accurate diagnosis and genetic counselling of affected individuals and their families. Moreover, characterizing the mechanisms by which disease-linked variants function will enhance our understanding of PD in general and potentially contribute to the development of future therapeutic strategies. In this review, we start by providing a brief introduction to PRKN-linked familial PD and the relevant molecular and cellular pathways, noting that these topics have been explored in several excellent in-depth and historical reviews [12,15,16,17,18,19,20]. Subsequently, we delve into the molecular mechanisms underlying PRKN loss of function and recent advances in computational prediction tools and high-throughput technologies for clinical assessment of PRKN gene variants.
PRKN-linked Parkinson’s disease
PRKN (also known as PARK2) is a well-characterized PD-linked gene, encoding the E3 ubiquitin-protein ligase Parkin [21,22] involved in mitochondrial quality control and the degradation of damaged mitochondria through mitophagy. PRKN gene variants associated with PD lead to a loss of Parkin function and range from single base pair substitutions to small deletions and splice site aberrations, to deletions that span thousands of nucleotides [23]. Obviously, gene variants with altered splicing or deletions of coding regions are expected to disrupt gene function completely or at least very strongly, and similarly strong effects are expected for single nucleotide substitutions resulting in early stop codons (nonsense variants). Conversely, the effects of missense variants, where one amino acid residue is exchanged with another, may range from a complete loss of function to neutral (harmless), and intriguingly even to enhanced activity [24,25].
PRKN variants are the most common cause of autosomal recessive PD (ARPD) regardless of gender and with an age of onset before 40 years of age in most individuals [21,26,27]. In addition, PRKN variants are the most common cause of early-onset PD (age of onset ≤ 20 years) [26] accounting for 42% of the familial early-onset PD cases [27] and the majority of PRKN-related PD patients with disease onset before 20 years of age carry bi-allelic PRKN variants [28]. In terms of the cardinal motor symptoms, PRKN-related PD patients demonstrate a clinical phenotype resembling that of sporadic PD patients [29]. The PRKN-specific clinical features include early disease onset, hyperreflexia and dystonia as some of the first signs of the disease besides slow disease progression [29]. Additionally, dementia is rarely reported [30]. Clinical pathology shows a selective loss of dopaminergic neurons in the substantia nigra and loss of noradrenergic neurons in the locus coeruleus with accompanying gliosis [31] without the presence of Lewy bodies, except in rare cases [7]. In most cases, PRKN-related PD patients respond well to l-DOPA treatment even at late stages, and L-DOPA is therefore often considered an efficient treatment for a sustained period of time. PRKN-linked PD patients in treatment with L-DOPA are, however, prone to develop l-DOPA-induced dyskinesia [26,30].
Induced pluripotent stem cells (iPSCs), derived from somatic cells or tissues, have enabled the investigation of the progressive development of disease-associated cellular changes in dopaminergic neurons from PD patients. Concurrently, advances in genome editing techniques have allowed correction or introduction of disease-causing variants in PD-relevant genes of importance for mitochondrial function. This in turn enables examination of isolated effects of PD-relevant protein variants, including Parkin, in developing and mature human dopaminergic neurons. Studies of PD patient iPSC-derived neurons with PRKN variants have documented increased oxidative stress, abnormal mitochondrial/lysosomal morphology and function, dysregulation of dopamine homeostasis, and to some extent α-synuclein accumulation [32,33,34,35,36,37,38].
Parkin structure and function
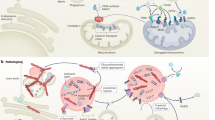
Parkin is a soluble multi-domain protein comprised of 465 amino acid residues [22], which make up an N-terminal ubiquitin-like (UBL) domain and four RING-like domains; RING0, RING1, in-between RING (IBR) and RING2 (Fig. 1). This arrangement of RING-IBR-RING (RBR) domains categorizes Parkin within the RBR E3 ubiquitin ligase protein family. Each RING domain co-ordinates two zinc ions that are critical for maintaining the protein structure [39]. In addition, Parkin contains a conserved nine-residue motif named the activating element (ACT) (residue 101–109) in the linker region between the UBL and RING0 domains [40], as well as a so-called repressor element (REP) located just upstream of the RING2 domain [41] (Fig. 1). These Parkin domains and motifs are highly conserved across species and play key roles for activity and regulation [40].
Parkin domain organization and auto-inhibited conformation. Schematic representation of the Parkin domain organization (upper panel). The AlphaFold2 (AF-O60260-F1) structure of Parkin is depicted, with colors corresponding to the domain organization (lower-left). Simplified representation of the Parkin structure in its auto-inhibited conformation (lower-right). In this conformation, both the active site (red dot within RING2) and the E2 binding site (yellow region in RING1) are occluded, thus preventing Parkin activity. Figure created with BioRender.com
As an E3 ubiquitin ligase, Parkin’s enzymatic activity is to catalyze covalent conjugation of the small protein ubiquitin to various cellular proteins. In this process, termed ubiquitination [42,43,44,45], a ubiquitin-loaded E2 enzyme will, in collaboration with an E3 enzyme, catalyze conjugation of the ubiquitin to the target protein. Typically this conjugation occurs via an isopeptide bond between the C-terminal carboxyl group of ubiquitin and the amino group of a lysine side chain in the target protein. Some E3 families catalyze the transfer of the ubiquitin moiety directly from the E2 to the target protein, whereas for others, including RBR-type E3s, such as Parkin, ubiquitin is first bound via a thioester bond to a cysteine residue in the E3, and subsequently transferred to the target protein [46,47,48]. In the case of Parkin, the active site cysteine residue is C431 located in the RING2 domain, while E2 binding occurs to the RING1 domain. Additional rounds of ubiquitination will target one or several of the lysine residues (K6, K11, K27, K29, K33, K48 and K63) in the conjugated ubiquitin, and lead to the formation of ubiquitin chains. Both cellular and in vitro studies have shown that Parkin catalyzes the formation of K6-, K11-, K48- and K63-linked ubiquitin chains [18]. Protein targets of Parkin catalyzed ubiquitination include several outer mitochondrial membrane (OMM) proteins including mitofusins MFN1 and MFN2 [49,50,51] but also a range of cytosolic proteins. This ubiquitination marks them for proteasomal degradation and autophagy, eventually leading to the clearance of damaged mitochondria (see below).
Considering these important cellular functions of Parkin, it is not surprising that its activity is tightly regulated and requires phosphorylation and structural rearrangements to become active [18,20,52]. Under basal conditions, Parkin adopts a closed, auto-inhibited conformation facilitated by its UBL domain associated with RING1. This auto-inhibitory state is governed by two key interactions: the REP element and UBL domain occluding the E2 binding site in RING1 and the RING0 domain blocking the active site (C431) in RING2 [53,54] (Fig. 1).
The structure of the individual domains and full-length Parkin have been solved [40,41,55,56,57,58,59,60,61,62], and the crystal structure of Parkin in its autoinhibited state is highly similar to the structure predicted by AlphaFold. Upon Parkin activation, auto-inhibited Parkin binds to ubiquitin phosphorylated by PTEN-induced kinase PINK1 (encoded by PINK1, the second most commonly ARPD-linked gene) at position S65 and translocates to damaged mitochondria. Parkin binding to phosphorylated ubiquitin retains it in an auto-inhibited conformation [55,58] but induces structural rearrangements of the UBL domain. In turn, this structural rearrangement allows for efficient Parkin phosphorylation by PINK1 at S65 in the UBL domain (the equivalent position in the UBL domain to S65 in ubiquitin) [41,55,57,58,59,63,64,65,66]. Next, binding of the phosphorylated UBL domain to a phospho-binding pocket in RING0 and the binding of the ACT element (previously bound by RING2 and REP) to RING0 cause a major structural reorganization that releases RING2 and the catalytic C431 residue from its inhibited state. This transforms Parkin into the fully activated Parkin conformation and promotes the binding of a ubiquitin-loaded E2 [24,40,54,57,60]. Subsequently, activated Parkin will ubiquitinate numerous OMM proteins [49,50,51] (Fig. 2).
Parkin activation. Parkin in its auto-inhibited conformation has the active site (red dot within RING2) and the E2 binding site (yellow region within RING1) occluded. PINK1 phosphorylation (red) of ubiquitin (UB) and the subsequent binding of phosphorylated ubiquitin result in the release of the UBL domain, priming it for phosphorylation by PINK1. Once phosphorylated, the UBL and ACT elements bind to RING0, leading to the release of REP and RING2. This release allows for the binding of E2 ~ UB, facilitating the transfer of ubiquitin from E2 to the catalytic C431 residue (red), and finally, the transfer of ubiquitin onto the substrate. Figure created with BioRender.com
Parkin regulates mitochondrial dynamics
Maintaining cellular homeostasis in response to stress conditions is crucial for cell function and survival. Mitochondria are double-membrane enclosed compartments that constantly adapt their shape and function to support the cell’s demands [67,68]. Since mitochondria generate high levels of reactive oxygen species as a natural by-product to their energy production, this makes them highly susceptible to mtDNA mutations and protein misfolding. Consequently, efficient and accurate quality control mechanisms are crucial for maintaining a healthy mitochondrial network and homeostasis [69]. This is of particular importance for non-dividing cells, like neurons, that are unable to dilute faulty mitochondria through cell division. These mechanisms include activation of the mitochondrial unfolded protein response (mtUPR) and integrated stress response (mtISR) pathways, as well as retro-translocation of misfolded mitochondrial proteins followed by degradation via the ubiquitin–proteasome system (UPS) (reviewed in [70,71,72]). These mechanisms provide an early first response system to alleviate the mitochondrial stress situation. However, as an additional layer of defense, cells also tightly regulate the biogenesis and degradation of mitochondria. Since mitochondria cannot be synthesized de novo, damaged or non-functional mitochondria are replaced by pre-existing ones, involving both fusion and fission processes. The balance between fusion and fission events affects the number, size and morphology of individual mitochondria, including small spherical shapes, long or short tube-like structures, or connected tubules [73,74]. Occurring on a regular basis, the two opposing processes, fusion and fission, comprise the mitochondrial dynamics.
Mitochondrial membrane fusion is orchestrated by three proteins belonging to the dynamin superfamily. The outer mitochondrial membrane (OMM) mitofusins, MFN1 and MFN2, required for fusion of the OMM, while the inner mitochondrial membrane (IMM) protein optic atrophy-1, OPA1, is responsible for fusion of the IMM. Fission occurs in response to cell division and stress conditions and requires the translocation of the dynamin-related protein 1, DRP1, to the OMM, where it divides the mitochondria into two [9,75]. Two distinct fusion events have been described: complete fusion and partial fusion [76]. Partial fusion and fission events ensure a continuous exchange of content between mitochondria, ensuring a homogenous mitochondrial population within the cell [9]. Although the function in mitochondrial quality control needs further exploration, based on the current understanding, fusion and fission events are assumed to aid the separation of damaged mitochondrial components from the functional mitochondrial network [74]. A recent study has demonstrated how the mitochondrion may display different membrane potentials along the mitochondrion [77,78], supporting the possibility of a mitochondrion having a high membrane potential in one part and a low membrane potential in another. This could explain the consistent observation of uneven fission events [79]. Here, daughter units displaying a high membrane potential were likely to reintegrate into the mitochondrial network by fusion, whereas daughter units exhibiting low membrane potentials were isolated from the network and subsequent likely to be eliminated by a selective form of autophagy, termed mitophagy.
Mitophagy is the process of autophagy-mediated selective degradation of abnormal or dysfunctional mitochondria in lysosomes, is a critical pathway for sustaining a network of healthy mitochondria. Parkin and PINK1 play pivotal roles in preventing fusion and promoting the segregation of damaged mitochondria for degradation through mitophagy [51,80,81,82,83]. To this end, Parkin mediates the ubiquitination and subsequent proteasomal degradation of multiple OMM proteins, such as MIRO, MFN1 and MFN2, important for mitochondrial fusion. At the same time, PINK1 induces mitochondrial fission by recruiting DRP1 to the OMM [84,85,86] (Fig. 3).
The role of Parkin in mitochondrial fission. Mitochondrial fission and subsequent degradation of damaged mitochondria (orange mitochondria) by Parkin/PINK1 dependent attachment of ubiquitin (purple) and consequent proteasomal degradation of mitofusins (MFN1/2) and MIRO, and PINK1 mediated recruitment of DRP1. Figure created with BioRender.com
Overall, Parkin can preserve mitochondrial integrity by regulating mitochondrial dynamics by mediating mitochondrial fission [51,80,81,82,83,87]. Mitochondrial fission is widely recognized as the initial step necessary for removal of damaged mitochondria via mitophagy [84,88].
PINK1 and Parkin-mediated mitophagy
Work in Drosophila provided the first links between PINK1, Parkin and mitophagy [89,90,91], showing that parkin and pink1 null mutants display similar phenotypes, including locomotor defects and male sterility, and linking these phenotypes to defects in mitochondrial morphology [89,90]. Later proteomics studies revealed that mitochondrial protein turnover was slowed in parkin mutants [72], thus showing that Parkin promotes mitophagy in vivo.
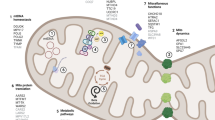
Accordingly, Parkin and PINK1 not only promote the segregation of damaged mitochondria, but also maintain mitochondrial homeostasis by mediating the degradation of damaged mitochondria [92,93]. When mitochondria are healthy, PINK1 is targeted to the mitochondria through its mitochondrial targeting sequence (MTS) [92]. Subsequently, it translocates through the translocase of the outer membrane (TOM) complex into the translocase of the inner membrane (TIM) complex [94]. Here, PINK1 is cleaved by the mitochondrial processing protease (MPP) [95], which catalyzes the cleavage of the N-terminal MTS of imported precursor proteins [96]. Following this, PINK1 is rapidly cleaved in its N-terminal transmembrane domain by the mitochondrial intramembrane PARL protease [97,98] and is retrotranslocated from the mitochondria to the cytosol for degradation through the ubiquitin–proteasome system (UPS) via the so-called N-end rule ubiquitin-conjugation pathway [99]. This PINK1 degradation mechanism ensures its low abundance in cells containing healthy mitochondria.
During conditions where mitochondria become damaged or dysfunctional, it is critical that these organelles are cleared from the cell. Defects in mitochondrial protein import may occur in both depolarized mitochondria (e.g., due to mitochondrial damage) [100] and polarized mitochondria (e.g., defective due to perturbed mitochondrial proteostasis) [88,101,102,103]. Impairment of protein import into the mitochondria prevents the translocation of PINK1, leading to PINK1 accumulation on the OMM in association with the TOM complex [104], thus flagging these mitochondria as damaged. Then, PINK1 on the OMM is activated through dimerization and trans-auto-phosphorylation [105,106,107]. Recent structural studies of PINK1 in an activated state have provided detailed insights into the order of events during PINK1 activation [107,108].
A recent study indicates an early first step in Parkin-PINK1 dependent mitophagy, prior to PINK1 activation of Parkin, involves the AMP-activated protein kinase AMPK that regulates autophagy and mitophagy through the serine/threonine protein kinase, ULK1 [109]. Here, phosphorylation of the highly conserved S108 in the Parkin ACT element by ULK1 appears to precede PINK1 activation of Parkin in response to mitochondrial damage [110]. Subsequently, PINK1 triggers Parkin activation a multi-step process. First, PINK1 phosphorylates S65 on pre-existing ubiquitin on the OMM [49,111,112,113,114]. The mitochondrial E3 ubiquitin ligase MITOL has been proposed to introduce the initial ubiquitin important for Parkin recruitment [115]. During Parkin activation, auto-inhibited Parkin binds to the phosphorylated ubiquitin, thus translocating to damaged mitochondria, where the mentioned structural rearrangements activate Parkin. PINK1 phosphorylates both monomeric ubiquitin and poly-ubiquitin chains that Parkin binds to, hence retaining Parkin on the OMM. This binding of Parkin to the OMM boosts the formation of additional mono- and poly-ubiquitinated substrate proteins, initiating a positive feedback loop in which PINK1 phosphorylates additional Parkin and ubiquitin molecules to activate and retain additional Parkin on the surface of dysfunctional mitochondria [49,116]. Furthermore, recent studies indicate Parkin activation independent of Parkin phosphorylation by PINK1, supporting the feed-forward mechanism and rapid degradation of mitochondria through the Parkin-PINK1 mitophagy pathway. In this scenario, the phosphor-binding pocket in the RING0 domain of Parkin binds to phosphorylated ubiquitin on the OMM, releasing the catalytic RING2 from the auto-inhibited position [61,62]. These findings may explain how Parkin, without its UBL domain, retains some ability to induce mitophagy [24,53,66].
The accumulation of ubiquitinated OMM proteins marks the mitochondria for degradation through both mitophagy and the UPS [18,51,83,100,117,118]. The initiation of mitophagy likely involves the prior degradation of some ubiquitinated OMM proteins through the UPS [51,83]. Subsequently, the ubiquitin chains present on the OMM recruit autophagic cargo receptors such as sequestosome 1 (SQSTM1/p62) and optineurin (OPTN). The dysfunctional mitochondria are then encapsulated by a double-membrane vesicle known as the autophagosome and delivered to lysosomes, where the lysosomal hydrolases finally break down the mitochondria [18] (Fig. 4).
Parkin-dependent mitophagy. When mitochondria are healthy PINK1 is translocated through TOM and cleaved by the mitochondrial intramembrane protease PARL (blue pac-man) and retrotranslocated from the mitochondria to the cytosol where it is degraded. Upon mitochondrial damage, PINK1 is stabilized at the outer mitochondrial membrane (OMM), where it phosphorylates (red) OMM protein such as ubiquitin (purple), causing Parkin recruitment to damaged mitochondria. Then, Parkin activation is induced by PINK1 phosphorylation and a feedforward mechanism involving phosphorylated ubiquitin. Activated Parkin then mediates the formation of ubiquitin chains on OMM proteins that directs certain OMM proteins for proteasomal degradation and some to be recognized by autophagy receptors, which leads to the sequestering of damaged mitochondria in auto-phagosomes that fuse with lysosomes. Figure created with BioRender.com
Negative regulation of Parkin-mediated mitophagy
Considering the tight regulation of Parkin activation, it is not surprising that Parkin-mediated mitophagy is also subject to negative regulation at several stages along the PINK1-Parkin signaling axis. Accordingly, the PTEN-L isoform of the PTEN phosphatase was shown to dephosphorylate ubiquitin and Parkin [119,120], thus forcing Parkin towards the auto-inhibited conformation, blocking its mitochondrial translocation, and suppressing mitophagy.
Similar to phosphorylation, ubiquitination is a reversible post-translation modification. Deubiquitinating enzymes (DUBs), also known as ubiquitin-specific proteases (USPs), mediate this reversal by cleaving isopeptide bonds between individual ubiquitin moieties or between ubiquitin and the protein target [121,122]. Accordingly, DUBs capable of trimming the ubiquitin or phospho-ubiquitin chains on OMM proteins antagonize mitophagy, and are thus obvious drug targets for PD [123,124,125,126]. Intriguingly, USP30, a K6-ubiquitin-chain-specific DUB localized on the OMM, has been shown to antagonize Parkin/PINK1-dependent mitophagy by deubiquitinating OMM proteins, including TOM20 [127,128,129]. Moreover, several other DUBs [130], including ataxin-3 [131,132], USP8 [133], and USP15 [134,135], have been reported to deubiquitinate mitochondrial proteins, and some redundancy between these enzymes is expected.
Other functions of Parkin
In addition to its role in mitochondrial quality control and PD, Parkin has been implicated in various other functions, too extensive to be discussed exhaustively here. However, notably a correlation between impaired Parkin activity and increased risk of cancer has been observed. In turn, this indicates that Parkin activity and efficient mitochondrial quality control provide a mechanism for tumor suppression [136,137]. As with this observation, most of the additional functions attributed to Parkin are still, albeit more indirectly, connected with mitochondrial homeostasis and dependent on PINK1. For instance, PINK1 and Parkin regulate PGC1α activation through degradation of the transcriptional repressor ZNF746 (also known as PARIS). PINK1 directly phosphorylates PARIS, priming it for ubiquitination by Parkin, which interacts with the C-terminal zinc finger domain of PARIS and tags it for degradation [138]. Since PGC1α stimulates the synthesis of mitochondrial DNA, protein, and membrane [139], Parkin may also in this way affect mitochondrial homeostasis. Accordingly, overexpression of PARIS leads to the loss of dopaminergic neurons, an effect that can be reversed by co-expression of either Parkin or PGC1α [138]. Studies have also shown that Parkin plays a role in ubiquitin-mediated autophagy of intracellular Mycobacterium tuberculosis [140], which may explain why certain non-coding PRKN polymorphisms have been linked to increased susceptibility to intracellular pathogens [141,142].
Finally, a link between immune signaling by the kinase TBK1 and Parkin-dependent mitophagy was recently shown to occur via OPTN [143]. Thus, after Parkin-dependent mitochondrial ubiquitination, OPTN assembly stimulates TBK1 which then further stimulates OPTN assembly [143]. This positive feedback loop eventually results in the elimination of damaged mitochondria and also connects Parkin and neurodegeneration with inflammatory processes.
The fragility of the PRKN gene
The PRKN gene is located in the so-called FRA6E region on chromosome 6, one of the most fragile areas of the human genome [144]. Typically, chromosomal fragile sites are hotspots for deletions and amplifications, and fragility of the FRA6E region is likely caused by replication problems linked to transcription of the extremely large PRKN gene [144,145,146]. Thus, even though the mature PRKN mRNA is only about 4 kb, the PRKN gene and primary transcript is a whopping 1.4 Mb which mainly consists of introns. Since, the size of the region is conserved in vertebrates, the non-coding DNA may possess some unknown regulatory function [144,147]. A recent study has shown that the large introns do not seem to be important for PRKN expression 146, but may through the fragility of the region contribute to an increased risk of de novo germline variants in PRKN. Moreover, the fragility of the PRKN gene likely increases the risk of somatic PRKN mutations, which based on Parkin’s possible role as a tumor suppressor [136,137], could have implications for cancer susceptibility.
PD-associated PRKN variants
To aid research and diagnosis of genetic disorders, information on gene variants and their significance to disease have been collected in databases such as ClinVar [23,148]. For certain diseases, specialized databases have been developed that focus on a single disease or a subset of diseases. In case of PRKN gene variants, these are recorded in both ClinVar and the Movement Disorder Society Genetic mutation database (MDSGene) [149]. Currently, ClinVar comprises information on more than 650 different PRKN variants reported and interpreted by clinical and research laboratories as either “benign”, “likely benign”, “likely pathogenic”, “pathogenic”, “conflicting interpretations” or “variant of uncertain significance (VUS)”. In addition to the clinical status of the reported PRKN variants, the databases also include information on the type of variant. Synonymous variants were once perceived as “silent” with no impact on protein abundance, structure and activity, but some synonymous variants have later been demonstrated to partake in various human diseases by the deletion or creation of splice sites, or by affecting RNA stability, miRNA targeting or the rate of translation, thus altering co-translational protein folding [150]. However, in general synonymous variants only rarely affect function, and in the case of PRKN none of the reported synonymous variants are classified as disease-linked in ClinVar or MDSGene [151]. From a total of 1350 PRKN sequence variants reported in MDSGene, most (43.5%) of the disease-linked variants are so-called structural variants causing exon rearrangements. These are followed by missense (22.3%), frameshift (16.5%), nonsense (7.9%), and splice-site variants (7.9%) [151]. Structural and nonsense variants are typically highly detrimental and are therefore generally predicted to cause loss of function. In contrast, the effect of missense variants can range from subtle to dramatic impact on protein structure, stability and activity, dependent on the position and nature of the amino acid substitution. Accordingly, predicting the impact of missense variants on protein stability and function is not straightforward, and missense PRKN variants therefore represent the largest group that is clinically classified as conflicting or as variants of unknown significance (VUS).
Inactivating mechanisms of disease-linked Parkin missense variants
Missense variants can inhibit gene function in multiple ways. However, two common mechanisms, which are not mutually exclusive, include either a direct ablation of protein activity e.g. by disruption of the active site, or indirectly by destabilizing the native fold (Fig. 5) [152]. In some cases, the position and nature of the substitution may provide insights into its mechanism of pathogenicity. Thus, Parkin missense variants that directly inhibit the catalytic activity are expected to occur at (or near) the catalytic C431 residue [54,153,154,155]. The pathogenic variants: T415N, G430D and P437L are all positioned close to the active site and were found to be expressed at high levels but displayed very low mitophagy upon mitochondrial damage [25], indicating that they may alter the catalytic site causing impaired functionality without strongly affecting Parkin folding and structural stability. The Parkin M1T and M1V variants, affecting the start codon, will disturb translation initiation. The consequence of using an alternative in-frame start codon found at position 80 would result in a version of Parkin lacking its UBL domain [156], which has been observed in human cell lines [155]. Despite the mentioned report that Parkin lacking its UBL domain is functional, the M1T and M1V variants are pathogenic. Another variant, K161N, found in a family with PD [157], affects a position involved in binding the phosphorylated UBL to the RING0 domain and abolishes Parkin’s ability to mediate mitophagy [25,158], emphasizing the importance of this residue and interaction for Parkin function.
Disease-linked missense variants located outside the active site or other critical functional positions are likely to affect protein folding or stability, thus more indirectly inhibiting function. Typically, structurally destabilized proteins are prone to interact with components of the cellular protein quality control (PQC) system, including molecular chaperones and components of the UPS, leading to rapid PQC-linked degradation and reduced abundance of the missense protein. For instance, PRKN variants affecting the zinc coordinating residues, such as the PD-associated variants, C212Y, C253Y and C441R, are of low abundance and display impaired capability to induce mitophagy in response to mitochondrial stress [25]. In addition, the well-described pathogenic variant R42P [21,159] introduces a steric clash in the UBL domain, causing global unfolding of the UBL domain and drastic loss of protein abundance due to its rapid proteasomal degradation [25,155,160,161]. However, as the active site in RING2 is intact, the R42P variant may still be enzymatically active. Indeed, Parkin R42P has been shown to retain function [25,53,158,162,163]. Accordingly, using the Muller classification of mutations [164], R42P is an example of a hypomorph variant (a variant with reduced activity), while an active site variant would display a complete loss of function and thus be categorized as amorphic. Obviously, also stable variants may retain some enzymatic function and far from all destabilized variants are expected to be hypomorphs, since some variants may affect both activity and structural stability of the protein, which in case of Parkin will likely include substitutions of residues coordinating the Zn2+ ions in RING2.
Potential for restoring Parkin variant abundance and activity
The distinction between hypomorphic and amorphic missense variants is important, since hypomorphic alleles can, in principle, be rescued, whereas amorphic variants, similar to a deletion, cannot. Thus, increasing the abundance of a hypomorphic Parkin variant above some critical threshold could result in reactivation of mitophagy and thus mitigate disease. Potentially, this could be accomplished in multiple ways, such as through small molecule stabilizers, or by boosting synthesis or blocking the PQC-linked degradation. Though this has yet to be attempted for Parkin, based on other genetic disorders, these strategies hold some promise. For instance, small molecule stabilizers of the CFTR protein have proven effective for cystic fibrosis [165,166,167], and in the case of the hereditary cancer predisposition disease known as Lynch syndrome, blocking PQC-linked degradation restores function and drug sensitivity in cell models [168,169].
Currently, it is unknown how many of the Parkin missense variants are hypomorphic. However, recent estimates based on computational studies and high-throughput experiments have shown that as much as 60% of disease-linked missense variants results in significantly reduced structural stability of the encoded protein [152,170], which in turn is expected to lead to PQC-mediated degradation and a reduced abundance of the protein [171,172]. Based on recent deep mutational scanning of the abundance of > 99% of all possible Parkin variants, 50% of the known disease-linked Parkin missense variants were found to result in dramatically reduced steady-state levels (Fig. 5) [163]. Among all possible Parkin missense variants, 40% displayed an abundance reduced by more than half the amount of wild-type Parkin [163], and at least some of these are likely to be hypomorphic. Indeed, since Parkin is a multi-domain protein, it is likely that many low abundance missense variants positioned outside the catalytic RING2 domain are hypomorphic similar to R42P. However, the modular nature of the Parkin structure also challenges the development of stabilizing small molecules, since likely such a molecule would only correct protein folding/stability locally in the domain to which it binds. Accordingly, studies on enhancing PRKN expression and blocking the degradation of destabilized Parkin variants are warranted.
Degradation of destabilized Parkin variants
Since evolution selects for function rather than stability, most proteins are not overly stable under physiological conditions [173,174,175], and are therefore susceptible to degradation under stress conditions. Accordingly, various stressors can cause a decrease in Parkin protein levels [176,177,178,179,180]. In addition, Parkin loss is also observed upon mitochondrial depolarization, through either Parkin-dependent mitophagy or auto-ubiquitination and proteasomal degradation [133,181,182,183,184]. A more recent study found that Parkin is targeted for proteasomal degradation in response to stress conditions, independently of auto-ubiquitination and mitophagy [176].
As protein folding and structural stability are determined by the amino acid sequence [185], missense variants may, similar to a stress situation, lead to a reduced thermodynamic stability of the protein structure. Accordingly, a larger fraction of such proteins will, relatively to the wild-type, be found in a partially or fully unfolded conformation, and are therefore prone to form non-specific interactions with other cell components, thus poisoning the intracellular environment. To mitigate this danger, cells are equipped with a protein quality control (PQC) system that catalyzes the refolding or degradation of these aberrant proteins [172,186]. Molecular chaperones promote protein folding [187,188], but also collaborate with ubiquitin-protein ligases and the proteasome to clear non-native proteins from the cytosol and nucleus [189,190,191,192,193].
It is well known, that certain pathogenic PRKN variants may lead to a structural destabilization of the protein, which in turn results in reduced solubility, aggregate formation, and increased degradation [155,194,195]. It is therefore not surprising that such Parkin variants are subject to regulation by the PQC system. For instance, Hsp70 overexpression has been observed to ameliorate phenotypes of Drosophila Parkin mutants [196]. In addition, increased expression of chaperones has been found to both prevent aggregation of wild-type Parkin and promote the folding of the W453stop nonsense variant [197], while the J-domain Hsp70 co-chaperone DNAJB2 has been found to restore function to the low-abundance C289G Parkin variant [194]. Conversely, BAG5, another Hsp70 co-chaperone, has been found to interact with Parkin, inhibit its ubiquitin-ligase activity and enhance Parkin localization in protein aggregates [198]. Accordingly, molecular chaperones have been suggested as targets for PD therapeutics [199,200].
In addition to PQC-mediated degradation of destabilized Parkin variants, certain Parkin missense variants in the N-terminal UBL domain such as R42P have been shown to disrupt the auto-inhibited conformation, leading to Parkin activation and auto-ubiquitination, followed by degradation [155,160,161,163]. Hence, unlike most PQC targets, structural destabilization of Parkin can trigger two separate mechanisms of Parkin ubiquitination (Fig. 6). However, importantly increased auto-ubiquitination does not preclude PQC-mediated ubiquitination, and the auto-ubiquitination-dependent clearance of Parkin variants will only apply to the subset of destabilized Parkin variants that are hypomorphs.
Ubiquitination and proteasomal degradation of destabilized Parkin variants. Parkin, which is structurally destabilized due to a missense variant (star), can be subject to both increased auto-ubiquitination and PQC-linked ubiquitination by other cellular E3s (orange). Figure created with BioRender.com
In PQC-dependent protein degradation, recent data suggest that the discriminating feature recognized by the degradation system is the exposure of degradation signals (degrons) through local or global unfolding events [171,172,201,202]. It has recently been shown that many such quality control degrons are enriched in hydrophobic residues and depleted for negatively charged residues [203,204,205,206,207,208]. Accordingly, these degrons are typically buried inside the native protein structure but become exposed upon unfolding or misfolding [201,209]. In a recent systematic study, we found several such PQC degrons are found within all of the structured domains in Parkin [163]. Presumably, upon introduction of a destabilizing amino acid substitution, local or global unfolding will result in exposure of some of these PQC degrons that in turn will recruit molecular chaperones and PQC E3s to facilitate ubiquitination and degradation (Fig. 7).
The role of degrons in the degradation of Parkin variants. Structurally destabilized Parkin variants may transiently expose buried PQC degrons leading to recognition by PQC components and degradation (left). Alternatively, surface exposed degrons may be generated as a direct result of a missense variant (yellow star). Such neo degrons also lead to degradation of the protein variant. However, in this case the protein is structurally stable and likely still functional (right). Figure created with BioRender.com
An alternative mechanism, by which amino acid substitutions may trigger degradation, is by generating degrons in exposed regions of the protein. In this situation, the amino acid substitution need not destabilize the native fold, since the degron is surface exposed and thus can be directly recognized by the E3. Although none of the known disease-linked Parkin variants have been shown to operate in this manner, specific missense variants in the disordered linker region between the UBL and RING0 domains can form such a neo-degron [163] (Fig. 7). As the region is unstructured, the prediction is that such degron forming variants will not affect the structural stability or function of the protein, and degron generating variants should therefore be hypomorphic. However, as the region is conserved through evolution and proximal to the ACT element involved in Parkin activation [40], it is possible that the degron formation via amino acid substitutions at this position is an indirect consequence of the sequence properties required for function of the ACT region. Thus, the functional requirement for an exposed, partially hydrophobic motif makes the ACT region prone to missense variants that introduce additional hydrophobic residues and the formation of a neo-degron.
Interestingly, in depth studies have shown that some Parkin missense variants, including V224A, W403A and F146A enhance Parkin-dependent mitophagy [24,25,210]. In addition, these were shown to rescue mitophagy when expressed in cis with several otherwise non-functional variants [25] and thus appear to function as intragenic second site suppressors of certain disease-linked Parkin variants, including variants in regions important for PINK1-mediated activation or E2 binding sites such as K211N and T240M, respectively, as well as low abundance variants such as R42P, V56E and R275W. Mechanistically, such suppressors could operate by increasing the specific enzyme activity, increasing phospho-ubiquitin binding, mitochondrial translocation, and/or by stabilizing the native conformation, thus increasing Parkin abundance. However, since several of the hyperactive variants actually appear to destabilize the closed inactive Parkin conformation [25,210], they likely operate by shifting Parkin towards the active conformation. In turn, this may lead to increased autoubiquitination and/or a general destabilization of the Parkin structure. This is supported by the observation that some of these variants display reduced steady-state levels [163], and therefore, paradoxically the hyperactive Parkin variants could also be hypomorphic. Hopefully, future studies will probe these observations further, which could potentially pave the way for developing therapeutics mimicking the second site suppressors.
Variants of uncertain significance
Like most severe monogenic disorders, PRKN-linked PD is rare. However, due to the increased speed and lowered cost of DNA sequencing, a growing number of individuals are being sequenced and diagnosed with PRKN-linked PD and other genetic diseases. In addition, the rise in genome sequencing also results in an increased number of newly identified gene variants, whose pathogenicity must be carefully determined. For instance, whole exome sequencing analyses return ~ 50,000 autosomal variants per individual compared to a reference genome [211]. Only a small fraction will cause monogenic disease while the majority are either harmless or may contribute to multifactorial disease. However, often there is insufficient evidence to classify newly observed variants as being either benign or pathogenic and instead these are designated as VUS [212]. In terms of diagnosis and genetic counseling of affected individuals or families, clinical sequencing is only relevant if variants are classified, which renders the large number of VUS highly problematic. To improve clinical diagnostic yield from genome sequencing there is a need to increase the speed of variant classification. As evident from the above, variant classification, in particular of missense variants, is not straightforward. While population-based studies are doing well at linking common variants with phenotypes, they require many individuals and are often inadequate in the case of rare variants [212]. Traditionally, the effect of a VUS is assessed in a low-throughput manner (Fig. 8), efforts that can take several months or years. Though such assays typically yield detailed and highly accurate results, this approach is not feasible considering the number of observed gene variants. Indeed, given the size of the human genome, the mutation rate, and the current global population, all single nucleotide variants compatible with life are likely to exist in the current human population [213]. For this reason, there is a need for assessing not only already observed variants, but essentially all variants [214], including the rare variants that have not yet been observed in the population.
Approaches to classification of variant effects. The effects of PRKN gene variants can be assessed by traditional and highly detailed, but often very costly low throughput approaches (upper panel). Computational studies offer an alternative approach, which is cheap and rapid, but also sometimes less precise (middle panel). Finally, high-throughput experimental approaches allow libraries of gene variants to be assayed in large, multiplexed experiments (lower panel). Figure created with BioRender.com
To approach this issue, both computational techniques and laboratory-based high-throughput assays have been developed to rapidly generate variant effect data. Computational models typically assess the impact of gene variants based on sequence conservation and/or on structural data, and models are trained to predict the variant effects on protein function. These in silico predictors provide a non-laborious and fast method to predict the impact of variants making them attractive (Fig. 8). With recent developments, precise variant effect predictions are possible [215,216], at least for the kinds of variants, genes and diseases for which we can accurately assess such pathogenicity predictions. However, in general computational tools do not inform on the mechanistic basis of the variant effects. The laboratory-based high-throughput assays, though demanding, offer an approach to variant classification [214] (Fig. 8), and generate data, which may also be useful for training future computer-based predictors. Our recent deep mutational scanning of Parkin variant abundance revealed that structural destabilization and degradation explain about half of the pathogenic Parkin variants [163], and is thus on its own, not highly useful for variant classification. Accordingly, additional comprehensive assessments of variant effects are required, such as high-content mapping of Parkin function e.g. by monitoring Parkin’s ability to initiate mitophagy.
Concluding remarks
Mitochondrial biogenesis, mitochondrial dynamics, and mitophagy form a continuous process to preserve optimal mitochondrial quality. Parkin, as a key protein, plays a pivotal role in the entire spectrum of mitochondrial quality control.
As evident from the above, studying the cellular and molecular functions of Parkin provides important insights into not only the monogenic forms of PD, but also indirectly increases our understanding of idiopathic PD. More specifically, studies on Parkin variant effects are useful for clinical genetics and variant classification, but also benefit our understanding of the structural, biophysical and cellular impact of gene variants, regardless of their frequency in the population [217].
Although variant classification based on high-throughput experimental or computational data is currently showing promising progression, these methods will still wrongly classify some variants and also do not provide highly detailed information on the molecular and cellular mechanisms involved. Meticulous low-throughput structural, biophysical, and animal studies are therefore still exceedingly important. This is exemplified by the studies showing second site suppressors of certain disease-linked Parkin variants [25,210], which may guide development of small molecules with similar effects. Recently, a positive allosteric modulator of Parkin activity, BIO-2007817, was described [218], but the compound did not increase mitophagy [125,218] and did not broadly stabilize low abundant Parkin variants in cell-based studies [163]. However, other small molecules that interact with Parkin could potentially activate Parkin or at least stabilize the protein and thus increase the intracellular amount of low abundance pathogenic variants [171,219]. Alternatively, increased cellular Parkin levels could also be achieved by boosting Parkin synthesis or blocking Parkin degradation, and for this reason further studies on Parkin proteostasis are required. For instance, although some chaperones and co-chaperones have been shown to triage Parkin variants [194,196,197,198], it is unknown if these engage broadly with structurally destabilized disease-linked variants. In addition, the identity of the E2 and E3 enzymes involved in the PQC-linked proteasomal degradation of structurally unstable Parkin variants are still unknown, despite their potential importance as drug targets for low-abundance variants. Other approaches to activate Parkin-mediated mitophagy are currently ongoing and include USP30 inhibitors [220] or activators of PINK1 or ULK1 (recently reviewed [125]). Importantly, such therapeutics may be relevant not only for PRKN-linked PD, but likely also for idiopathic PD.
Availability of data and material
Not applicable.
References
Poewe W et al (2017) Parkinson disease. Nat Rev Dis Primers 3:17013
Balestrino R, Schapira AHV (2020) Parkinson disease. Eur J Neurol 27:27–42
Vázquez-Vélez GE, Zoghbi HY (2021) Parkinson’s disease genetics and pathophysiology. Annu Rev Neurosci 44:87–108
Cheng HC, Ulane CM, Burke RE (2010) Clinical progression in Parkinson disease and the neurobiology of axons. Ann Neurol 67:715–725
Armstrong MJ, Okun MS (2020) Diagnosis and treatment of Parkinson disease: a review. JAMA 323:548–560
Goedert M (2001) Alpha-synuclein and neurodegenerative diseases. Nat Rev Neurosci 2:492–501
Savitt JM, Dawson VL, Dawson TM (2006) Diagnosis and treatment of Parkinson disease: molecules to medicine. J Clin Invest 116:1744–1754
East DA, Campanella M (2016) Mitophagy and the therapeutic clearance of damaged mitochondria for neuroprotection. Int J Biochem Cell Biol 79:382–387
Chan DC (2020) Mitochondrial dynamics and its involvement in disease. Annu Rev Pathol 15:235–259
William Langston J, Ballard P, Tetrud JW, Irwin I (1983) Chronic Parkinsonism in humans due to a product of meperidine-analog synthesis. Science 219:979–980
Park JS, Davis RL, Sue CM (2018) Mitochondrial Dysfunction in Parkinson’s Disease: New Mechanistic Insights and Therapeutic Perspectives. Curr Neurol Neurosci Rep 18:1–11
Pickrell AM, Youle RJ (2015) The roles of PINK1, parkin, and mitochondrial fidelity in Parkinson’s disease. Neuron 85:257–273
Bandres-Ciga S, Diez-Fairen M, Kim JJ, Singleton AB (2020) Genetics of Parkinson’s disease: an introspection of its journey towards precision medicine. Neurobiol Dis 137:104782
Nalls MA et al (2019) Identification of novel risk loci, causal insights, and heritable risk for Parkinson’s disease: a meta-analysis of genome-wide association studies. Lancet Neurol 18:1091–1102
Truban D, Hou X, Caulfield TR, Fiesel FC, Springer W (2017) PINK1, Parkin, and mitochondrial quality control: what can we learn about Parkinson’s disease pathobiology? J Parkinsons Dis 7:13–29
Bayne AN, Trempe JF (2019) Mechanisms of PINK1, ubiquitin and Parkin interactions in mitochondrial quality control and beyond. Cell Mol Life Sci 76:4589–4611
Trempe JF, Gehring K (2023) Structural mechanisms of mitochondrial quality control mediated by PINK1 and parkin. J Mol Biol 435:168090
Wade Harper J, Ordureau A, Heo JM (2018) Building and decoding ubiquitin chains for mitophagy. Nat Rev Mol Cell Biol 19:93–108
Matsuda N, Yamano K (2020) Two sides of a coin: Physiological significance and molecular mechanisms for damage-induced mitochondrial localization of PINK1 and Parkin. Neurosci Res 159:16–24
Tanaka K (2020) The PINK1-Parkin axis: an overview. Neurosci Res 159:9–15
Shimura H et al (2000) Familial Parkinson disease gene product, parkin, is a ubiquitin-protein ligase. Nat Genet 25:302–305
Kitada T et al (1998) Mutations in the parkin gene cause autosomal recessive juvenile Parkinsonism. Nature 392:605–608
Landrum MJ et al (2020) ClinVar: improvements to accessing data. Nucleic Acids Res 48:D835–D844
Tang MY et al (2017) Structure-guided mutagenesis reveals a hierarchical mechanism of Parkin activation. Nat Commun 8:1–14
Yi W et al (2019) The landscape of Parkin variants reveals pathogenic mechanisms and therapeutic targets in Parkinson’s disease. Hum Mol Genet 28:2811
Brüggemann N Klein, C. Parkin Type of Early-Onset Parkinson Disease. GeneReviews® (2020).
Lesage S et al (2020) Characterization of recessive Parkinson disease in a large multicenter study. Ann Neurol 88:843–850
Lücking CB et al (2000) Association between early-onset Parkinson’s disease and mutations in the parkin gene. N Engl J Med 342:1560–1567
Corti O, Lesage S, Brice A (2011) What genetics tells us about the causes and mechanisms of Parkinson’s disease. Physiol Rev 91:1161–1218
Wasner K, Grünewald A, Klein C (2020) Parkin-linked Parkinson’s disease: From clinical insights to pathogenic mechanisms and novel therapeutic approaches. Neurosci Res 159:34–39
Mori H et al (1998) Pathologic and biochemical studies of juvenile parkinsonism linked to chromosome 6q. Neurology 51:890–892
Bogetofte H et al (2019) PARK2 mutation causes metabolic disturbances and impaired survival of human iPSC-derived neurons. Front Cell Neurosci 13:297
Bogetofte H et al (2019) Perturbations in RhoA signalling cause altered migration and impaired neuritogenesis in human iPSC-derived neural cells with PARK2 mutation. Neurobiol Dis 132:104581
Okarmus J et al (2020) Lysosomal perturbations in human dopaminergic neurons derived from induced pluripotent stem cells with PARK2 mutation. Sci Rep 10:10278
Okarmus J et al (2021) Identification of bioactive metabolites in human iPSC-derived dopaminergic neurons with PARK2 mutation: altered mitochondrial and energy metabolism. Stem Cell Rep 16:1510–1526
Wei PC et al (2019) Neuroprotection of indole-derivative compound NC001–8 by the regulation of the NRF2 pathway in Parkinson’s disease cell models. Oxid Med Cell Longev 2019:5074367
Shaltouki A et al (2015) Mitochondrial alterations by PARKIN in dopaminergic neurons using PARK2 patient-specific and PARK2 knockout isogenic iPSC lines. Stem Cell Reports 4:847–859
Imaizumi Y et al (2012) Mitochondrial dysfunction associated with increased oxidative stress and α-synuclein accumulation in PARK2 iPSC-derived neurons and postmortem brain tissue. Mol Brain 5:35
Seirafi M, Kozlov G, Gehring K (2015) Parkin structure and function. FEBS J 282:2076–2088
Gladkova C, Maslen SL, Skehel JM, Komander D (2018) Mechanism of parkin activation by PINK1. Nature 559:410–414
Trempe JF et al (2013) Structure of parkin reveals mechanisms for ubiquitin ligase activation. Science 340:1451–1455
Hershko A, Ciechanover A (1998) The ubiquitin system. Annu Rev Biochem 67:425–479
Zheng N, Shabek N (2017) Ubiquitin ligases: structure, function, and regulation. Annu Rev Biochem 86:129–157
Varshavsky A (2017) The ubiquitin system, autophagy, and regulated protein degradation. Annu Rev Biochem 86:123–128
Kwon YT, Ciechanover A (2017) The ubiquitin code in the ubiquitin-proteasome system and autophagy. Trends Biochem Sci 42:873–886
Wenzel DM, Lissounov A, Brzovic PS, Klevit RE (2011) UBCH7 reactivity profile reveals parkin and HHARI to be RING/HECT hybrids. Nature 474:105–108
Dye BT, Schulman BA (2007) Structural mechanisms underlying posttranslational modification by ubiquitin-like proteins. Annu Rev Biophys Biomol Struct 36:131–150
Wenzel DM, Klevit RE (2012) Following Ariadne’s thread: a new perspective on RBR ubiquitin ligases. BMC Biol 10:24
Ordureau A et al (2014) Quantitative proteomics reveal a feedforward mechanism for mitochondrial pARKIN translocation and ubiquitin chain synthesis. Mol Cell 56:360–375
Ordureau A et al (2020) Global landscape and dynamics of Parkin and USP30-dependent ubiquitylomes in ineurons during mitophagic signaling. Mol Cell 77:1124-1142.e10
Chan NC et al (2011) Broad activation of the ubiquitin-proteasome system by Parkin is critical for mitophagy. Hum Mol Genet 20:1726–1737
Walden H, Martinez-Torres RJ (2012) Regulation of Parkin E3 ubiquitin ligase activity. Cell Mol Life Sci 69:3053–3067
Chaugule VK et al (2011) Autoregulation of Parkin activity through its ubiquitin-like domain. EMBO J 30:2853
Wauer T, Komander D (2013) Structure of the human Parkin ligase domain in an autoinhibited state. EMBO J 32:2099–2112
Kumar A et al (2017) Parkin-phosphoubiquitin complex reveals cryptic ubiquitin-binding site required for RBR ligase activity. Nat Struct Mol Biol 24:475–483
McWilliams TG et al (2018) Phosphorylation of Parkin at serine 65 is essential for its activation in vivo. Open Biol 8:180108
Kumar A et al (2015) Disruption of the autoinhibited state primes the E3 ligase parkin for activation and catalysis. EMBO J 34:2506–2521
Wauer T, Simicek M, Schubert A, Komander D (2015) Mechanism of phospho-ubiquitin-induced PARKIN activation. Nature 524:370–374
Sauvé V et al (2015) A Ubl/ubiquitin switch in the activation of Parkin. EMBO J 34:2492–2505
Sauvé V et al (2018) Mechanism of parkin activation by phosphorylation. Nat Struct Mol Biol 25:623–630
Sauvé V et al (2022) Structural basis for feedforward control in the PINK1/Parkin pathway. EMBO J 41:e109460
Fakih R, Sauvé V, Gehring K (2022) Structure of the second phosphoubiquitin-binding site in parkin. J Biol Chem 298:102114
Kondapalli C et al (2012) PINK1 is activated by mitochondrial membrane potential depolarization and stimulates Parkin E3 ligase activity by phosphorylating Serine 65. Open Biol 2:120080
Iguchi M et al (2013) Parkin-catalyzed ubiquitin-ester transfer is triggered by PINK1-dependent phosphorylation. J Biol Chem 288:22019–22032
Schubert AF et al (2017) Structure of PINK1 in complex with its substrate ubiquitin. Nature 552:51–56
Shiba-Fukushima K et al (2012) PINK1-mediated phosphorylation of the Parkin ubiquitin-like domain primes mitochondrial translocation of Parkin and regulates mitophagy. Sci Rep 2:1–8
Herst PM, Rowe MR, Carson GM, Berridge MV (2017) Functional mitochondria in health and disease. Front Endocrinol (Lausanne) 8:296
Friedman JR, Nunnari J (2014) Mitochondrial form and function. Nature 505:335–343
Scheibye-Knudsen M, Fang EF, Croteau DL, Wilson DM, Bohr VA (2015) Protecting the mitochondrial powerhouse. Trends Cell Biol 25:158–170
Cheng YW, Liu J, Finkel T (2023) Mitohormesis. Cell Metab 35:1872–1886
Karbowski M, Oshima Y, Verhoeven N (2022) Mitochondrial proteotoxicity: implications and ubiquitin-dependent quality control mechanisms. Cell Mol Life Sci 79:1–23
Liu S, Liu S, Jiang H (2022) Multifaceted roles of mitochondrial stress responses under ETC dysfunction—repair, destruction and pathogenesis. FEBS J 289:6994–7013
Ploumi C, Daskalaki I, Tavernarakis N (2017) Mitochondrial biogenesis and clearance: a balancing act. FEBS J 284:183–195
Gottlieb RA et al (2021) At the heart of mitochondrial quality control: many roads to the top. Cell Mol Life Sci 78:3791–3801
Shen Y et al (2023) Mitochondrial dynamics in neurological diseases: a narrative review. Ann Transl Med 11:264–264
Liu X, Weaver D, Shirihai O, Hajnóczky G (2009) Mitochondrial ‘kiss-and-run’’: interplay between mitochondrial motility and fusion–fission dynamics’. EMBO J 28:3074
Wolf DM et al (2019) Individual cristae within the same mitochondrion display different membrane potentials and are functionally independent. EMBO J 38:e101056
Loriette V et al (2023) Dynamics of mitochondrial membranes under photo-oxidative stress with high spatiotemporal resolution. Front Cell Dev Biol 11:1307502
Twig G et al (2008) Fission and selective fusion govern mitochondrial segregation and elimination by autophagy. EMBO J 27:433–446
Wang X et al (2011) PINK1 and Parkin target miro for phosphorylation and degradation to arrest mitochondrial motility. Cell 147:893–906
Shlevkov E, Kramer T, Schapansky J, Lavoie MJ, Schwarz TL (2016) Miro phosphorylation sites regulate Parkin recruitment and mitochondrial motility. Proc Natl Acad Sci U S A 113:E6097–E6106
Gegg ME et al (2010) Mitofusin 1 and mitofusin 2 are ubiquitinated in a PINK1/parkin-dependent manner upon induction of mitophagy. Hum Mol Genet 19:4861–4870
Tanaka A et al (2010) Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J Cell Biol 191:1367–1380
Buhlman L et al (2014) Functional interplay between Parkin and Drp1 in mitochondrial fission and clearance. Biochim Biophys Acta 1843:2012–2026
Pryde KR, Smith HL, Chau KY, Schapira AHV (2016) PINK1 disables the anti-fission machinery to segregate damaged mitochondria for mitophagy. J Cell Biol 213:163–171
Xu J, Deng Z, Shang S, Wang C, Han H (2023) FUNDC1 collaborates with PINK1 in regulating mitochondrial Fission and compensating for PINK1 deficiency. Biochem Biophys Res Commun 687:149210
Chen X, Wang Q, Li S, Li XJ, Yang W (2022) Mitochondrial-Dependent and Independent Functions of PINK1. Front Cell Dev Biol 10:954536
Burman JL et al (2017) Mitochondrial fission facilitates the selective mitophagy of protein aggregates. J Cell Biol 216:3231–3247
Greene JC et al (2003) Mitochondrial pathology and apoptotic muscle degeneration in Drosophila parkin mutants. Proc Natl Acad Sci U S A 100:4078–4083
Clark IE et al (2006) Drosophila pink1 is required for mitochondrial function and interacts genetically with parkin. Nature 441:1162–1166
Vincow ES et al (2013) The PINK1-Parkin pathway promotes both mitophagy and selective respiratory chain turnover in vivo. Proc Natl Acad Sci U S A 110:6400–6405
Valente EM et al (2004) Hereditary early-onset Parkinson’s disease caused by mutations in PINK1. Science 304:1158–1160
Kilarski LL et al (2012) Systematic review and UK-based study of PARK2 (parkin), PINK1, PARK7 (DJ-1) and LRRK2 in early-onset Parkinson’s disease. Mov Disord 27:1522–1529
Neupert W, Herrmann JM (2007) Translocation of proteins into mitochondria. Annu Rev Biochem 76:723–749
Greene AW et al (2012) Mitochondrial processing peptidase regulates PINK1 processing, import and Parkin recruitment. EMBO Rep 13:378–385
Kunová N et al (2022) Mitochondrial Processing Peptidases-Structure, Function and the Role in Human Diseases. Int J Mol Sci 23:1297
Jin SM et al (2010) Mitochondrial membrane potential regulates PINK1 import and proteolytic destabilization by PARL. J Cell Biol 191:933–942
Meissner C, Lorenz H, Weihofen A, Selkoe DJ, Lemberg MK (2011) The mitochondrial intramembrane protease PARL cleaves human Pink1 to regulate Pink1 trafficking. J Neurochem 117:856–867
Yamano K, Youle RJ (2013) PINK1 is degraded through the N-end rule pathway. Autophagy 9:1758–1769
Narendra D, Tanaka A, Suen DF, Youle RJ (2008) Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J Cell Biol 183:795–803
Michaelis JB et al (2022) Protein import motor complex reacts to mitochondrial misfolding by reducing protein import and activating mitophagy. Nat Commun 13:5164
Jin SM, Youle RJ (2013) The accumulation of misfolded proteins in the mitochondrial matrix is sensed by PINK1 to induce PARK2/Parkin-mediated mitophagy of polarized mitochondria. Autophagy 9:1750–1757
Rubio-Peña K et al (2020) Mitophagy of polarized sperm-derived mitochondria after fertilization. iScience 24:102029
Lazarou M, Jin SM, Kane LA, Youle RJ (2012) Role of PINK1 binding to the TOM complex and alternate intracellular membranes in recruitment and activation of the E3 ligase Parkin. Dev Cell 22:320–333
Okatsu K et al (2012) PINK1 autophosphorylation upon membrane potential dissipation is essential for Parkin recruitment to damaged mitochondria. Nat Commun 3:1016
Aerts L, Craessaerts K, De Strooper B, Morais VA (2015) PINK1 kinase catalytic activity is regulated by phosphorylation on serines 228 and 402. J Biol Chem 290:2798–2811
Gan ZY et al (2021) Activation mechanism of PINK1. Nature 602:328–335
Rasool S et al (2022) Mechanism of PINK1 activation by autophosphorylation and insights into assembly on the TOM complex. Mol Cell 82:44-59.e6
Herzig S, Shaw RJ (2017) (2017) AMPK: guardian of metabolism and mitochondrial homeostasis. Nat Rev Mol Cell Biol 19:121–135
Hung CM et al (2021) AMPK/ULK1-mediated phosphorylation of Parkin ACT domain mediates an early step in mitophagy. Sci Adv 7:eabg4544
Kazlauskaite A et al (2014) Accelerated publication: Parkin is activated by PINK1-dependent phosphorylation of ubiquitin at Ser65. Biochem J 460:127–139
Koyano F et al (2014) (2014) Ubiquitin is phosphorylated by PINK1 to activate Parkin. Nature 510:162–166
Kane LA et al (2014) PINK1 phosphorylates ubiquitin to activate Parkin E3 ubiquitin ligase activity. J Cell Biol 205:143–153
Wauer T et al (2015) Ubiquitin Ser65 phosphorylation affects ubiquitin structure, chain assembly and hydrolysis. EMBO J 34:307–325
Koyano F, Yamano K, Kosako H, Tanaka K, Matsuda N (2019) Parkin recruitment to impaired mitochondria for nonselective ubiquitylation is facilitated by MITOL. J Biol Chem 294:10300–10314
Ordureau A et al (2015) Defining roles of PARKIN and ubiquitin phosphorylation by PINK1 in mitochondrial quality control using a ubiquitin replacement strategy. Proc Natl Acad Sci U S A 112:6637–6642
Rose CM et al (2016) Highly multiplexed quantitative mass spectrometry analysis of ubiquitylomes. Cell Syst 3:395-403.e4
Sarraf SA et al (2013) Landscape of the PARKIN-dependent ubiquitylome in response to mitochondrial depolarization. Nature 496:372–376
Wang L, Wang J, Tang Y, Shen HM (2018) PTEN-L puts a brake on mitophagy. Autophagy 14:2023–2025
Wang L et al (2018) PTEN-L is a novel protein phosphatase for ubiquitin dephosphorylation to inhibit PINK1-Parkin-mediated mitophagy. Cell Res 28:787–802
Suresh HG, Pascoe N, Andrews B (2020) The structure and function of deubiquitinases: lessons from budding yeast. Open Biol 10:200279
Clague MJ, Urbé S, Komander D (2019) Breaking the chains: deubiquitylating enzyme specificity begets function. Nat Rev Mol Cell Biol 20:338–352
Okarmus J et al (2024) (2014) USP30 inhibition induces mitophagy and reduces oxidative stress in parkin-deficient human neurons. Cell Death Dis 15:1–13
Lange SM, Armstrong LA, Kulathu Y (2022) Deubiquitinases: From mechanisms to their inhibition by small molecules. Mol Cell 82:15–29
Silvian LF (2022) PINK1/Parkin Pathway activation for mitochondrial quality control—which is the best molecular target for therapy? Front Aging Neurosci 14:890823
Tsefou E, Ketteler R (2022) Targeting deubiquitinating enzymes (DUBs) that regulate mitophagy via direct or indirect interaction with Parkin. Int J Mol Sci 23:12105
Sato Y et al (2017) Structural basis for specific cleavage of Lys6-linked polyubiquitin chains by USP30. Nat Struct Mol Biol 24:911–919
Gersch M et al (2017) Mechanism and regulation of the Lys6-selective deubiquitinase USP30. Nat Struct Mol Biol 24:920–930
Bingol B et al (2014) (2014) The mitochondrial deubiquitinase USP30 opposes parkin-mediated mitophagy. Nature 510:370–375
Chakraborty J, Ziviani, E (2020) Deubiquitinating enzymes in Parkinson’s disease. Front Physiol 11:535
Durcan TM, Kontogiannea M, Bedard N, Wing SS, Fon EA (2012) Ataxin-3 deubiquitination is coupled to Parkin ubiquitination via E2 ubiquitin-conjugating enzyme. J Biol Chem 287:531–541
Durcan TM, Fon EA (2011) Mutant ataxin-3 promotes the autophagic degradation of parkin. Autophagy 7:233–234
Durcan TM et al (2014) USP8 regulates mitophagy by removing K6-linked ubiquitin conjugates from parkin. EMBO J 33:2473–2491
Cornelissen T et al (2018) Deficiency of parkin and PINK1 impairs age-dependent mitophagy in Drosophila. Elife 7:e35878
Cornelissen T et al (2014) The deubiquitinase USP15 antagonizes Parkin-mediated mitochondrial ubiquitination and mitophagy. Hum Mol Genet 23:5227–5242
Wang F et al (2004) Parkin gene alterations in hepatocellular carcinoma. Genes Chromosomes Cancer 40:85–96
Bernardini JP, Lazarou M, Dewson G (2017) Parkin and mitophagy in cancer. Oncogene 36:1315–1327
Shin JH et al (2011) PARIS (ZNF746) repression of PGC-1α contributes to neurodegeneration in Parkinson’s disease. Cell 144:689–702
Finck BN, Kelly DP (2006) PGC-1 coactivators: inducible regulators of energy metabolism in health and disease. J Clin Invest 116:615–622
Manzanillo PS et al (2013) The ubiquitin ligase parkin mediates resistance to intracellular pathogens. Nature 501:512–516
Ali S et al (2006) PARK2/PACRG polymorphisms and susceptibility to typhoid and paratyphoid fever. Clin Exp Immunol 144:425–431
Mira MT et al (2004) Susceptibility to leprosy is associated with PARK2 and PACRG. Nature 427:636–640
Yamano K et al (2024) Optineurin provides a mitophagy contact site for TBK1 activation. EMBO J 43:754
Pentzold C et al (2018) FANCD2 binding identifies conserved fragile sites at large transcribed genes in avian cells. Nucleic Acids Res 46:1280–1294
Voutsinos V, Munk SHN, Oestergaard VH (2018) Common chromosomal fragile sites-conserved failure stories. Genes (Basel) 9:580
Munk SHN, Voutsinos V, Oestergaard VH (2021) Large intronic deletion of the fragile site gene PRKN dramatically lowers its fragility without impacting gene expression. Front Genet 12:695172
Hattori N, Mizuno Y (2017) Twenty years since the discovery of the parkin gene. J Neural Transm (Vienna) 124:1037–1054
Landrum MJ et al (2018) ClinVar: improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46:D1062–D1067
Lill CM et al (2016) Launching the movement disorders society genetic mutation database (MDSGene). Mov Disord 31:607–609
Sauna ZE, Kimchi-Sarfaty C (2011) Understanding the contribution of synonymous mutations to human disease. Nat Rev Genet 12(10):683–691
Kasten M et al (2018) Genotype-phenotype relations for the Parkinson’s disease genes parkin, PINK1, DJ1: MDSGene systematic review. Mov Disord 33:730–741
Cagiada M et al (2021) Understanding the origins of loss of protein function by analyzing the effects of thousands of variants on activity and abundance. Mol Biol Evol 38:3235–3246
Wang C et al (2005) Alterations in the solubility and intracellular localization of parkin by several familial Parkinson’s disease-linked point mutations. J Neurochem 93:422–431
Fiesel FC et al (2015) Structural and functional impact of Parkinson disease-associated mutations in the E3 ubiquitin ligase parkin. Hum Mutat 36:774–786
Henn IH, Gostner JM, Lackner P, Tatzelt J, Winklhofer KF (2005) Pathogenic mutations inactivate parkin by distinct mechanisms. J Neurochem 92:114–122
Zhang BR et al (2010) Mutation analysis of parkin and PINK1 genes in early-onset Parkinson’s disease in China. Neurosci Lett 477:19–22
Abbas N et al (1999) A wide variety of mutations in the parkin gene are responsible for autosomal recessive parkinsonism in Europe. French Parkinson’s Disease Genetics Study Group and the European Consortium on Genetic Susceptibility in Parkinson’s Disease. Hum Mol Genet 8:567–574
Sriram SR et al (2005) Familial-associated mutations differentially disrupt the solubility, localization, binding and ubiquitination properties of parkin. Hum Mol Genet 14:2571–2586
Terreni L, Calabrese E, Calella AM, Forloni G, Mariani C (2001) New mutation (R42P) of the parkin gene in the ubiquitinlike domain associated with Parkinsonism. Neurology 56:463–466
Safadi SS, Shaw GS (2007) A disease state mutation unfolds the parkin ubiquitin-like domain. Biochemistry 46:14162–14169
Hampe C, Ardila-Osorio H, Fournier M, Brice A, Corti O (2006) Biochemical analysis of Parkinson’s disease-causing variants of Parkin, an E3 ubiquitin-protein ligase with monoubiquitylation capacity. Hum Mol Genet 15:2059–2075
Jensen LD, Vinther-Jensen T, Kahns S, Sundbye S, Jensen PH (2006) Cellular parkin mutants are soluble under non-stress conditions. NeuroReport 17:1205–1208
Clausen L et al (2024) A mutational atlas for Parkin proteostasis. Nat Commun 15:1541
Muller HJ (1932) Further studies on the nature and causes of gene mutations. In: Proceedings of the 6th International Congress of genetics, pp 213–255
Okiyoneda T et al (2013) Mechanism-based corrector combination restores ΔF508-CFTR folding and function. Nat Chem Biol 9:444–454
Brusa I et al (2022) Proteostasis regulators in cystic fibrosis: current development and future perspectives. J Med Chem 65:5212–5243
Hutt DM et al (2010) Reduced histone deacetylase 7 activity restores function to misfolded CFTR in cystic fibrosis. Nat Chem Biol 6:25–33
Arlow T, Scott K, Wagenseller A, Gammie A (2013) Proteasome inhibition rescues clinically significant unstable variants of the mismatch repair protein Msh2. Proc Natl Acad Sci U S A 110:246–251
Kampmeyer C et al (2017) Blocking protein quality control to counter hereditary cancers. Genes Chromosomes Cancer 56:823–831
Yin Y, Moult J (2019) Characterizing and comparing missense variants in monogenic disease and in cancer. bioRxiv. https://doi.org/10.1101/534693
Stein A, Fowler DM, Hartmann-Petersen R, Lindorff-Larsen K (2019) Biophysical and mechanistic models for disease-causing protein variants. Trends Biochem Sci 44:575–588
Clausen L et al (2019) Protein stability and degradation in health and disease. Adv Protein Chem Struct Biol 114:61–83
Bartlett AI, Radford SE (2009) An expanding arsenal of experimental methods yields an explosion of insights into protein folding mechanisms. Nat Struct Mol Biol 16:582–588
Kim YE, Hipp MS, Bracher A, Hayer-Hartl M, Ulrich Hartl F (2013) Molecular chaperone functions in protein folding and proteostasis. Annu Rev Biochem 82:323–355
Maxwell KL et al (2005) Protein folding: defining a ‘standard’ set of experimental conditions and a preliminary kinetic data set of two-state proteins. Protein Sci 14:602–616
Kovalchuke L, Mosharov EV, Levy OA, Greene LA (2019) Stress-induced phospho-ubiquitin formation causes parkin degradation. Sci Rep 9:11682
Park HM et al (2009) The serine protease HtrA2/Omi cleaves Parkin and irreversibly inactivates its E3 ubiquitin ligase activity. Biochem Biophys Res Commun 387:537–542
Sonia Angeline M, Chaterjee P, Anand K, Ambasta RK, Kumar P (2012) Rotenone-induced parkinsonism elicits behavioral impairments and differential expression of parkin, heat shock proteins and caspases in the rat. Neuroscience 220:291–301
Aimé P et al (2015) Trib3 Is Elevated in Parkinson’s disease and mediates death in Parkinson’s disease models. J Neurosci 35:10731
Sun X et al (2013) ATF4 protects against neuronal death in cellular Parkinson’s disease models by maintaining levels of parkin. J Neurosci 33:2398
Rakovic A et al (2013) Phosphatase and tensin homolog (PTEN)-induced putative kinase 1 (PINK1)-dependent ubiquitination of endogenous Parkin attenuates mitophagy: study in human primary fibroblasts and induced pluripotent stem cell-derived neurons. J Biol Chem 288:2223–2237
Soleimanpour SA et al (2014) The diabetes susceptibility gene Clec16a regulates mitophagy. Cell 157:1577
Potting C et al (2018) Genome-wide CRISPR screen for PARKIN regulators reveals transcriptional repression as a determinant of mitophagy. Proc Natl Acad Sci USA 115:E180–E189
McLelland GL, Soubannier V, Chen CX, McBride HM, Fon EA (2014) Parkin and PINK1 function in a vesicular trafficking pathway regulating mitochondrial quality control. EMBO J 33:282–295
Anfinsen CB, Haber E, Sela M, White FH (1961) The kinetics of formation of native ribonuclease during oxidation of the reduced polypeptide chain. Proc Natl Acad Sci U S A 47:1309–1314
Hartl FU, Bracher A, Hayer-Hartl M (2011) Molecular chaperones in protein folding and proteostasis. Nature 475:324–332
Hartl FU, Hayer-Hartl M (2009) Converging concepts of protein folding in vitro and in vivo. Nat Struct Mol Biol 16:574–581
Rosenzweig R, Nillegoda NB, Mayer MP, Bukau B (2019) The Hsp70 chaperone network. Nat Rev Mol Cell Biol 20:665–680
Hickey CM, Breckel C, Zhang M, Theune WC, Hochstrasser M (2021) Protein quality control degron-containing substrates are differentially targeted in the cytoplasm and nucleus by ubiquitin ligases. Genetics 217:1–19
Enam C, Geffen Y, Ravid T, Gardner RG (2018) Protein quality control degradation in the nucleus. Annu Rev Biochem 87:725–749
Abildgaard AB et al (2020) Co-Chaperones in targeting and delivery of misfolded proteins to the 26S proteasome. Biomolecules 10:1–24
Alberti S, Esser C, Höhfeld J (2003) BAG-1–a nucleotide exchange factor of Hsc70 with multiple cellular functions. Cell Stress Chaperones 8:225–231
Andersen KM, Hofmann K, Hartmann-Petersen R (2005) Ubiquitin-binding proteins: similar, but different. Essays Biochem 41:49–67
Rose JM, Novoselov SS, Robinson PA, Cheetham ME (2011) Molecular chaperone-mediated rescue of mitophagy by a Parkin RING1 domain mutant. Hum Mol Genet 20:16–27
Schlehe JS et al (2008) Aberrant folding of pathogenic Parkin mutants: aggregation versus degradation. J Biol Chem 283:13771–13779
Zhang CW et al (2016) Pharmacological or genetic activation of Hsp70 protects against loss of parkin function. Neurodegener Dis 16:304–316
Winklhofer KF, Henn IH, Kay-Jackson PC, Heller U, Tatzelt J (2003) Inactivation of parkin by oxidative stress and C-terminal truncations: a protective role of molecular chaperones. J Biol Chem 278:47199–47208
Kalia SK et al (2004) BAG5 inhibits parkin and enhances dopaminergic neuron degeneration. Neuron 44:931–945
Hu S et al (2021) Molecular chaperones and Parkinson’s disease. Neurobiol Dis 160:105527
Kalia KS, Kalia LV, McLean JP (2010) Molecular chaperones as rational drug targets for Parkinson’s disease therapeutics. CNS Neurol Disord Drug Targets 9:741–753
Kampmeyer C et al (2022) Disease-linked mutations cause exposure of a protein quality control degron. Structure 30:1245-1253.e5
Timms RT, Koren I (2020) Tying up loose ends: the N-degron and C-degron pathways of protein degradation. Biochem Soc Trans 48:1557–1567
Johansson KE, Mashahreh B, Hartmann-Petersen R, Ravid T, Lindorff-Larsen K (2023) Prediction of quality-control degradation signals in yeast proteins. J Mol Biol 435:167915
Mashahreh B et al (2022) Conserved degronome features governing quality control associated proteolysis. Nat Commun 13:7588
Koren I et al (2018) The eukaryotic proteome is shaped by E3 ubiquitin ligases targeting C-terminal degrons. Cell 173:1622-1635.e14
Abildgaard AB et al (2023) HSP70-binding motifs function as protein quality control degrons. Cell Mol Life Sci 80:32
Maurer MJ et al (2016) Degradation signals for ubiquitin-proteasome dependent cytosolic protein quality control (CytoQC) in yeast. G3 (Bethesda) 6:1853–1866
Zhang Z et al (2023) Elucidation of E3 ubiquitin ligase specificity through proteome-wide internal degron mapping. Mol Cell 83:3377-3392.e6
Ravid T, Hochstrasser M (2008) Diversity of degradation signals in the ubiquitin-proteasome system. Nat Rev Mol Cell Biol 9:679–689
Stevens MU et al (2023) Structure-based design and characterization of Parkin-activating mutations. Life Sci Alliance 6:e202201419
Van Hout CV et al (2020) Exome sequencing and characterization of 49,960 individuals in the UK Biobank. Nature 586:749–756
Starita LM et al (2017) Variant interpretation: functional assays to the rescue. Am J Human Genet 101:315–325
Weile J, Roth FP (2018) Multiplexed assays of variant effects contribute to a growing genotype-phenotype atlas. Hum Genet 137:665–678
Fowler DM et al (2023) An Atlas of Variant Effects to understand the genome at nucleotide resolution. Genome Biol 24:147
Cheng J et al (2023) Accurate proteome-wide missense variant effect prediction with AlphaMissense. Science 381:eadg7492
Frazer J et al (2021) Disease variant prediction with deep generative models of evolutionary data. Nature 599:91–95
Tabet D, Parikh V, Mali P, Roth FP, Claussnitzer M (2022) Scalable functional assays for the interpretation of human genetic variation. Annu Rev Genet 56:441–465
Shlevkov E et al (2022) Discovery of small-molecule positive allosteric modulators of Parkin E3 ligase. iScience 25:103650
Chiti F, Kelly JW (2022) Small molecule protein binding to correct cellular folding or stabilize the native state against misfolding and aggregation. Curr Opin Struct Biol 72:267–278
Wang F et al (2022) USP30: structure, emerging physiological role, and target inhibition. Front Pharmacol 13:851654
Acknowledgements
We thank all members of the PRISM center and the Linderstrøm-Lang Centre for Protein Science for helpful discussions. Figures were created with BioRender.com.
Funding
Open access funding provided by Copenhagen University. We are funded by the Novo Nordisk Foundation (https:// novonordiskfonden.dk) challenge program PRISM (to K.L.-L., & R.H.-P.), the Lundbeck Foundation (https://www.lundbeckfonden.com) R249-2017–510 (to L.C. & R.H.-P.), the Danish Council for Independent Research (Det Frie Forskningsråd) (https://dff.dk) https://doi.org/10.46540/2032-00007B (to R.H.P.) and the Danish Parkinson Foundation (to J.O. & M.M.).
Author information
Authors and Affiliations
Contributions
L.C. prepared the figures. L.C., J.O., V.V., M.M., K.L-L., and R.H.-P. wrote and revised the paper.
Corresponding author
Ethics declarations
Conflict of interest
K.L.-L. holds stock options in and is a consultant for Peptone Ltd. All other authors have no relevant financial or nonfinancial interests to disclose.
Ethics approval and consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Clausen, L., Okarmus, J., Voutsinos, V. et al. PRKN-linked familial Parkinson’s disease: cellular and molecular mechanisms of disease-linked variants. Cell. Mol. Life Sci. 81, 223 (2024). https://doi.org/10.1007/s00018-024-05262-8
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-024-05262-8